Abstract
Key message
Phosphorus deficiency in soil is a worldwide constraint threatening maize production. Through a genome-wide association study, we identified molecular markers and associated candidate genes and molecular pathways for low-phosphorus stress tolerance.
Abstract
Phosphorus deficiency in soils will severely affect maize (Zea mays L.) growth and development, thus decreasing the final yield. Deciphering the genetic basis of yield-related traits can benefit our understanding of maize tolerance to low-phosphorus stress. However, considering that yield-related traits should be evaluated under field condition with large populations rather than under hydroponic condition at a single-plant level, searching for appropriate field experimental sites and target traits for low-phosphorus stress tolerance is still very challenging. In this study, a genome-wide association analysis using two natural populations was performed to detect candidate genes in response to low-phosphorus stress at two experimental sites representative of different climate and soil types. In total, 259 candidate genes were identified and these candidate genes are mainly involved in four major pathways: transcriptional regulation, reactive oxygen scavenging, hormone regulation, and remodeling of cell wall. Among these candidate genes, 98 showed differential expression by transcriptome data. Based on a haplotype analysis of grain number under phosphorus deficiency condition, the positive haplotypes with favorable alleles across five loci increased grain number by 42% than those without favorable alleles. For further verifying the feasibility of genomic selection for improving maize low-phosphorus tolerance, we also validated the predictive ability of five genomic selection methods and suggested that moderate-density SNPs were sufficient to make accurate predictions for low-phosphorus tolerance traits. All these results will facilitate elucidating genetic basis of maize tolerance to low-phosphorus stress and improving marker-assisted selection efficiency in breeding process.






Similar content being viewed by others
References
Araus JL, Cairns JE (2014) Field high-throughput phenotyping: the new crop breeding frontier. Trends Plant Sci 19(1):52–61. https://doi.org/10.1016/j.tplants.2013.09.008
Bari R, Pant BD, Stitt M, Scheible W-R (2006) PHO2, MicroRNA399, and PHR1 define a phosphate-signaling pathway in plants. Plant Physiol 141(3):988–999. https://doi.org/10.1104/pp.106.079707
Barrett JC, Fry B, Maller J, Daly MJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21(2):263–265. https://doi.org/10.1093/bioinformatics/bth457
Bayuelo-Jiménez JS, Gallardo-Valdéz M, Pérez-Decelis VA, Magdaleno-Armas L, Ochoa I, Lynch JP (2011) Genotypic variation for root traits of maize (Zea mays L.) from the Purhepecha Plateau under contrasting phosphorus availability. Field Crop Res 121(3):350–362. https://doi.org/10.1016/j.fcr.2011.01.001
Bernhardt J, Stich K, Schwarz-Sommer Z, Saedler H, Wienand U (1998) Molecular analysis of a second functional A1 gene (dihydroflavonol 4-reductase) in Zea mays. Plant J 14(4):483–488. https://doi.org/10.1046/j.1365-313X.1998.00142.x
Boeven PHG, Longin CFH, Leiser WL, Kollers S, Ebmeyer E, Würschum T (2016) Genetic architecture of male floral traits required for hybrid wheat breeding. Theor Appl Genet 129(12):1–15. https://doi.org/10.1007/s00122-016-2771-6
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23(19):2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Bustos R, Castrillo G, Linhares F, Puga ML, Rubio V, Pérez-Pérez J, Solano R, Leyva A, Paz-Ares J (2010) A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis. PLoS Genet 6(9):e1001102. https://doi.org/10.1371/journal.pgen.1001102
