Abstract
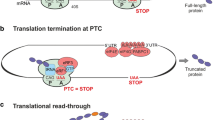
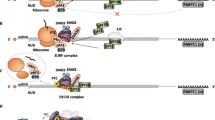
About 11% of all human disease-associated gene lesions are nonsense mutations, resulting in the introduction of an in-frame premature translation-termination codon (PTC) into the protein-coding gene sequence. When translated, PTC-containing mRNAs originate truncated and often dysfunctional proteins that might be non-functional or have gain-of-function or dominant-negative effects. Therapeutic strategies aimed at suppressing PTCs to restore deficient protein function—the so-called nonsense suppression (or PTC readthrough) therapies—have the potential to provide a therapeutic benefit for many patients and in a broad range of genetic disorders, including cancer. These therapeutic approaches comprise the use of translational readthrough-inducing compounds that make the translational machinery recode an in-frame PTC into a sense codon. However, most of the mRNAs carrying a PTC can be rapidly degraded by the surveillance mechanism of nonsense-mediated decay (NMD), thus decreasing the levels of PTC-containing mRNAs in the cell and their availability for PTC readthrough. Accordingly, the use of NMD inhibitors, or readthrough-compound potentiators, may enhance the efficiency of PTC suppression. Here, we review the mechanisms of PTC readthrough and their regulation, as well as the recent advances in the development of novel approaches for PTC suppression, and their role in personalized medicine.



Similar content being viewed by others
References
Mort M, Ivanov D, Cooper DN, Chuzhanova NA (2008) A meta-analysis of nonsense mutations causing human genetic disease. Hum Mutat 29:1037–1047
Savas S, Tuzmen S, Ozcelik H (2006) Human SNPs resulting in premature stop codons and protein truncation. Hum Genomics 2:274–286
Maquat LE, Kinniburgh AJ, Ross J (1981) Unstable β-globin mRNA in β-thalassemia. Cell 27:543–553
Zhang J, Maquat LE (1996) Evidence that the decay of nucleus-associated nonsense mRNA for human triosephosphate isomerase involves nonsense codon recognition after splicing. RNA 2:235–243
Holbrook JA, Neu-Yilik G, Hentze MW, Kulozik AE (2004) Nonsense-mediated decay approaches the clinic. Nat Genet 36:801–808
Popp MW-L, Maquat LE (2013) Organizing principles of mammalian nonsense-mediated mRNA decay. Annu Rev Genet 47:139–165
Frischmeyer PA, Dietz HC (1999) Nonsense-mediated mRNA decay in health and disease. Hum Mol Genet 8:1893–1900
Kugler W, Enssle J, Hentze MW, Kulozik AE (1995) Nuclear degradation of nonsense mutated β-globin mRNA: a post-transcriptional mechanism to protect heterozygotes from severe clinical manifestations of β-thalassemia? Nucleic Acids Res 13:413–418
Ho PJ, Wickramasinghe SN, Rees DC, Lee MJ, EdenThein A (1997) Erythroblastic inclusions in dominantly inherited beta thalassemias. Blood 89:322–328
Keeling KM, Xue X, Gunn G, Bedwell DM (2014) Therapeutics based on stop codon readthrough. Annu Rev Genomics Hum Genet 15:371–394
Keeling KM, Bedwell DM (2011) Suppression of nonsense mutations as a therapeutic approach to treat genetic diseases. Wiley Interdiscip Rev RNA 2:837–852
Lee H-LR, Dougherty JP (2012) Pharmaceutical therapies to recode nonsense mutations in inherited diseases. Pharmacol Ther 136:227–266
Morais P, Adachi H, Yu YT (2020) Suppression of nonsense mutations by new emerging technologies. Int J Mol Sci 21:4394
Hinnebusch AG (2014) The scanning mechanism of eukaryotic translation initiation. Annu Rev Biochem 83:779–812
Jackson RJ, Hellen CUT, Pestova TV (2010) The mechanism of eukaryotic translation initiation. Nat Rev Mol Cell Biol 11:113–127
Lomakin Ivan B, Steitz Thomas A (2013) The initiation of mammalian protein synthesis and the mechanism of scanning. Nature 500:307–311
Shirokikh NE, Preiss T (2018) Translation initiation by cap-dependent ribosome recruitment: recent insights and open questions. Wiley Interdiscip Rev RNA 9:e1473
Dever TE, Green R (2012) The elongation, termination, and recycling phases of translation in eukaryotes. Cold Spring Harb Perspect Biol 4:1–16
Dever TE, Dinman JD, Green R (2018) Translation elongation and recoding in eukaryotes. Cold Spring Harb Perspect Biol 10:a032649
Schuller AP, Green R (2018) Roadblocks and resolutions in eukaryotic translation. Nat Rev Mol Cell Biol 19:526–541
Frolova L, Goff XL, Rasmussen HH et al (1994) A highly conserved eukaryotic protein family possessing properties of polypeptide chain release factor. Nature 372:701–703
Brown A, Shao S, Murray J, Hegde RS, Ramakrishnan V (2015) Structural basis for stop codon recognition in eukaryotes. Nature 524:493–496
Song H, Mugnier P, Das Amit K, Tuite MF, Hemmings BA, Barford D (2000) The crystal structure of human eukaryotic release factor eRF1—mechanism of stop codon recognition and peptidyl-tRNA hydrolysis. Cell 100:311–321
Zhouravleva G, Frolova L, Goff XL, Guellec RL, Inge-vechtomov S, Kisselev L, Philippe M, Zhouravleva G (1995) Termination of translation in eukaryotes is governed by two interacting polypeptide chain release factors, eRF1 and eRF3 Xenopus SUP35C. EMBO J 14:4065–4072
Stansfield I, Jones KM, Kushnirov VV, Dagkesamanskaya AR, Poznyakovski AI, Paushkin SV, Nierras CR, Cox BS, Ter-Avanesyan MD, Tuite MF (1995) The products of the SUP45 (eRF1) and SUP35 genes interact to mediate translation termination in Saccharomyces cerevisiae. EMBO J 14:4365–4373
Hellen CUT (2018) Translation termination and ribosome recycling in eukaryotes. Cold Spring Harb Perspect Biol 10:1–19
