Abstract
Key message
Genetic diversity, population structure, LD decay, and selective sweeps in 687 wheat accessions were analyzed, providing relevant guidelines to facilitate the use of the germplasm in wheat breeding.
Abstract
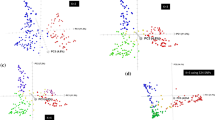
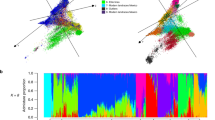
Common wheat (Triticum aestivum L.) is one of the most widely grown crops in the world. Landraces were subjected to strong human-mediated selection in developing high-yielding, good quality, and widely adapted cultivars. To investigate the genome-wide patterns of allelic variation, population structure and patterns of selective sweeps during modern wheat breeding, we tested 687 wheat accessions, including landraces (148) and cultivars (539) mainly from China and Pakistan in a wheat 90 K single nucleotide polymorphism array. Population structure analysis revealed that cultivars and landraces from China and Pakistan comprised three relatively independent genetic clusters. Cultivars displayed lower nucleotide diversity and a wider average LD decay across whole genome, indicating allelic erosion and a diversity bottleneck due to the modern breeding. Analysis of genetic differentiation between landraces and cultivars from China and Pakistan identified allelic variants subjected to selection during modern breeding. In total, 477 unique genome regions showed signatures of selection, where 109 were identified in both China and Pakistan germplasm. The majority of genomic regions were located in the B genome (225), followed by the A genome (175), and only 77 regions were located in the D genome. EigenGWAS was further used to identify key selection loci in modern wheat cultivars from China and Pakistan by comparing with global winter wheat and spring wheat diversity panels, respectively. A few known functional genes or loci found within these genome regions corresponded to known phenotypes for disease resistance, vernalization, quality, adaptability and yield-related traits. This study uncovered molecular footprints of modern wheat breeding and explained the genetic basis of polygenic adaptation in wheat. The results will be useful for understanding targets of modern wheat breeding, and in devising future breeding strategies to target beneficial alleles currently not pursued.






Similar content being viewed by others
Abbreviations
- CL:
-
Chinese landraces
- CMC:
-
Chinese modern cultivars
- FMC:
-
Foreign modern cultivars
- F ST :
-
F-statistics
- GWAS:
-
Genome-wide association study
- IWGSC:
-
International Wheat Genome Sequencing Consortium
- LD:
-
Linkage disequilibrium
- MAF:
-
Minor allele frequency
- MAS:
-
Marker-assisted selection
- NJ:
-
Neighbor-joining
- PCA:
-
Principal components analysis
- PL:
-
Pakistan landraces
- PMC:
-
Pakistan modern cultivars
- SNP:
-
Single nucleotide polymorphism
References
Ain QU, Rasheed A, Anwar A, Mahmood T, Imtiaz M, He ZH, Quraishi UM (2015) Genome-wide association for grain yield under rainfed conditions in historical wheat cultivars from Pakistan. Front Plant Sci 6:743
Akagi T, Hanada T, Yaegaki H, Gradziel TM, Tao R (2016) Genome-wide view of genetic diversity reveals paths of selection and cultivar differentiation in peach domestication. DNA Res 23:271–282
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19:1655–1664
Allen AM, Winfield MO, Burridge AJ, Downie RC, Benbow HR, Barker GL, Scopes G (2016) Characterization of a wheat breeders, array suitable for high-throughput SNP genotyping of global accessions of hexaploid bread wheat (Triticum aestivum). Plant Biotechnol J 15:390–401
Baenziger PS, DePauw RM (2009) Wheat breeding: procedures and strategies. In: Carver BF (ed) Wheat: science and trade. Wiley-Blackwell Publishing, Ames, pp 275–308
Betran FJ, Ribaut JM, Beck D, De Leon DG (2003) Genetic diversity, specific combining ability, and heterosis in tropical maize under stress and nonstress environments. Crop Sci 43:797–806