Calderon-Vazquez C, Ibarra-Laclette E, Caballero-Perez J, Herrera-Estrella L (2008) Transcript profiling of Zea mays roots reveals gene responses to phosphate deficiency at the plant- and species-specific levels. J Exp Bot 59(9):2479–2497. https://doi.org/10.1093/jxb/ern115
Calderón-Vázquez C, Sawers RJH, Herrera-Estrella L (2011) Phosphate deprivation in maize: genetics and genomics. Plant Physiol 156(3):1067–1077. https://doi.org/10.1104/pp.111.174987
Chen Z-H, Nimmo GA, Jenkins GI, Nimmo HG (2007) BHLH32 modulates several biochemical and morphological processes that respond to Pi starvation in Arabidopsis. Biochem J 405:191–198. https://doi.org/10.1042/BJ20070102
Chen Y-F, Li L-Q, Xu Q, Kong Y-H, Wang H, Wu W-H (2009) The WRKY6 transcription factor modulates PHOSPHATE1 expression in response to low Pi stress in Arabidopsis. Plant Cell 21:3554–3566. https://doi.org/10.1105/tpc.108.064980
Chuck GS, Brown PJ, Meeley R, Hake S (2014) Maize SBP-box transcription factors unbranched2 and unbranched3 affect yield traits by regulating the rate of lateral primordia initiation. Proc Natl Acad Sci USA 111(52):18775–18780. https://doi.org/10.1073/pnas.1407401112
Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w 1118; iso-2; iso-3. Fly 6(2):80–92. https://doi.org/10.4161/fly.19695
Dai X, Wang Y, Yang A, Zhang W-H (2012) OsMYB2P-1, an R2R3 MYB transcription factor, is involved in the regulation of phosphate starvation responses and root architecture in rice. Plant Physiol 159:169–183. https://doi.org/10.1104/pp.112.194217
Dai X, Wang Y, Zhang W-H (2016) OsWRKY74, a WRKY transcription factor, modulates tolerance to phosphate starvation in rice. J Exp Bot 67(3):947–960. https://doi.org/10.1093/jxb/erv515
Devaiah BN, Karthikeyan AS, Raghothama KG (2007a) WRKY75 transcription factor is a modulator of phosphate acquisition and root development in Arabidopsis. Plant Physiol 143:1789–1801. https://doi.org/10.1104/pp.106.093971
Devaiah BN, Nagarajan VK, Raghothama KG (2007b) Phosphate homeostasis and root development in Arabidopsis are synchronized by the zinc finger transcription factor ZAT6. Plant Physiol 145:147–159. https://doi.org/10.1104/pp.107.101691
Devaiah BN, Madhuvanthi R, Karthikeyan AS, Raghothama KG (2009) Phosphate starvation responses and gibberellic acid biosynthesis are regulated by the MYB62 transcription factor in Arabidopsis. Mol Plant 2:43–58. https://doi.org/10.1093/mp/ssn081
Du Q, Wang K, Xu C, Zou C, Xie C, Xu Y, Li W-X (2016) Strand-specific RNA-Seq transcriptome analysis of genotypes with and without low-phosphorus tolerance provides novel insights into phosphorus-use efficiency in maize. BMC Plant Biol 16(1):222. https://doi.org/10.1186/s12870-016-0903-4
Duncan DR, Kriz AL, Paiva R, Widholm JM (2003) Globulin-1 gene expression in regenerable Zea mays (maize) callus. Plant Cell Rep 21(7):684–689. https://doi.org/10.1007/s00299-002-0568-3
El-Moneim DA, Contreras R, Silva-Navas J, Gallego FJ, Figueiras AM, Benito C (2015) On the consequences of aluminium stress in rye: repression of two mitochondrial malate dehydrogenase mRNAs. Plant Biol 17(1):123–133. https://doi.org/10.1111/plb.12219
Fujii H, Chiou T-J, Lin S-I, Aung K, Zhu J-K (2005) A miRNA involved in phosphate-starvation response in Arabidopsis. Curr Biol 15(22):2038–2043. https://doi.org/10.1016/j.cub.2005.10.016
Gabriel SB, Schaffner SF, Nguyen H, Moore JM, Roy J, Blumenstiel B, Higgins J, DeFelice M, Lochner A, Faggart M, Liu-Cordero SN, Rotimi C, Adeyemo A, Cooper R, Ward R, Lander ES, Daly MJ, Altshuler D (2002) The structure of haplotype blocks in the human genome. Science 296(5576):2225–2229. https://doi.org/10.1126/science.1069424