Ivanov A, Mikhailova T, Eliseev B, Yeramala L, Sokolova E, Susorov D, Shuvalov A, Schaffitzel C, Alkalaeva E (2016) PABP enhances release factor recruitment and stop codon recognition during translation termination. Nucleic Acids Res 44:7766–7776
Mikhailova T, Shuvalova E, Ivanov A, Susorov D, Shuvalov A, Kolosov PM, Alkalaeva E (2017) RNA helicase DDX19 stabilizes ribosomal elongation and termination complexes. Nucleic Acids Res 45:1307–1318
Beißel C, Neumann B, Uhse S, Hampe I, Karki P, Krebber H (2019) Translation termination depends on the sequential ribosomal entry of eRF1 and eRF3. Nucleic Acids Res 47:4798–4813
Pisarev AV, Skabkin MA, Pisareva VP, Skabkina OV, Rakotondrafara AM, Hentze MW, Hellen CUT, Pestova TV (2010) The role of ABCE1 in eukaryotic posttermination ribosomal recycling. Mol Cell 37:196–210
Annibaldis G, Domanski M, Dreos R, Contu L, Carl S, Kläy N, Mühlemann O (2020) Readthrough of stop codons under limiting ABCE1 concentration involves frameshifting and inhibits nonsense-mediated mRNA decay. Nucleic Acids Res 48:10259–10279
Pisarev AV, Hellen CUTT, Pestova TV (2007) Recycling of eukaryotic posttermination ribosomal complexes. Cell 131:286–299
Drugeon G, Jean-Jean O, Frolova L, Le G, Philippe M, Kisselev L, Haenni AL (1997) Eukaryotic release factor 1 (eRF1) abolishes readthrough and competes with suppressor tRNAs at all three termination codons in messenger RNA. Nucleic Acids Res 25:2254–2258
Bonetti B, Bedwell DM (1995) The efficiency of translation termination is determined by a synergistic interplay between upstream and downstream sequences in Saccharomyces cerevisiae. J Mol Biol 251:334–345
Dabrowski M, Bukowy-Bieryllo Z, Zietkiewicz E (2015) Translational readthrough potential of natural termination codons in eucaryotes—the impact of RNA sequence. RNA Biol 12:950–958
Fearon K, Mcclendon V, Bonetti B, Bedwell DM (1994) Premature translation termination mutations are efficiently suppressed in a highly conserved region of yeast Ste6p, a member of the ATP-binding cassette (ABC) transporter family. J Biol Chem 269:17802–17808
Floquet C, Hatin I, Rousset J-P, Bidou L (2012) Statistical analysis of readthrough levels for nonsense mutations in mammalian cells reveals a major determinant of response to gentamicin. PLoS Genet 8:e1002608
Loughran G, Chou M-Y, Ivanov IP, Jungreis I, Kellis M, Kiran AM, Baranov PV, Atkins JF (2014) Evidence of efficient stop codon readthrough in four mammalian genes. Nucleic Acids Res 42:8928–8938
Namy O, Hatin I, Rousset J-P (2001) Impact of the six nucleotides downstream of the stop codon on translation termination. EMBO Rep 2:787–793
Cridge AG, Crowe-McAuliffe C, Mathew SF, Tate WP (2018) Eukaryotic translational termination efficiency is influenced by the 3′ nucleotides within the ribosomal mRNA channel. Nucleic Acids Res 46:1927–1944
McCaughan KK, Brown CM, Dalphin ME, Berry MJ, Tate WP (1995) Translational termination efficiency in mammals is influenced by the base following the stop codon. Proc Natl Acad Sci USA 92:5431–5435
Hofhuis J, Dieterle S, George R, Schueren F, Thoms S (2017) Dual reporter systems for the analysis of translational readthrough in mammals. Methods Mol Biol 1595:81–92
Peccarelli M, Kebaara BW (2014) Regulation of natural mRNAs by the nonsense-mediated mRNA decay pathway. Eukarytic Cell 13:1126–1135
Wolin SL, Maquat LE (2019) Cellular RNA surveillance in health and disease. Science 366:822–882
Steneberg P, Samakovlis C (2001) A novel stop codon readthrough mechanism produces functional Headcase protein in Drosophila trachea. EMBO Rep 2:593–597
Firth AE, Wills NM, Gesteland RF, Atkins JF (2011) Stimulation of stop codon readthrough: frequent presence of an extended 3’ RNA structural element. Nucleic Acids Res 39:6679–6691
Karijolich J, Yu Y-T (2011) Converting nonsense codons into sense codons by targeted pseudouridylation. Nature 474:395–398
Amrani N, Ganesan R, Kervestin S, Mangus DA, Ghosh S, Jacobson A (2004) A faux 3′-UTR promotes aberrant termination and triggers nonsense-mediated mRNA decay. Nature 432:112–118
Beznosková P, Pavlíková Z, Zeman J, Aitken CE, Valášek LS (2019) Yeast applied readthrough inducing system (YARIS): an invivo assay for the comprehensive study of translational readthrough. Nucleic Acids Res 47:6339–6350
Blanchet S, Rowe M, Haar TVD, Fabret C, Demais S, Howard MJ, Namy O (2015) New insights into stop codon recognition by eRF1. Nucleic Acids Res 43:3298–3308
Roy B, Leszyk JD, Mangus DA, Jacobson A (2015) Nonsense suppression by near-cognate tRNAs employs alternative base pairing at codon positions 1 and 3. Proc Natl Acad Sci USA 112:3038–3043
Loenarz C, Sekirnik R, Armin Thalhammer A et al (2014) Hydroxylation of the eukaryotic ribosomal decoding center affects translational accuracy. Proc Natl Acad Sci USA 111:4019–4024
Bidou L, Allamand V, Rousset J-P, Namy O (2012) Sense from nonsense: therapies for premature stop codon diseases. Trends Mol Med 18:679–688
Bidou L, Hatin I, Perez N, Allamand V, Panthier JJ, Rousset JP (2004) Premature stop codons involved in muscular dystrophies show a broad spectrum of readthrough efficiencies in response to gentamicin treatment. Gene Ther 11:619–627
Jungreis I, Lin MF, Spokony R, Chan CS, Negre N, Victorsen A, White KP, Kellis M (2011) Evidence of abundant stop codon readthrough in Drosophila and other metazoa. Genome Res 21:2096–2113
Manuvakhova M, Keeling K, Bedwell DM (2000) Aminoglycoside antibiotics mediate context-dependent suppression of termination codons in a mammalian translation system. RNA 6:1044–1055
Howard MT, Shirts BH, Petros LM, Flanigan KM, Gesteland RF, Atkins JF (2000) Sequence specificity of aminoglycoside-induced stop condon readthrough: potential implications for treatment of Duchenne muscular dystrophy. Ann Neurol 48:164–116