Bonman JM, Babiker EM, Cuesta-Marcos A, Esvelt-Klos K, Brown-Guedira G, Chao S, See D, Chen JL, Akhunov E, Zhang JL, Bockelman HE, Gordon TC (2015) Genetic diversity among wheat accessions from the USDA National Small Grains Collection. Crop Sci 55:1243–1253
Bosse M, Spurgin LG, Laine VN, Cole EF, Firth JA, Gienapp P, Gosler AG, McMahon K, Poissant J, Verhagen I, Groenen MA (2017) Recent natural selection causes adaptive evolution of an avian polygenic trait. Science 358:365–368
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Brenchley R, Spannagl M, Pfeifer M, Barker GLA, D’Amore R, Allen AM, McKenzie N, Kramer M, Kerhornou A, Bolser D, Kay S, Waite D, Trick M, Bancroft I, Gu Y, Huo NX, Luo MC, Sehgal S, Gill B, Kianian S, Anderson O, Kersey P, Dvorak J, McCombie WR, Hall A, Mayer KFX, Edwards KJ, Bevan MW, Hall N (2012) Analysis of the bread wheat genome using whole-genome shotgun sequencing. Nature 491:705–710
Breseghello F, Sorrells ME (2006) Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. Genetics 172:1165–1177
Calus MPL, De Roos APW, Veerkamp RF (2008) Accuracy of genomic selection using different methods to define haplotypes. Genetics 178:553–561
Cavanagh CR, Chao S, Wang S, Huang BE, Stephen S, Kiani S, For-rest K, Saintenac C, Brown-Guedira GL, Akhunova A, See D, Bai G, Pumphrey M, Tomar L, Wong D, Kong S, Reynolds M, da Silva ML, Bockelman H, Talbert L, Anderson JA, Dreisi-gacker S, Baenziger S, Carter A, Korzun V, Morrell PL, Dubcovsky J, Morell MK, Sorrells ME, Hayden MJ, Akhunov E (2013) Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc Natl Acad Sci USA 110:8057–8062
Chao SM, Zhang WJ, Akhunov E, Sherman J, Ma YQ, Luo MC, Dubcovsky J (2009) Analysis of gene derived SNP marker polymorphism in US wheat (Triticum aestivum L.) cultivars. Mol Breed 23:23–33
Chen X, Min D, Yasir TA, Hu YG (2012) Genetic diversity, population structure and linkage disequilibrium in elite Chinese winter wheat investigated with SSR markers. PLoS ONE 7:e44510
Chen GB, Lee SH, Zhu ZX, Benyamin B, Robinson MR (2016) EigenGWAS: finding loci under selection through genome-wide association studies of eigenvectors in structured populations. Heredity 117:51–61
Choulet F, Alberti A, Theil S, Glover N, Barbe V, Daron J, Pingault L, Sourdille P, Couloux A, Paux E, Leroy P, Mangenot S, Guilhot N, Gouis JL, Balfourier F, Alaux M, Jamilloux V, Poulain J, Durand C, Bellec A, Gaspin C, Safar J, Dolezel J, Rogers J, Vandepoele K, Aury JM, Mayer K, Berges H, Quesneville H, Wincker P, Feuillet C (2014) Structural and functional partitioning of bread wheat chromosome 3B. Science 345:1249721
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Handsaker RE, Lunter G, Marth GT, McVean G, Durbin R (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of domestication. Cell 127:1309–1321
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure from small quantities of fresh leaf tissues. Phytochem Bull 19:11–15
Dvorak J, Terlizzi PD, Zhang HB, Resta P (1993) The evolution of polyploid wheats: identification of the A genome donor species. Genome 36:21–31
Edae EA, Byrne PF, Haley SD, Lopes MS, Reynolds MP (2014) Genome-wide association mapping of yield and yield components of spring wheat under contrasting moisture regimes. Theor Appl Genet 127:791–807
Flint-Garcia SA, Thornsberry JM, Buckler ES (2003) Structure of linkage disequilibrium in plants. Annu Rev Plant Biol 54:357–374
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitckhiking and background selection. Genetics 147:915–925
Gao LF, Zhao GY, Huang DW, Jia JZ (2017) Candidate loci involved in domestication and improvement detected by a published 90 K wheat SNP array. Sci Rep-UK 7:44530
Gianibelli MC, Echaide M, Larroque OR, Carrillo JM, Dubcovsky J (2002) Biochemical and molecular characterization of Glu-1 loci in Argentinean wheat cultivars. Euphytica 128:61–73
Gupta PK, Rustgi S, Mir RR (2008) Array-based high-throughput DNA markers for crop improvement. Heredity 101:5–18
Han YP, Zhao X, Liu DY, Li YH, Lightfoot DA, Yang ZJ, Zhao L, Zhou G, Wang ZK, Huang L, ZW Zhang, Qiu LJ, Zheng HK, Li WB (2016) Domestication footprints anchor genomic regions of agronomic importance in soybeans. New Phytol 209:871–884