Gamuyao R, Chin JH, Pariasca-Tanaka J, Pesaresi P, Catausan S, Dalid C, Slamet-Loedin I, Tecson-Mendoza EM, Wissuwa M, Heuer S (2012) The protein kinase Pstol1 from traditional rice confers tolerance of phosphorus deficiency. Nature 488(7412):535–539. https://doi.org/10.1038/nature11346
Gao W, Lu L, Qiu W, Wang C, Shou H (2017) OsPAP26 encodes a major purple acid phosphatase and regulates phosphate remobilization in rice. Plant Cell Physiol 58(5):885–892. https://doi.org/10.1093/pcp/pcx041
Gilbert N (2009) The disappearing nutrient. Nature 461(7265):716–718. https://doi.org/10.1038/461716a
Gómez LD, Vanacker H, Buchner P, Noctor G, Foyer CH (2004) Intercellular distribution of glutathione synthesis in maize leaves and its response to short-term chilling. Plant Physiol 134(4):1662–1671. https://doi.org/10.1104/pp.103.033027
Gu R, Chen F, Long L, Cai H, Liu Z, Yang J, Wang L, Li H, Li J, Liu W, Mi G, Zhang F, Yuan L (2016) Enhancing phosphorus uptake efficiency through QTL-based selection for root system architecture in maize. J Genet Genomics 43(11):663–672. https://doi.org/10.1016/j.jgg.2016.11.002
Guo M, Ruan W, Li C, Huang F, Zeng M, Liu Y, Yu Y, Ding X, Wu Y, Wu Z, Mao C, Yi K, Wu P, Mo X (2015) Integrative comparison of the role of the PHOSPHATE STARVATION RESPONSE1 subfamily in phosphate signaling and homeostasis in rice. Plant Physiol 168(4):1762–1776. https://doi.org/10.1104/pp.15.00736
Hamburger D, Rezzonico E, Petétot JMC, Somerville C, Poirier Y (2002) Identification and characterization of the Arabidopsis PHO1 gene involved in phosphate loading to the xylem. Plant Cell 14:889–902. https://doi.org/10.1105/tpc.000745
Hufnagel B, De Sousa SM, Assis L, Guimaraes CT, Leiser W, Azevedo GC, Negri B, Larson BG, Shaff JE, Pastina MM, Barros BA, Weltzien E, Frederick H, Rattunde W, Viana JH, Clark RT, Falcão A, Gazaffi R, Garcia AAF, Schaffert RE, Kochian LV, Magalhaes JV (2014) Duplicate and conquer: multiple homologs of PHOSPHORUS-STARVATION TOLERANCE1 enhance phosphorus acquisition and sorghum performance on low-phosphorus soils. Plant Physiol 166(2):659–677. https://doi.org/10.1104/pp.114.243949
Klinge B, Lange T, Werr W (1997) The IBP gene of maize are expressed in non-meristematic, elongating cells of the seedling and in abortive floral organs. Mol Gen Genet 255(3):248–257. https://doi.org/10.1007/s004380050495
Landi P, Giuliani S, Salvi S, Ferri M, Tuberosa R, Sanguineti MC (2010) Characterization of root-yield-1.06, a major constitutive QTL for root and agronomic traits in maize across water regimes. J Exp Bot 61(13):3553–3562. https://doi.org/10.1093/jxb/erq192
Li Z, Gao Q, Liu Y, He C, Zhang X, Zhang J (2011) Overexpression of transcription factor ZmPTF1 improves low phosphate tolerance of maize by regulating carbon metabolism and root growth. Planta 233:1129–1143. https://doi.org/10.1007/s00425-011-1368-1
Li Z, Xu C, Li K, Yan S, Qu X, Zhang J (2012) Phosphate starvation of maize inhibits lateral root formation and alters gene expression in the lateral root primordium zone. BMC Plant Biol 12(1):89. https://doi.org/10.1186/1471-2229-12-89
Li Y, Zhang J, Zhang X, Fan H, Gu M, Qu H, Xu G (2015) Phosphate transporter OsPht1; 8 in rice plays an important role in phosphorus redistribution from source to sink organs and allocation between embryo and endosperm of seeds. Plant Sci 230:23–32. https://doi.org/10.1016/j.plantsci.2014.10.001
Li C, Sun B, Li Y, Liu C, Wu X, Zhang D, Shi Y, Song Y, Buckler ES, Zhang Z, Wang T, Li Y (2016a) Numerous genetic loci identified for drought tolerance in the maize nested association mapping populations. BMC Genom 17(1):894. https://doi.org/10.1186/s12864-016-3170-8