Li G, Rice CM (1993) The signal for translational readthrough of a UGA codon in Sindbis virus RNA involves a single cytidine residue immediately downstream of the termination codon. J Virol 67:5062–5067
Wangen JR, Green R (2020) Stop codon context influences genome-wide stimulation of termination codon readthrough by aminoglycosides. Chromosom Gene Expr 9:e52611
Harrell L, Melcher U, Atkins JF (2017) Predominance of six different hexanucleotide recoding signals 3′ of read-through stop codons. Nucleic Acids Res 30:2011–2017
Chevance FFV, Guyon SL, Hughes KT (2014) The effects of codon context on in vivo translation speed. PLoS Genet 10:e1004392
Tork S, Hatin I, Rousset JP, Fabret C (2004) The major 5′ determinant in stop codon read-through involves two adjacent adenines. Nucleic Acids Res 32:415–421
Gross T, Siepmann A, Sturm D, Windgassen M, Scarcelli JJ, Seedorf M, Cole CN, Krebber H (2007) The DEAD-box RNA helicase Dbp5 functions in translation termination. Science 315:646–649
Freitag J, Ast J, Bölker M (2012) Cryptic peroxisomal targeting via alternative splicing and stop codon read-through in fungi. Nature 485:522–525
Dunn JG (2013) Ribosome profiling reveals pervasive and regulated stop codon readthrough in Drosophila melanogaster. Cell Biol Evol Biol 2:e01179
Loughran G, Jungreis I, Tzani I, Power M, Dmitriev RI, Ivanov IP, Kellis M, Atkins JF (2018) Stop codon readthrough generates a C-terminally extended variant of the human vitamin D receptor with reduced calcitriol response. J Biol Chem 293:4434–4444
Yamaguchi Y, Hayashi A, Campagnoni CW, Kimura A, Inuzuka T, Baba H (2012) L-MPZ, a novel isoform of myelin P0, is produced by stop codon readthrough. J Biol Chem 287:17765–17776
Eswarappa SM, Potdar AA, Koch WJ, Fan Y, Vasu K, Lindner D, Willard B, Graham LM, DiCorleto PE, Fox PL (2014) Programmed translational readthrough generates antiangiogenic VEGF-Ax. Cell 157:1605–1618
Schueren F, Lingner T, George R, Hofhuis J, Dickel C, Gärtner J, Thoms S (2014) Peroxisomal lactate dehydrogenase is generated by translational readthrough in mammals. Cell Biol Evol Biol 3:e03640
Stiebler AC, Freitag J, Schink KO, Stehlik T, Tillmann BAM, Ast J, Bölker M (2014) Ribosomal readthrough at a short UGA stop codon context triggers dual localization of metabolic enzymes in Fungi and animals. PLoS Genet 10:e1004685
Beznosková P, Wagner S, Jansen ME, Haar TVD, Valášek LS (2015) Translation initiation factor eIF3 promotes programmed stop codon readthrough. Nucleic Acids Res 43:5099–50111
Singh A, Manjunath LE, Kundu P, Sahoo S, Das A, Suma HR, Fox PL, Eswarappa SM (2019) Let-7a-regulated translational readthrough of mammalian AGO1 generates a microRNA pathway inhibitor. EMBO J 38:e100727
Silva AL, Romão L (2009) The mammalian nonsense-mediated mRNA decay pathway: to decay or not to decay! Which players make the decision? FEBS Lett 583:499–505
Rehwinkel J, Raes J, Izaurralde E (2006) Nonsense-mediated mRNA decay: target genes and functional diversification of effectors. Trends Biochem Sci 31:639–646
Mendell JT, Sharifi NA, Meyers JL, Martinez-Murillo F, Dietz HC (2004) Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise. Nat Genet 36:1073–1078
Mendell JT, Dietz HC (2001) When the message goes awry: disease-producing mutations that influence mRNA content and performance. Cell 107:411–414
Nicholson P, Yepiskoposyan H, Metze S, Orozco RZ, Kleinschmidt N, Mühlemann O (2010) Nonsense-mediated mRNA decay in human cells: mechanistic insights, functions beyond quality control and the double-life of NMD factors. Cell Mol Life Sci 67:677–700
Hug N, Longman D, Cáceres JF (2016) Mechanism and regulation of the nonsense-mediated decay pathway. Nucleic Acids Res 44:1483–1495
Mühlemann O, Lykke-Andersen J (2010) How and where are nonsense mRNAs degraded in mammalian cells? RNA Biol 7:28–32
Lee WC, Hou BH, Hou CY, Tsao SM, Kao P, Chen HM (2020) Widespread exon junction complex footprints in the RNA degradome mark mRNA degradation before steady state translation. Plant Cell 32:904–922
Pawlicka K, Kalathiya U, Alfaro J (2020) Nonsense-mediated mRNA decay: Pathologies and the potential for novel therapeutics. Cancers 12:1–17
Behm-Ansmant I, Izaurralde E (2006) Quality control of gene expression: a stepwise assembly pathway for the surveillance complex that triggers nonsense-mediated mRNA decay. Genes Dev 20:391–398
Nasif S, Contu L, Mühlemann O (2018) Beyond quality control: The role of nonsense-mediated mRNA decay (NMD) in regulating gene expression. Semin Cell Dev Biol 75:78–87
Lykke-Andersen S, Jensen TH (2015) Nonsense-mediated mRNA decay: an intricate machinery that shapes transcriptomes. Nat Rev Mol Cell Biol 16:665–677
Ramani AK, Nelson AC, Kapranov P, Bell I, Gingeras TR, Fraser AG (2009) High resolution transcriptome maps for wild-type and nonsense-mediated decay-defective Caenorhabditis elegans. Genome Biol 10:R101
Tani H, Mizutani R, Salam KA, Tano K, Ijiri K, Wakamatsu A, Isogai T, Suzuki Y, Akimitsu N (2012) Genome-wide determination of RNA stability reveals hundreds of short-lived noncoding transcripts in mammals. Genome Res 22:947–956
Tani H, Akimitsu N (2012) Genome-wide technology for determining RNA stability in mammalian cells: historical perspective and recent advantages based on modified nucleotide labeling. RNA Biol 9:1233–1238
Yepiskoposyan H, Aeschimann F, Nilsson D, Okoniewski M, Mühlemann O (2011) Autoregulation of the nonsense-mediated mRNA decay pathway in human cells. RNA 17:2108–2118