Hao CY, Wang LF, Ge HM, Dong YC, Zhang XY (2011) Genetic diversity and linkage disequilibrium in Chinese bread wheat (Triticum aestivum L.) revealed by SSR markers. PLoS ONE 6:e17279
Hao CY, Wang YQ, Chao SAM, Li T, Liu HX, Wang LF, Zhang XY (2017) The iSelect 9 K SNP analysis revealed polyploidization induced revolutionary changes and intense human selection causing strong haplotype blocks in wheat. Sci Rep-UK 7:41247
Heffner EL, Sorrells ME, Jannink JL (2009) Genomic selection for crop improvement. Crop Sci 49:1–12
Helyar SJ, Hemmer-Hansen J, Bekkevold D, Taylor MI, Ogden R, Limborg MT, Cariani A, Maes JE, Diopere E, Carvalho GR, Nielsen EE (2011) Application of SNPs for population genetics of nonmodel organisms: new opportunities and challenges. Mol Ecol Resour 11:123–136
Heslot N, Jannink JL, Sorrells ME (2015) Perspectives for genomic selection applications and research in plants. Crop Sci 55:1–12
Hill TA, Ashrafi H, Reyes-Chin-Wo S, Yao J, Stoffel K, Truco MJ, Kozik A, Michelmore RW, Deynze AV, Van Deynze A (2013) Characterization of Capsicum annuum genetic diversity and population structure based on parallel polymorphism discovery with a 30 K unigene pepper gene chip. PLoS ONE 8:e56200
Holsinger KE, Weir BS (2009) Genetics in geographically structured populations: defining, estimating and interpreting F ST. Nat Rev Genet 10:639–650
Hudson RR, Slatkin M, Maddison WP (1992) Estimation of levels of gene flow from DNA sequence data. Genetics 132:583–589
Hulse-Kemp AM, Lemm J, Plieske J, Ashrafi H, Buyyarapu R, Fang DD et al (2015) Development of a 63 K SNP array for cotton and high-density mapping of intra-and inter-specific populations of Gossypium spp. G3-Genes Genom Genet 5:1187–1209
Hutter S, Vilella AJ, Rozas J (2006) Genome-wide DNA polymorphism analyses using VariScan. BMC Bioinform 7:1–10
International Wheat Genome Sequencing Consortium (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788
Jiao YP, Zhao HN, Ren LH, Song WB, Zeng B, Guo JJ, Wang BB, Liu ZP, Chen J, Zhang M, Lai JS (2012) Genome-wide genetic changes during modern breeding of maize. Nat Genet 44:812
Kihara H (1944) Discovery of the DD-analyser, one of the ancestors of Triticum vulgare. Agric Hortic 19:889–890
Kim KW, Bennison C, Hemmings N, Brookes L, Hurley LL, Griffith SC, Burke T, Birkhead TR, Slate J (2017) A sex-linked supergene controls sperm morphology and swimming speed in a songbird. Nat Ecol Evol 1:1168–1176
Li YH, Zhao SC, Ma JX, Li D, Yan L, Li J, Chang RZ et al (2013) Molecular footprints of domestication and improvement in soybean revealed by whole genome re-sequencing. BMC Genom 14:579
Li J, Wan HS, Yang WY (2014) Synthetic hexaploid wheat enhances variation and adaptive evolution of bread wheat in breeding processes. J Syst Evol 52:735–742
Liu K, Muse SV (2005) PowerMarker, an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Lopes M, Dreisigacker S, Peña R, Sukumaran S, Reynolds M (2014) Genetic characterization of the Wheat Association Mapping Initiative (WAMI) panel for dissection of complex traits in spring wheat. Theor Appl Genet 128:453–464
Lopes MS, El-Basyoni I, Baenziger PS, Singh S, Royo C, Ozbek K, Aktas H, Ozer E, Ozdemir F, Manickavelu A, Ban T, Vikram P (2015) Exploiting genetic diversity from landraces in wheat breeding for adaptation to climate change. J Exp Bot 66:3477–3486
Luo MC, Yang ZL, You FM, Kawahara T, Waines JG, Dvorak J (2007) The structure of wild and domesticated emmer wheat populations, gene flow between them, and the site of emmer domestication. Theor Appl Genet 114:947–959
Lupton FGH (ed) (1987) Wheat breeding: its scientific basis. Chapman and Hall Ltd, London
Marcussen T, Sandve SR, Heier L, Spannagl M, Pfeifer M, Jakobsen KS, Jakobsen KS, The International Wheat Genome Sequencing Consortium, Wulff BBH, Steuernage B, Mayer KFX, Olsen OA (2014) Ancient hybridizations among the ancestral genomes of bread wheat. Science 345:1250092