Li Z, Zhang X, Liu X, Zhao Y, Wang B, Zhang J (2016b) miRNA alterations are important mechanism in maize adaptations to low-phosphate environments. Plant Sci 252:103–117. https://doi.org/10.1016/j.plantsci.2016.07.009
Lid SE, Meeley RB, Min Z, Nichols S, Olsen O-A (2004) Knock-out mutants of two members of the AGL2 subfamily of MADS-box genes expressed during maize kernel development. Plant Sci 167(3):575–582. https://doi.org/10.1016/j.plantsci.2004.04.031
Lin W-Y, Lin S-I, Chiou T-J (2009) Molecular regulators of phosphate homeostasis in plants. J Exp Bot 60:1427–1438. https://doi.org/10.1093/jxb/ern303
Lin S-I, Santi C, Jobet E, Lacut E, Kholti NE, Karlowski WM, Verdeil J-L, Breitler JC, Périn C, Ko S-S, Guiderdoni E, Chiou T-J, Echeverria M (2010) Complex regulation of two target genes encoding SPX-MFS proteins by rice miR827 in response to phosphate starvation. Plant Cell Physiol 51(12):2119–2131. https://doi.org/10.1093/pcp/pcq170
Liu X, Huang M, Fan B, Buckler ES, Zhang Z (2016) Iterative usage of fixed and random effect models for powerful and efficient genome-wide association studies. PLoS Genet 12(2):e1005767. https://doi.org/10.1371/journal.pgen.1005767
Lu L, Qiu W, Gao W, Tyerman SD, Shou H, Wang C (2016) OsPAP10c, a novel secreted acid phosphatase in rice, plays an important role in the utilization of external organic phosphorus. Plant, Cell Environ 39:2247–2259. https://doi.org/10.1111/pce.12794
Matias FI, Galli G, Granato ISC, Fritsche-Neto R (2017) Genomic prediction of autogamous and allogamous plants by SNPs and haplotypes. Crop Sci 57:2951–2958. https://doi.org/10.2135/cropsci2017.01.0022
Mcgonigle B, Keeler SJ, Lau SMC, Koeppe MK, O’Keefe DP (2000) A genomics approach to the comprehensive analysis of the glutathione S-transferase gene family in soybean and maize. Plant Physiol 124(3):1105–1120. https://doi.org/10.1104/pp.124.3.1105
National Bureau of Statistics of China (2016) China Statistical Yearbook 2016. China Statistics Press, Beijing
Pariasca-Tanaka J, Chin JH, Dramé KN, Dalid C, Heuer S, Wissuwa M (2014) A novel allele of the P-starvation tolerance gene OsPSTOL1 from African rice (Oryza glaberrima Steud) and its distribution in the genus Oryza. Theor Appl Genet 127(6):1387–1398. https://doi.org/10.1007/s00122-014-2306-y
Patterson HD, Williams ER (1976) A new class of resolvable incomplete block designs. Biometrika 63(1):83–92. https://doi.org/10.2307/2335087
Qin Y, Ye H, Tang N, Xiong L (2009) Systematic identification of X1-homologous genes reveals a family involved in stress responses in rice. Plant Mol Biol 71(4):483–496. https://doi.org/10.1007/s11103-009-9535-5
Ruan W, Guo M, Wu P, Yi K (2017) Phosphate starvation induced OsPHR4 mediates Pi-signaling and homeostasis in rice. Plant Mol Biol 93:327–340. https://doi.org/10.1007/s11103-016-0564-6
Shin H, Shin H-S, Dewbre GR, Harrison MJ (2004) Phosphate transport in Arabidopsis: Pht1;1 and Pht1;4 play a major role in phosphate acquisition from both low- and high-phosphate environments. Plant J 39(4):629–642. https://doi.org/10.1111/j.1365-313X.2004.02161.x
Shin R, Berg RH, Schachtman DP (2005) Reactive oxygen species and root hairs in Arabidopsis root response to nitrogen, phosphorus and potassium deficiency. Plant Cell Physiol 46:1350–1357. https://doi.org/10.1093/pcp/pci145
Si L, Chen J, Huang X, Gong H, Luo J, Hou Q, Zhou T, Lu T, Zhu J, Shangguan Y, Chen E, Gong C, Zhao Q, Jing Y, Zhao Y, Li Y, Cui L, Fan D, Lu Y, Weng Q, Wang Y, Zhan Q, Liu K, Wei X, An K, An G, Han B (2016) OsSPL13 controls grain size in cultivated rice. Nat Genet 48(4):447–457. https://doi.org/10.1038/ng.3518