Durand S, Cougot N, Mahuteau-Betzer F, Nguyen C-H, Grierson DS, Bertrand E, Tazi J, Lejeune F (2007) Inhibition of nonsense-mediated mRNA decay (NMD) by a new chemical molecule reveals the dynamic of NMD factors in P-bodies. J Cell Biol 178:1145–1160
Oliveira CC, McCarthy JE (1995) The relationship between eukaryotic translation and mRNA stability. A short upstream open reading frame strongly inhibits translational initiation and greatly accelerates mRNA degradation in the yeast Saccharomyces cerevisiae. J Biol Chem 270:8936–8943
Eberle AB, Stalder L, Mathys H, Orozco RZ, Mühlemann O (2008) Posttranscriptional gene regulation by spatial rearrangement of the 3′ untranslated region. PLoS Biol 6:e92
Singh G, Rebbapragada I, Lykke-Andersen J (2008) A competition between stimulators and antagonists of Upf complex recruitment governs human nonsense-mediated mRNA decay. PLoS Biol 6:e111
Kishor A, Fritz SE, Hogg JR (2019) Nonsense-mediated mRNA decay: the challenge of telling right from wrong in a complex transcriptome. Wiley Interdiscip Rev RNA 10:e1548
Fernandes R, Nogueira G, da Costa PJ, Pinto F, Romão L (2019) Nonsense-mediated mRNA decay in development, stress and cancer. Adv Exp Med Biol 1157:41–83
Cutting GR, Kasch LM, Rosenstein BJ, Zielenski J, Tsui LC, Antonarakis SE Jr, Kazazian HH (1990) A cluster of cystic fibrosis mutations in the first nucleotide-binding fold of the cystic fibrosis conductance regulator protein. Nature 346:366–369
Kerem BS, Zielenski J, Markiewicz D, Bozon D, Gazit E, Yahav J, Kennedy D, Riordan JR, Collins FS, Rommens JM (1990) Identification of mutations in regions corresponding to the two putative nucleotide (ATP)-binding folds of the cystic fibrosis gene. Proc Natl Acad Sci USA 87:8447–8451
Veit G, Avramescu RG, Chiang AN et al (2016) From CFTR biology toward combinatorial pharmacotherapy: expanded classification of cystic fibrosis mutations. Mol Biol Cell 27:424–433
Dekkers JF, Berkers G, Kruisselbrink E et al (2016) Characterizing responses to CFTR-modulating drugs using rectal organoids derived from subjects with cystic fibrosis. Sci Transl Med 8:344–384
Borgatti M, Altamura E, Salvatori F, D’Aversa E, Altamura N (2020) Screening readthrough compounds to suppress nonsense mutations: possible application to β-thalassemia. J Clin Med 9:289
Karousis ED, Gurzeler L-A, Annibaldis G, Dreos R, Mühlemann O (2020) Human NMD ensues independently of stable ribosome stalling. Nat Commun 11:1–12
Blanchet S, Cornu D, Hatin I, Grosjean H, Bertin P, Namy O (2018) Deciphering the reading of the genetic code by near-cognate tRNA. Proc Natl Acad Sci USA 115:3018–3023
Burke JF, Mogg AE (1985) Suppression of a nonsense mutation in mammalian cells in vivo by the aminoglycoside antibiotics G-418 and paromomycin. Nucleic Acids Res 13:6265–6272
Nikolaus N, Strehlitz B (2014) DNA-aptamers binding aminoglycoside antibiotics. Sensors 14:3737–3755
Davies J, Gilbert W, Gorini L (1964) Streptomycin, suppression, and the code. Proc Natl Acad Sci USA 51:883–890
Forge A, Schacht J (2000) Aminoglycoside antibiotics. Audiol Neurootol 5:3–22
Krause KM, Serio AW, Kane TR, Connolly LE (2016) Aminoglycosides: an overview. Cold Spring Harb Perspect Med 6:a027029
Obrecht D, Bernardini F, Dale G, Dembowsky K (2011) Emerging new therapeutics against key gram-negative pathogens. Annu Rep Med 46:245–262
Fan-Minogue H, Bedwell DM (2008) Eukaryotic ribosomal RNA determinants of aminoglycoside resistance and their role in translational fidelity. RNA 14:148–157
Moazed D, Noller HF (1987) Interaction of antibiotics with functional sites in 16S ribosomal RNA. Nature 327:389–394
François B, Russell RJM, Murray JB, Aboul-ela F, Masquida B, Vicens Q, Westhof E (2005) Crystal structures of complexes between aminoglycosides and decoding A site oligonucleotides: role of the number of rings and positive charges in the specific binding leading to miscoding. Nucleic Acids Res 33:5677–5699
Recht MI, Douthwaite S, Puglisi JD (1999) Basis for prokaryotic specificity of action of aminoglycoside antibiotics. EMBO J 18:3133–3138
Lynch SR, Puglisi JD (2001) Structural origins of aminoglycoside specificity for prokaryotic ribosomes. J Mol Biol 306:1037–1058
Keeling KM, Bedwell DM (2005) Pharmacological suppression of premature stop mutations that cause genetic diseases. Curr Pharmacogenomics 3:259–269
Pfister P, Hobbie S, Vicens Q, Böttger EC, Westhof E (2011) The molecular basis for A-site mutations conferring aminoglycoside resistance: relationship between ribosomal susceptibility and X-ray crystal structures. ChemBioChem 4:1078–1088
Eustice DC, Wilhelm JM (1984) Fidelity of the eukaryotic codon-anticodon interaction: interference by aminoglycoside antibiotics. Biochemisty 23:1462–1467
Howard MT, Shirts BH, Petros LM, Flanigan KM, Gesteland RF, Atkins JF (2001) Sequence specificity of aminoglycoside-induced stop codon readthrough: potential implications for treatment of Duchenne muscular dystrophy. Ann Neurol 48:164–116
Keeling KM, Wang D, Conard SE, Bedwell DM (2012) Suppression of premature termination codons as a therapeutic approach. Crit Rev Biochem Mol Biol 47:444–463
Keeling KM (2016) Nonsense suppression as an approach to treat lysosomal storage diseases. Diseases 4:32
Martorell L, Cortina V, Parra R, Barquinero J, Vidal F (2020) Variable readthrough responsiveness of nonsense mutations in Hemophilia A. Haematologica 105:508–518
Howard M, Frizzell RA, Bedwell DM (1996) Aminoglycoside antibiotics restore CFTR function by overcoming premature stop mutations. Nat Med 2:467–469