Mascher M, Gundlach H, Himmelbach A, Beier S, Twardziok SO, Wicker T et al (2017) A chromosome conformation capture ordered sequence of the barley genome. Nature 544:427–433
McFadden ES, Sears ER (1946) The origin of Triticum spelta and its free-threshing hexaploid relatives. J Hered 37:107–116
Meyer RS, Purugganan MD (2013) Evolution of crop species: genetics of domestication and diversification. Nat Rev Genet 14:840–852
Miranda LM, Murphy JP, Marshall D, Leath S (2006) Pm34: a new powdery mildew resistance gene transferred from Aegilops tauschii Coss. to common wheat (Triticum aestivum L.). Theor Appl Genet 113:1497–1504
Morrell PL, Buckler ES, Ross-Ibarra J (2011) Crop genomics: advances and applications. Nat Rev Genet 13:85–96
Narum SR, Hess JE (2011) Comparison of F ST outlier tests for SNP loci under selection. Mol Ecol Resour 11:184–194
Olson EL, Rouse MN, Pumphrey MO, Bowden RL, Gill BS, Poland JA (2013) Introgression of stem rust resistance genes SrTA10187 and SrTA10171 from Aegilops tauschii to wheat. Theor Appl Genet 126:2477–2484
Parolo S, Lacroix S, Kaput J, Scott-Boyer MP (2017) Ancestors’ dietary patterns and environments could drive positive selection in genes involved in micronutrient metabolism-the case of cofactor transporters. Gene Nutr 12:28
Peng JX, Ronin Y, Fahima T, Röder MS, Li YC, Nevo E, Korol A (2003) Domestication quantitative trait loci in Triticum dicoccoides, the progenitor of wheat. Proc Natl Acad Sci USA 100:2489–2494
Petersen G, Seberg O, Yde M, Berthelsen K (2006) Phylogenetic relationships of Triticum and Aegilops and evidence for the origin of the A, B, and D genomes of common wheat (Triticum aestivum). Mol Phylogenet Evol 39:70–82
Pfeifer M, Kugler KG, Sandve SR, Zhan B, Rudi H, Hvidsten TR, International Wheat Genome Sequencing Consortium, Mayer KFX, Olsen OA (2014) Genome interplay in the grain transcriptome of hexaploid bread wheat. Science 345:1250091
Pickrell JK, Coop G, Novembre J, Kudaravalli S, Li JZ, Absher D, Srinivasan BS, Barsh GS, Myers RM, Feldman MW, Pritchard JK (2009) Signals of recent positive selection in a worldwide sample of human populations. Genome Res 19:826–837
Qin L, Hao CY, Hou J, Wang YQ, Li T, Wang LF, Ma ZQ, Zhang XY (2014) Homologous haplotypes, expression, genetic effects and geographic distribution of the wheat yield gene TaGW2. BMC Plant Biol 14:107
Ralph P, Coop G (2010) Parallel adaptation: one or many waves of advance of an advantageous allele? Genetics 186:647–668
Rasheed A, Xia XC, Mahmood T, Quraishi UM, Aziz A, Bux H, Mahmood Z, Mirza JI, Mujeeb-Kazi A, He ZH (2016) Comparison of economically important loci in landraces and improved wheat cultivars from Pakistan. Crop Sci 56:1–15
Rasheed A, Mujeeb-Kazi A, Ogbonnaya FC, He ZH, Rajaram S (2017) Wheat genetic resources in the post-genomics era: promise and challenges. Ann Bot-Lond 121:603–616
Rasheed A, Ogbonnaya FC, Lagudah E, Appels R, He ZH (2018) The goat grass genome’s role in wheat improvement. Nat Plants 4:56
Sukumaran S, Dreisigacker S, Lopes M, Chavez P, Reynolds MP (2015) Genome-wide association study for grain yield and related traits in an elite spring wheat population grown in temperate irrigated environments. Theor Appl Genet 128:353–363
Swarts K, Gutaker RM, Benz B, Blake M, Bukowski R, Holland J, Kruse-Peeples M, Lepak N, Prim L, Romay MC, Ross-Ibarra J, Gonzalez JJS, Schmidt C, Schuenemann VJ, Krause J, Matson RG, Weige D, Buckler ES, Burbano HA (2017) Genomic estimation of complex traits reveals ancient maize adaptation to temperate North America. Science 57:512–515
Tajima F (1983) Evolutionary relationship of DNA sequences in finite populations. Genetics 105:437–460
Tanksley SD, McCouch SR (1997) Seed banks and molecular maps: unlocking genetic potential from the wild. Science 277:1063–1066
Tanno KI, Willcox G (2006) How fast was wild wheat domesticated. Science 311:1886
Trethowan RM, Mujeeb-Kazi A (2008) Novel germplasm resources for improving environmental stress tolerance of hexaploid wheat. Crop Sci 48:1255–1265
van Heerwaarden J, Hufford MB, Ross-Ibarra J (2012) Historical genomics of North American maize. Proc Natl Acad Sci USA 109:12420–12425