Sosso D, Luo D, Li Q-B, Sasse J, Yang J, Gendrot G, Suzuki M, Koch KE, McCarty DR, Chourey PS, Rogowsky PM, Ross-Ibarra J, Yang B, Frommer WB (2015) Seed filling in domesticated maize and rice depends on sweet-mediated hexose transport. Nat Genet 47(12):1489–1493. https://doi.org/10.1038/ng.3422
Spindel J, Begum H, Akdemir D, Virk P, Collard B, Redoña E, Atlin G, Jannink J-L, McCouch SR (2015) Genomic selection and association mapping in rice (Oryza sativa): effect of trait genetic architecture, training population composition, marker number and statistical model on accuracy of rice genomic selection in elite, tropical rice breeding lines. PLoS Genet 11(2):e1004982. https://doi.org/10.1371/journal.pgen.1004982
Sun S, Gu M, Cao Y, Huang X, Zhang X, Ai P, Zhao J, Fan X, Xu G (2012) A constitutive expressed phosphate transporter, OsPht1;1, modulates phosphate uptake and translocation in phosphate-replete rice. Plant Physiol 159(4):1571–1581. https://doi.org/10.1104/pp.112.196345
Sun L, Song L, Zhang Y, Zheng Z, Liu D (2016) Arabidopsis PHL2 and PHR1 act redundantly as the key components of the central regulatory system controlling transcriptional responses to phosphate starvation. Plant Physiol 170(1):499–514. https://doi.org/10.1104/pp.15.01336
Syers JK, Johnston AE, Curtin D (2008) Efficiency of soil and fertilizer phosphorus use: Reconciling changing concepts of soil phosphorus behavior with agronomic information. FAO and Fertilizer and Plant Nutrition Bulletin 18. Rome: Food and Agricultural Organization
Trull MC, Guiltinan MJ, Lynch JP, Deikman J (1997) The responses of wild-type and ABA mutant Arabidopsis thaliana plants to phosphorus starvation. Plant, Cell Environ 20:85–92. https://doi.org/10.1046/j.1365-3040.1997.d01-4.x
Tuberosa R, Sanguineti MC, Landi P, Giuliani MM, Salvi S, Conti S (2002) Identification of QTLs for root characteristics in maize grown in hydroponics and analysis of their overlap with QTLs for grain yield in the field at two water regimes. Plant Mol Biol 48(5):697–712. https://doi.org/10.1023/A:1014897607670
Wang S, Yin L, Tanaka H, Tanaka K, Tsujimoto H (2010a) Identification of wheat alien chromosome addition lines for breeding wheat with high phosphorus efficiency. Breeding Sci 60(4):371–379. https://doi.org/10.1270/jsbbs.60.371
Wang X, Yan X, Liao H (2010b) Genetic improvement for phosphorus efficiency in soybean: a radical approach. Ann Bot 106(1):215–222. https://doi.org/10.1093/aob/mcq029
Wang H, Xu Q, Kong Y-H, Chen Y, Duan J-Y, Wu W-H, Chen Y-F (2014) Arabidopsis WRKY45 transcription factor activates PHOSPHATE TRANSPORTER1;1 expression in response to phosphate starvation. Plant Physiol 164(4):2020–2029. https://doi.org/10.1104/pp.113.235077
Wang H, Xu C, Liu X, Guo Z, Xu X, Wang S, Xie C, Li W-X, Zou C, Xu Y (2017) Development of a multiple-hybrid population for genome-wide association studies: theoretical consideration and genetic mapping of flowering traits in maize. Sci Rep 7:40239. https://doi.org/10.1038/srep40239
Xiao Y, Liu H, Wu L, Warburton M, Yan J (2017) Genome-wide association studies in maize: praise and stargaze. Mol Plant 10(3):359–374. https://doi.org/10.1016/j.molp.2016.12.008
Xu Y (2016) Envirotyping for deciphering environmental impacts on crop plants. Theor Appl Genet 129(4):653–673. https://doi.org/10.1007/s00122-016-2691-5
Xu C, Ren Y, Jian Y, Guo Z, Zhang Y, Xie C, Fu J, Wang H, Wang G, Xu Y, Li P, Zou C (2017) Development of a maize 55 K SNP array with improved genome coverage for molecular breeding. Mol Breed 37(3):20. https://doi.org/10.1007/s11032-017-0622-z
Yamaji N, Takemoto Y, Miyaji T, Mitani-Ueno N, Yoshida KT, Ma JF (2016) Reducing phosphorus accumulation in rice grains with an impaired transporter in the node. Nature 541(7635):92–95. https://doi.org/10.1038/nature20610