Bedwell DM, Kaenjak A, Benos DJ, Bebok Z, Bubien JK, Hong J, Tousson A, Clancy JP, Sorscher EJ (1997) Suppression of a CFTR premature stop mutation in a bronchial epithelial cell line. Nat Med 3:1280–1284
DrugBank (2005) Geneticin. In: DrugBank. https://www.drugbank.ca/drugs/DB04263. Accessed 15 Aug 2020
Bar-Nun S, Beckmann JS (1983) G-418, an elongation inhibitor of 80 S ribosomes. Biochim Biophys Acta Gene Struct Expr 741:123–127
Kuschal C, Khan SG, Enk B, DiGiovanna JJ, Kraemer KH (2015) Readthrough of stop codons by use of aminoglycosides in cells from xeroderma pigmentosum group C patients. Exp Dermatol 24:296–297
Moosajee M, Gregory-Evans K, Ellis CD, Seabra MC, Gregory-Evans CY (2008) Translational bypass of nonsense mutations in zebrafish rep1, pax2.1 and lamb1 highlights a viable therapeutic option for untreatable genetic eye disease. Hum Mol Genet 17:3987–4000
Popescu AC, Sidorova E, Zhang G, Eubanks JH (2010) Aminoglycoside-mediated partial suppression of MECP2 nonsense mutations responsible for Rett syndrome in vitro. J Neurosci Res 88:2316–2324
Lai C-H, Chun HH, Nahas SA, Mitui M, Gamo KM, Du L, Gatti RA (2004) Correction of ATM gene function by aminoglycoside-induced read-through of premature termination codons. Proc Natl Acad Sci USA 101:15676–15681
Nudelman I, Rebibo-Sabbah A, Cherniavsky M, Belakhov V, Hainrichson M, Chen F, Schacht J, Pilch DS, Ben-Yosef T, Baasov T (2009) Development of novel aminoglycoside (NB54) with reduced toxicity and enhanced suppression of disease-causing premature stop mutations. J Med Chem 52:2836–2845
Zilberberg A, Lahav L, Rosin-Arbesfeld R (2010) Restoration of APC gene function in colorectal cancer cells by aminoglycoside- and macrolide-induced read-through of premature termination codons. Gut 59:496–507
Keeling KM, Bedwell DM (2002) Clinically relevant aminoglycosides can suppress disease-associated premature stop mutations in the IDUA and P53 cDNAs in a mammalian translation system. J Mol Med 80:367–376
Floquet C, Deforges J, Rousset J-P, Bidou L (2011) Rescue of non-sense mutated p53 tumor suppressor gene by aminoglycosides. Nucleic Acids Res 39:3350–3362
Du M, Keeling KM, Fan L, Liu X, Kovaçs T, Sorscher E, Bedwell DM (2006) Clinical doses of amikacin provide more effective suppression of the human CFTR-G542X stop mutation than gentamicin in a transgenic CF mouse model. J Mol Med 84:573–582
Wolstencroft EC, Mattis V, Bajer AA, Young PJ, Lorson CL (2005) A non-sequence-specific requirement for SMN protein activity: the role of aminoglycosides in inducing elevated SMN protein levels. Hum Mol Genet 14:1199–1210
Mattis VB, Rai R, Wang J, Chang C-WT, Coady T, Lorson CL (2006) Novel aminoglycosides increase SMN levels in spinal muscular atrophy fibroblasts. Hum Genomics 120:589–601
Heier CR, DiDonato CJ (2009) Translational readthrough by the aminoglycoside geneticin (G418) modulates SMN stability in vitro and improves motor function in SMA mice in vivo. Hum Mol Genet 18:1310–1322
Zingman LV, Park S, Olson TM, Alekseev AE, Terzic A (2007) Aminoglycoside-induced translational read-through in disease: overcoming nonsense mutations by pharmacogenetic therapy. Clin Pharmacol Ther 81:99–103
Bidou L, Bugaud O, Belakhov V, Baasov T, Namyb O (2017) Characterization of new-generation aminoglycoside promoting premature termination codon readthrough in cancer cells. RNA Biol 14:378–388
Laurent G, Carlier MB, Rollman B, Van Hoof F, Tulkens P (1982) Mechanism of aminoglycoside-induced lysosomal phospholipidosis: in vitro and in vivo studies with gentamicin and amikacin. Biochem Pharmacol 31:3861–3870
Priuska EM, Schacht J (1995) Formation of free radicals by gentamicin and iron and evidence for an iron/gentamicin complex. Biochem Pharmacol 50:1749–1752
Jiang M, Karasawa T, Steyger PS (2017) Aminoglycoside-induced cochleotoxicity: a review. Front Cell Neurosci 11:308
Hobbie SN, Akshay S, Kalapala SK, Bruell CM, Shcherbakov D, Böttger EC (2008) Genetic analysis of interactions with eukaryotic rRNA identify the mitoribosome as target in aminoglycoside ototoxicity. Proc Natl Acad Sci USA 105:3244–3249
Huth ME, Ricci AJ, Cheng AG (2011) Mechanisms of aminoglycoside ototoxicity and targets of hair cell protection. Int J Otolaryngol 2011:937861
Mingeot-Leclercq MP, Tulkens PM (1999) Aminoglycosides: nephrotoxicity. Antimicrob Agents Chemother 43:1003–1012
Begg EJ, Barclay ML (1995) Aminoglycosides—50 years on. Br J Clin Pharmacol 39:597–603
Moestrup SK, Cui S, Vorum H, Bregengård C, Bjørn SE, Norris K, Gliemann J, Christensen EI (1995) Evidence that epithelial glycoprotein 330/megalin mediates uptake of polybasic drugs. J Clin Invest 96:1404–1413
Selimoglu E (2007) Aminoglycoside-induced ototoxicity. Curr Pharm Des 13:119–126
Schroeder R, Waldsich C, Wank H (2000) Modulation of RNA function by aminoglycoside antibiotics. EMBO J 19:1–9
Clifford RE, Coleman JKM, Balough BJ, Liu J, Kopke RD, Jackson RL (2011) Low-dose d-methionine and N-acetyl-l-cysteine for protection from permanent noise-induced hearing loss in chinchillas. Otolaryngol Head Neck Surg 145:999–1006
Trüeb RM (2009) Oxidative stress in ageing of hair. Int J Trichol 1:6–14
Thibault N, Grenier L, Simard M, Bergeron MG, Beauchamp D (1994) Attenuation by daptomycin of gentamicin-induced experimental nephrotoxicity. Antimicrob Agents Chemother 38:1027–1035
Sha SH, Schacht J (1999) Salicylate attenuates gentamicin-induced ototoxicity. Lab Invest 79:807–813
Li G, Sha S-H, Zotova E, Arezzo J, Water TVD, Schacht J (2002) Salicylate protects hearing and kidney function from cisplatin toxicity without compromising its oncolytic action. Lab Invest 82:585–596