Wang S, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L, Mastrangelo AM, Stephen S, Barker G, Wieseke R, Plieske J, International Wheat Genome Sequencing Consortium, Lillemo M, Mather D, Appels R, Dulferos R, Brown-Guedira G, Korol A, Akhunova AR, Feuillet C, Salse J, Morgante M, Pozniak C, Luo MC, Dvorak J, Morell M, Dubcovsky J, Ganal M, Tuberosa R, Lawley C, Mikoulitch I, Cavanagh C, Edwards KJ, Hayden M, Akhunov E (2014) Characterization of polyploid wheat genomic diversity using the high-density 90,000 SNP array. Plant Biotech J 12:787–796
Watanabe N, Fujii Y, Takesada N, Martinek P (2006) Cytological and microsatellite mapping of genes for brittle rachis in a Triticum aestivum-Aegilops tauschii introgression line. Euphytica 151:63–69
Watterson GA (1975) On the number of segregating sites in genetical models without recombination. Theor Popul Biol 7:256–276
Wei DY, Cui YX, He YJ, Xiong Q, Qian LW, Tong CB, Lu GY, Ding YJ, Li JN, Jung C, Qian W (2017) A genome-wide survey with different rapeseed ecotypes uncovers footprints of domestication and breeding. J Exp Bot 68:4791–4801
Weir BS (1996) Genetic data analysis II, 2nd edn. Sinauer, Sunderland
Winfield MO, Allen AM, Burridge AJ, Barker GL, Benbow HR, Wilkinson PA, Coghill J (2015) High-density SNP genotyping array for hexaploid wheat and its secondary and tertiary gene pool. Plant Biotechnol J 14:1195–1206
Winfield MO, Allen AM, Wilkinson PA, Burridge AJ, Barker GL, Coghill J, Waterfall C, Wingen LU, Griffiths S, Edwards KJ (2017) High density genotyping of the AE Watkins Collection of hexaploid landraces identifies a large molecular diversity compared to elite bread wheat. Plant Biotechnol J. https://doi.org/10.1111/pbi.12757
Wright SI, Bi IV, Schroeder SG, Yamasaki M, Doebley JF, McMullen MD, Brandon S, Gaut BS (2005) The effects of artificial selection on the maize genome. Science 308:1310–1314
Yamasaki M, Wright SI, McMullen MD (2007) Genomic screening for artificial selection during domestication and improvement in maize. Ann Bot-Lond 100:967–973
Yan Y, Zheng J, Xiao Y, Yu J, Hu Y, Cai M, Li Y, Hsam SLK, Zeller FJ (2004) Identification and molecular characterization of a novel y-type Glu-Dt1 glutenin gene of Aegilops tauschii. Theor Appl Genet 108:1349–1358
Yao H, Gray AD, Auger DL, Birchler JA (2013) Genomic dosage effects on heterosis in triploid maize. Proc Natl Acad Sci USA 110:2665–2669
Yu J, Buckler ES (2006) Genetic association mapping and genome organization of maize. Curr Opin Biotechnol 17:155–160
Zhou Y, Chen, ZX, Cheng MP, Chen J, Zhu TT, Wang R, Liu YX, Qi PF, Chen GY, Jiang QT, Wei YM, Luo MC, Nevo E, Allaby RG, Liu DC, Wang JR, Dvorak J, Zheng YL (2018) Uncovering the dispersion history, adaptive evolution and selection of wheat in China. Plant Biotechnol J 16:280–291
Acknowledgements
We thank Prof. R. A. McIntosh, at Plant Breeding Institute, University of Sydney, for reviewing this manuscript. We are grateful to Dr. Huihui Li, at CAAS, for the help in statistical analysis. We acknowledge the Triticeae Toolbox (http://triticeaetoolbox.org; T3) for 90K data availability used in this manuscript. This work was funded by the National Natural Science Foundation of China (31461143021, 31550110212), the National Key Research and Development Program of China (2016YFD0101802, 2016YFE0108600, 2014BAD01B05), and CAAS Science and Technology Innovation Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
We declare no conflicts of interest in regard to this manuscript.
Ethical standards
We declare that these experiments comply with the ethical standards in China.
Additional information
Communicated by Gary Muehlbauer.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, J., Rasheed, A., He, Z. et al. Genome-wide variation patterns between landraces and cultivars uncover divergent selection during modern wheat breeding. Theor Appl Genet 132, 2509–2523 (2019). https://doi.org/10.1007/s00122-019-03367-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-019-03367-4