Yang N, Lu Y, Yang X, Huang J, Zhou Y, Ali F, Wen W, Liu J, Li J, Yan J (2014) Genome wide association studies using a new nonparametric model reveal the genetic architecture of 17 agronomic traits in an enlarged maize association panel. PLoS Genet 10(9):e1004573. https://doi.org/10.1371/journal.pgen.1004573
Ying Y, Yue W, Wang S, Li S, Wang M, Zhao Y, Wang C, Mao C, Whelan J, Shou H (2017) Two h-type thioredoxins interact with the E2 ubiquitin conjugase PHO2 to fine-tune phosphate homeostasis in rice. Plant Physiol 173(1):812–824. https://doi.org/10.1104/pp.16.01639
Yue E, Liu Z, Li C, Li Y, Liu Q, Xu J-H (2017) Overexpression of miR529a confers enhanced resistance to oxidative stress in rice (Oryza sativa L.). Plant Cell Rep 36:1171–1182. https://doi.org/10.1007/s00299-017-2146-8
Zhang H, Uddin MS, Zou C, Xie C, Xu Y, Li W-X (2014a) Meta-analysis and candidate gene mining of low-phosphorus tolerance in maize. J Integr Plant Biol 56(3):262–270. https://doi.org/10.1111/jipb.12168
Zhang L, Li J, Rong T, Gao S, Wu F, Xu J, Li M, Cao M, Wang J, Hu E, Liu Y, Lu Y (2014b) Large-scale screening maize germplasm for low-phosphorus tolerance using multiple selection criteria. Euphytica 197(3):435–446. https://doi.org/10.1007/s10681-014-1079-3
Zhang Z, Liao H, Lucas WJ (2014c) Molecular mechanisms underlying phosphate sensing, signaling, and adaptation in plants. J Integr Plant Biol 56(3):192–220. https://doi.org/10.1111/jipb.12163
Zhang H, Xu R, Xie C, Huang C, Liao H, Xu Y, Wang J, Li W-X (2015a) Large-scale evaluation of maize germplasm for low-phosphorus tolerance. PLoS ONE 10(5):e0124212. https://doi.org/10.1371/journal.pone.0124212
Zhang S-D, Ling L-Z, Zhang Q-F, Xu J-D, Cheng L (2015b) Evolutionary comparison of two combinatorial regulators of SBP-box genes, MiR156 and MiR529, in plants. PLoS ONE 10(4):e0124621. https://doi.org/10.1371/journal.pone.0124621
Zhang X, Pérez-Rodríguez P, Semagn K, Beyene Y, Babu R, López-Cruz MA, Vicente FS, Olsen M, Buckler ES, Jannink J-L, Prasanna BM, Crossa J (2015c) Genomic prediction in biparental tropical maize populations in water-stressed and well-watered environments using low-density and GBS SNPs. Heredity 114(3):291–299. https://doi.org/10.1038/hdy.2014.99
Zhu D, Scandalios JG (1994) Differential accumulation of manganese-superoxide dismutase transcripts in maize in response to abscisic acid and high osmoticum. Plant Physiol 106(1):173–178. https://doi.org/10.1104/pp.106.1.173
Acknowledgments
The research was supported by the National Natural Science Foundation of China–CGIAR International Collaborative Program (31361140364), National Key Research and Development Program of China (2016YFD0101803), Agricultural Science and Technology Innovation Program (ASTIP) of Chinese Academy of Agricultural Sciences (CAAS), Fundamental Research Funds for Central Non-Profit of Institute of Crop Sciences, CAAS (1610092016124), Bill and Melinda Gates Foundation, and CGIAR Research Program MAIZE.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Benjamin Stich.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2018_3108_MOESM1_ESM.docx
Manhattan plots on ten maize chromosomes for six yield-related traits under two P conditions tested in Gansu, 2015. a Significant SNPs under NP (normal-phosphorus) condition. b Significant SNPs under LP (low-phosphorus) condition. EL: ear length; RN: row number; GNPR: grain number per row; GN: grain number; HGW: hundred grain weight; GWPP: grain weight per plant (DOCX 1561 kb)
122_2018_3108_MOESM2_ESM.docx