Yukihara M, Ito K, Tanoue O, Goto K, Matsushita T, Matsumoto Y, Masuda M, Kimura S, Ueoka R (2011) Effective drug delivery system for Duchenne muscular dystrophy using hybrid liposomes including gentamicin along with reduced toxicity. Biol Pharm Bull 34:712–716
Schiffelers R, Storm G, Bakker-Woudenberg I (2001) Liposome-encapsulated aminoglycosides in pre-clinical and clinical studies. J Antimicrob Chemother 48:333–344
Gilbert DN, Wood CA, Kohlhepp SJ, Kohnen PW, Houghton DC, Finkbeiner HC, Lindsley J, Bennett WM (1989) Polyaspartic acid prevents experimental aminoglycoside nephrotoxicity. J Infect Dis 159:945–953
Du M, Keeling KM, Fan L, Liu X, Bedwell DM (2009) Poly-l-aspartic acid enhances and prolongs gentamicin-mediated suppression of the CFTR-G542X mutation in a cystic fibrosis mouse model. J Biol Chem 284:6885–6892
Nudelman I, Rebibo-Sabbah A, Shallom-Shezifi D, Hainrichson M, Stahl I, Ben-Yosef T, Baasov T (2006) Redesign of aminoglycosides for treatment of human genetic diseases caused by premature stop mutations. Bioorg Med Chem Lett 16:6310–6315
Goldmann T, Rebibo-Sabbah A, Overlack N, Nudelman I, Belakhov V, Baasov T, Ben-Yosef T, Wolfrum U, Nagel-Wolfrum K (2010) Beneficial read-through of a USH1C nonsense mutation by designed aminoglycoside NB30 in the retina. Physiol Pharmacol 51:6671–6680
Goldmann T, Overlack N, Möller F, Belakhov V, Wyk MV, Baasov T, Wolfrum U, Nagel-Wolfrum K (2012) A comparative evaluation of NB30, NB54 and PTC124 in translational read-through efficacy for treatment of an USH1C nonsense mutation. EMBO Mol Med 4:1186–1199
Rowe SM, Bedwell DM (2011) Suppression of CFTR premature termination codons and rescue of CFTR protein and function by the synthetic aminoglycoside NB54. J Mol Med 89:1149–1161
Nudelman I, Glikin D, Smolkin B, Hainrichson M, Belakhov V, Baasov T (2010) Repairing faulty genes by aminoglycosides: development of new derivatives of geneticin (G418) with enhanced suppression of diseases-causing nonsense mutations. Bioorgan Med Chem 18:3735–3746
Brendel C, Belakhov V, Werner H, Wegener E, Gärtner J, Nudelman I, Baasov T, Huppke P (2011) Readthrough of nonsense mutations in Rett syndrome: evaluation of novel aminoglycosides and generation of a new mouse model. J Mol Med 89:389–398
Mattis VB, Ebert AD, Fosso MY, Chang C-W, Lorson CL (2009) Delivery of a read-through inducing compound, TC007, lessens the severity of a spinal muscular atrophy animal model. Hum Mol Genet 18:3906–3913
Chang C-WT, Hui Y, Elchert B, Wang J, Li J, Rai R (2004) Pyranmycins, a novel class of aminoglycosides with improved acid stability: the SAR of d-pyranoses on ring III of pyranmycin. Org Lett 4:4603–4606
Mattis VB, Fosso MY, Chang CW, Lorson CL (2009) Subcutaneous administration of TC007 reduces disease severity in an animal model of SMA. BMC Neurosci 10:142
Bezzerri V, Api M, Allegri M, Fabrizzi B, Corey SJ, Cipolli M (2020) Nonsense suppression therapy: new hypothesis for the treatment of inherited bone marrow failure syndromes. Int J Mol Sci 21:4672
Leubitz A, Frydman-Marom A, Sharpe N, Duzer JV, Campbell KCM, Vanhoutte F (2019) Safety, tolerability, and pharmacokinetics of single ascending doses of ELX-02, a potential treatment for genetic disorders caused by nonsense mutations, in healthy volunteers. Clin Pharmacol Drug Dev 8:984–994
Xue X, Mutyam V, Tang L, Rowe SM (2014) Synthetic aminoglycosides efficiently suppress cystic fibrosis transmembrane conductance regulator nonsense mutations and are enhanced by ivacaftor. Am J Respir Cell Mol Biol 50:805–816
Brasell EJ, Chu LL, Akpa MM, Eshkar-Oren I, Alroy I, Corsini R, Gilfix BM, Yamanaka Y, Huertas P, Goodyer P (2019) The novel aminoglycoside, ELX-02, permits CTNSW138X translational read-through and restores lysosomal cystine efflux in cystinosis. PLoS ONE 14:e0223954
Welch EM, Barton ER, Zhuo J et al (2007) PTC124 targets genetic disorders caused by nonsense mutations. Nature 447:87–91
Hirawat S, Welch EM, Elfring GL et al (2007) Safety, tolerability, and pharmacokinetics of PTC124, a nonaminoglycoside nonsense mutation suppressor, following single- and multiple-dose administration to healthy male and female adult volunteers. J Clin Pharmacol 47:430–444
Krall M, Htun S, Slavotinek A (2019) Use of PTC124 for nonsense suppression therapy targeting BMP4 nonsense variants in vitro and the bmp4st72 allele in zebrafish. PLoS ONE 14:e0212121
Ruggiero L, Iodice R, Esposito M, Dubbioso R, Tozza S, Vitale F, Santoro L, Manganelli F (2018) One-year follow up of three Italian patients with Duchenne muscular dystrophy treated with Ataluren: is earlier better? Ther Adv Neurol Disord 11:1756286418809588
Ebrahimi-Fakhari D, Dillmann U, Flotats-Bastardas M, Poryo M, Abdul-Khaliq H, Shamdeen MG, Mischo B, Zemlin M, Meyer S (2018) Off-label use of Ataluren in four non-ambulatory patients with Duchenne muscular dystrophy: effects on cardiac and pulmonary function and muscle strength. Front Pediatr 6:316
Peltz SW, Morsy M, Welch EM, Jacobson A (2013) Ataluren as an agent for therapeutic nonsense suppression. Annu Rev Med 64:407–425
Roy B, Friesen WJ, Tomizawa Y, Jacobson A (2016) Ataluren stimulates ribosomal selection of near-cognate tRNAs to promote nonsense suppression. Proc Natl Acad Sci USA 113:12508–12513
Allamand V, Pascale G (2008) Drug-induced readthrough of premature stop codons leads to the stabilization of laminin alpha2 chain mRNA in CMD myotubes. J Gene Med 10:217–224
Tutone M, Pibiri I, Lentini L, Pace A (2019) Deciphering the nonsense readthrough mechanism of action of Ataluren: an insilico compared study. ACS Med Chem Lett 10:522–527
Auld DS, Thorne N, Maguire WF, Inglese J (2009) Mechanism of PTC124 activity in cell-based luciferase assays of nonsense codon suppression. Proc Natl Acad Sci USA 106:3585–3590
Auld DS, Lovell S, Thorne N, Lea WA, Maloney DJ, Shen M, Rai G, Battaile KP, Thomas CJ, Simeonov A, Hanzlik RP, Inglese J (2010) Molecular basis for the high-affinity binding and stabilization of firefly luciferase by PTC124. Proc Natl Acad Sci USA 107:4878–4883
Bolze F, Mocek S, Zimmermann A, Klingenspor M (2017) Aminoglycosides, but not PTC124 (Ataluren), rescue nonsense mutations in the leptin receptor and in luciferase reporter genes. Sci Rep 7:1020
Torriano S, Erkilic N, Baux D, Cereso N, De Luca V, Meunier I, Moosajee M, Roux AF, Hamel CP, Kalatzis V (2018) The effect of PTC124 on choroideremia fibroblasts and iPSC-derived RPE raises considerations for therapy. Sci Rep 8:1–15
Harmer SC, Mohal JS, Kemp D, Tinker A (2012) Readthrough of long-QT syndrome type 1 nonsense mutations rescues function but alters the biophysical properties of the channel. Biochem J 443:635–642
Kosmidis G, Veerman CC, Casini S, Verkerk AO, Pas SVD, Bellin M, Wilde AAM, Mummery CL, Bezzina CR (2016) SCN5A, Readthrough-promoting drugs gentamicin and PTC124 fail to rescue Nav1.5 function of human-induced pluripotent stem cell-derived cardiomyocytes carrying nonsense mutations in the sodium channel gene. Circ Arrhythmia Electrophysiol 9:e004227
Hamada M, Takeuchi T, Kondo S, Ikeda Y, Naganawa H, Maeda K, Okami Y, Umezawa H (1970) A new antibiotic, negamycin. J Antibiot 23:170–171
Taguchi A, Hamada K, Hayashi Y (2018) Chemotherapeutics overcoming nonsense mutation-associated genetic diseases: medicinal chemistry of negamycin. J Antibiot 71:205–214
Polikanov YS, Szal T, Jiang F, Gupta P, Matsuda R, Shiozuka M, Steitz TA, Vázquez-Laslop N, Mankin AS (2015) Negamycin interferes with decoding and translocation by simultaneous interaction with rRNA and tRNA. Mol Cell 56:541–550
Schroeder SJ, Blaha G, Moore PB (2007) Negamycin binds to the wall of the nascent chain exit tunnel of the 50S ribosomal subunit. Antimicrob Agents Chemother 51:4462–4465
Uehara Y, Hori M, Kondo S, Hamada M, Umezawa H (1976) Structure-activity relationships among negamycin analogs. J Antibiot 29:937–943
Arakawa M, Shiozuka M, Nakayama Y, Hara T, Matsuda R (2003) Negamycin restores dystrophin expression in skeletal and cardiac muscles of mdx mice. J Biochem 134:751–758
Pranke I, Bidou L, Martin N et al (2018) Factors influencing readthrough therapy for frequent cystic fibrosis premature termination codons. ERJ Open Res 4:00080–02017
Floquet C, RoussetJ-P BL (2011) Readthrough of premature termination codons in the adenomatous polyposis coli gene restores its biological activity in human cancer cells. PLoS ONE 6:e24125
Taguchi A, Nishiguchi S, Shiozuka M (2012) Negamycin analogue with readthrough-promoting activity as a potential drug candidate for Duchenne muscular dystrophy. ACS Med Chem Lett 3:2736–2740
Hamada K, Omura N, Taguchi A, Baradaran-Heravi A, Kotake M, Arai M, Takayama K, Taniguchi A, Roberge M, Hayashi Y (2019) New negamycin-based potent readthrough derivative effective against TGA-type nonsense mutations. ACS Med Chem Lett 10:1450–1456
Baltz RH, Seno ET (1988) Genetics of Streptomyces fradiae and tylosin biosynthesis. Annu Rev Microbiol 42:547–574
Garza-Ramos G, Xiong L, Zhong P, Mankin A (2001) Binding site of macrolide antibiotics on the ribosome: new resistance mutation identifies a specific interaction of ketolides with rRNA. J Bacteriol 183:6898–6907
Novotny GW, Jakobsen L, Andersen NM, Poehlsgaard J, Douthwaite S (2004) Ketolide antimicrobial activity persists after disruption of interactions with domain II of 23S rRNA. Antimicrob Agents Chemother 48:3677–3683
Brisson-Noël A, Trieu-Cuot P, Courvalin P (1988) Mechanism of action of spiramycin and other macrolides. J Antimicrob Chemother 22:13–23
Du L, Jung ME, Damoiseaux R, Completo G, Fike F, Ku J-M, Nahas S, Piao C, Hu H, Gatti RA (2013) A new series of small molecular weight compounds induce read through of all three types of nonsense mutations in the ATM gene. Mol Ther 21:1653–1660
Kayali R, Ku J-M, Bertoni C (2012) Read-through compound 13 restores dystrophin expression and improves muscle function in the mdx mouse model for Duchenne muscular dystrophy. Hum Mol Genet 21:4007–4020
Du L, Gatti RA (2009) Non-aminoglycoside compounds induce readthrough of nonsense mutations. J Exp Med 206:2285–2297
Tutone M, Pibiri I, Perriera R, Campofelice A, Culletta G, Melfi R, Pace A, Almerico AM, Lentini L (2020) Pharmacophore-based design of new chemical scaffolds as translational readthrough-inducing drugs (TRIDs). ACS Med Chem Lett 11:747–753
Friesen WJ, Trotta CR, Tomizawa Y, Zhuo J, Welch EM (2017) The nucleoside analog clitocine is a potent and efficacious readthrough agent. RNA 23:567–577
Trzaska C, Amand S, Bailly C, Leroy C, Marchand V (2020) 2,6-Diaminopurine as a highly potent corrector of UGA nonsense mutations. Nat Commun 11:1509
Mutyam V, Du M, Xue X, Keeling KM, White LE, Rowe SM (2016) Discovery of clinically approved agents that promote suppression of cystic fibrosis transmembrane conductance regulator nonsense mutations. Am J Respir Crit Care Med 194:1092–1103
Gonzalez-Hilarion S, Beghyn T, Jia J, Debreuck N, Berte G, Mamchaoui K, Mouly V, Gruenert DC, Déprez B, Lejeune F (2012) Rescue of nonsense mutations by amlexanox in human cells. Orphanet J Rare Dis 7:58
Atanasova VS, Jiang Q, Prisco M, Gruber C, South AP (2017) Amlexanox enhances premature termination codon read-through in COL7A1 and expression of full length type VII collagen: potential therapy for recessive dystrophic epidermolysis bullosa. J Invest Dermatol 137:1842–1849