Manhattan plots on ten maize chromosomes for seven traits tested under two P conditions in Guangdong. a Significant SNPs under normal-phosphorus (NP) condition in Guangdong. b Significant SNPs under low-phosphorus (LP) condition in Guangdong. FEW: fresh ear weight; LN: leaf number; PH: plant height. The numbers after the trait abbreviations indicate the number of days after planting (DOCX 1575 kb)
122_2018_3108_MOESM5_ESM.docx
Gene ontology classification of candidate genes under low-phosphorus condition. Input means the input genes detected by this study; background means the background genes of inbred line B73 (DOCX 303 kb)
122_2018_3108_MOESM6_ESM.docx
Venn diagram summarizing three types of gene expression. “2 Day vs. 8 Day” indicates differential expression between 2 days and 8 days (after initiation of low-phosphorus treatment); “31778 vs. CCM454” indicates differential expression between tolerant line CCM454 and sensitive line 31778; “Normal P vs. Low P” indicates differential expression between low-phosphorus (LP) and normal-phosphorus (NP) treatments (DOCX 161 kb)
122_2018_3108_MOESM7_ESM.docx
Prediction accuracy of five GS models for 11 tested traits under normal-phosphorus (NP) condition in 2014 and 2015 using genome-wide SNPs. a NP condition in 2014. b NP condition in 2015. PH: plant height; EL: ear length; RN: row number; GNPR: grain number per row; GN: grain number; HGW: hundred grain weight; DTT: days to tassel; DTA: days to anthesis; DTS: days to silk; ASI: anthesis-silking interval; GWPP: grain weight per plant (DOCX 655 kb)
122_2018_3108_MOESM8_ESM.docx
Prediction accuracy of five tested traits under normal-phosphorus (NP) and low-phosphorus (LP) conditions in 2014 and 2015 with 100-20,000 SNPs using GBLUP. a NP condition. b LP condition. PH: plant height; EL: ear length; RN: row number; DTT: days to tassel; GWPP: grain weight per plant (DOCX 523 kb)
122_2018_3108_MOESM9_ESM.docx
Prediction accuracy of BayesB and haplotype-based BayesB for 11 tested traits under low-phosphorus (LP) conditions in 2014 and 2015 using genome-wide SNPs. a LP condition in 2014. b LP condition in 2015. PH: plant height; EL: ear length; RN: row number; GNPR: grain number per row; GN: grain number; HGW: hundred grain weight; DTT: days to tassel; DTA: days to anthesis; DTS: days to silk; ASI: anthesis-silking interval; GWPP: grain weight per plant (DOCX 432 kb)
122_2018_3108_MOESM10_ESM.docx
Prediction accuracy of BayesB and haplotype-based BayesB for 11 tested traits under normal phosphorus (NP) in 2014 and 2015 using genome-wide SNPs. a NP condition in 2014. b NP condition in 2015. PH: plant height; EL: ear length; RN: row number; GNPR: grain number per row; GN: grain number; HGW: hundred grain weight; DTT: days to tassel; DTA: days to anthesis; DTS: days to silk; ASI: anthesis-silking interval; GWPP: grain weight per plant (DOCX 444 kb)
122_2018_3108_MOESM11_ESM.docx
Gene ontology classification of candidate genes related to LPTI. Input means the input genes detected by this study; background means the background genes of inbred line B7311 (DOCX 279 kb)
122_2018_3108_MOESM12_ESM.docx
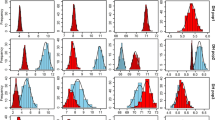
Phenotype distribution of 11 tested traits under normal-phosphorus (NP) and low-phosphorus (LP) conditions (DOCX 628 kb)
122_2018_3108_MOESM13_ESM.xlsx
SLI, LPPI, NPPI, and the extremely tolerant and susceptible inbred lines in 410 inbred lines tested in Gansu (XLSX 35 kb)
Rights and permissions
About this article
Cite this article
Xu, C., Zhang, H., Sun, J. et al. Genome-wide association study dissects yield components associated with low-phosphorus stress tolerance in maize. Theor Appl Genet 131, 1699–1714 (2018). https://doi.org/10.1007/s00122-018-3108-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-018-3108-4