Banning A, Schiff M, Tikkanen R (2018) Amlexanox provides a potential therapy for nonsense mutations in the lysosomal storage disorder aspartylglucosaminuria. Biochim Biophys Acta Mol Basis Dis 1864:668–675
Tarrasó G, Real-Martinez A, Parés M et al (2020) Absence of p.R50X Pygm read-through in McArdle disease cellular models. Dis Model Mech 13:d043281
Rijal K, Maraia RJ, Arimbasseri AG (2015) A methods review on use of nonsense suppression to study 3′ end formation and other aspects of tRNA biogenesis. Gene 556:35–50
Temple GF, Dozy AM, Roy KL, Kan YW (1982) Construction of a functional human suppressor tRNA gene: an approach to gene therapy for beta-thalassaemia. Nature 296:5857
Sako Y, Usuki F, Suga H (2006) A novel therapeutic approach for genetic diseases by introduction of suppressor tRNA. Nucleic Acids Symp Ser 50:239–240
Kiselev AV, Ostapenko OV, Rogozhkina EV, Kholod NS, Seit-Nebi AS, Baranov AN (2002) Suppression of nonsense mutations in the dystrophin gene by a suppressor tRNA gene. Mol Biol 36:30–33
Lueck JD, Infield DT, Mackey AL, Pope MR, McCray PB, Ahern CA (2016) Engineered tRNA suppression of a CFTR nonsense mutation. In: bioRxiv. https://www.biorxiv.org/content/https://doi.org/10.1101/088690v1. Accessed 24 Aug 2020
Panchal RG, Wang S, McDermott J Jr, Link CJ (1999) Partial functional correction of xeroderma pigmentosum group A cells by suppressor tRNA. Hum Gene Ther 10:2209–2219
Bordeira-Carriço R, Ferreira D, Oliveira C (2014) Rescue of wild-type E-cadherin expression from nonsense-mutated cancer cells by a suppressor-tRNA. Eur J Hum Genet 22:1085–1092
Lueck JD, Yoon JS, Perales-Puchalt A, Mackey AL, Infield DT, Ahern CA (2019) Engineered transfer RNAs for suppression of premature termination codons. Nat Commun 10:822
Hagemeijer MC, Siegwart DJ, Strug LJ, Cebotaru L, Torres MJ, Sofoluwe A, Beekman JM (2018) Translational research to enable personalized treatment of cystic fibrosis. J Cyst Fibros 17:S46–S51
Herring CD, Blattner FR (2004) Global transcriptional effects of a suppressor tRNA and the inactivation of the regulator frmR. J Bacteriol 186:6714–6720
Ganot P, Bortolin ML, Kiss T (1997) Site-specific pseudouridine formation in preribosomal RNA is guided by small nucleolar RNAs. Cell 89:799–809
Kiss T, Jády BE (2010) Box H/ACA small ribonucleoproteins. Mol Cell 37:597–606
De Zoysa MD, Yu YT (2017) Posttranscriptional RNA pseudouridylation. Enzymes 41:151–167
Junhui G, Yi-Tao Y (2014) RNA pseudouridylation: new insights into an old modification. Trends Biochem Sci 38:210–218
De Zoysa MD, Wu G, Katz R, Yu Y-T (2018) Guide-substrate base-pairing requirement for box H/ACA RNA-guided RNA pseudouridylation. RNA 24:1106–1117
Huang C, Wu G, Yu YT (2012) Inducing nonsense suppression by targeted pseudouridylation. Nat Protoc 7:789–800
Adachi H, Yu Y-T (2020) Pseudouridine-mediated stop codon read-through in S. cerevisiae is sequence context-independent. RNA 26:1247–1256
Benne R, Van den Burg J, Brakenhoff JP, Sloof P, Van Boom JH, Tromp MC (1986) Major transcript of the frameshifted coxII gene from trypanosome mitochondria contains four nucleotides that are not encoded in the DNA. Cell 46:819–826
Ohman M, Kallman AM, Bass BL (2000) In vitro analysis of the binding of ADAR2 to the pre-mRNA encoding the GluR-B R/G site. RNA 6:687–697
Kung C-P, Leonard BM Jr, Weber JD (2018) The role of RNA editing in cancer development and metabolic disorders. Front Endocrinol 9:762
Montiel-Gonzalez MF, Quiroz JFD, Rosenthal JJC (2020) Current strategies for site-directed RNA editing using ADARs. Methods 1:16–24
Wettengel J, Reautschnig P, Geisler S, Kahle PJ, Stafforst T (2017) Harnessing human ADAR2 for RNA repair—recoding a PINK1 mutation rescues mitophagy. Nucleic Acids Res 45:2797–2808
Montiel-González MF, Vallecillo-Viejo IC, Rosenthal JJC (2016) An efficient system for selectively altering genetic information within mRNAs. Nucleic Acids Res 44:e157
Vogel P, Moschref M, Li Q, Merkle T, Selvasaravanan KD, Li JB, Stafforst T (2018) Efficient and precise editing of endogenous transcripts with SNAP-tagged ADARs. Nat Med 15:535–538
Basilio C, Wahba AJ, Lengyel P, Speyer JF, Ochoa S (1962) Synthetic polynucleotides and the amino acid code. Proc Natl Acad Sci USA 48:613–616
Bhakta S, Azad TA, Tsukahara T (2018) Genetic code restoration by artificial RNA editing of Ochre stop codon with ADAR1 deaminase. Protein Eng Des Sel 31:471–478
Montiel-Gonzalez MF, Vallecillo-Viejo I, Yudowski GA, Rosenthal JJC (2013) Correction of mutations within the cystic fibrosis transmembrane conductance regulator by site-directed RNA editing. Proc Natl Acad Sci USA 110:18285–18290
Cox DBT (2017) RNA editing with CRISPR-Cas13. Science 358:1019–1027
Zhang C, Konermann S (2018) Structural basis for the RNA-guided ribonuclease activity of CRISPR-Cas13d. Cell 175:212–223
Li H, Yang Y, Hong W, Huang M, Wu M, Zhao X (2020) Applications of genome editing technology in the targeted therapy of human diseases: mechanisms, advances and prospects. Signal Transduct Target Ther 5:1–23
Rudin N, Sugarman E, Haber JE (1989) Genetic and physical analysis of double-strand break repair and recombination in Saccharomyces cerevisiae. Genetics 122:519–534
Capecchi MR (1989) Altering the genome by homologous recombination. Science 244:1288–1292
Urnov FD, Miller JC, Lee Y-L (2005) Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435:646–651
Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, Lahaye T, Nickstadt A, Bonas U (2009) Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326:1509–1512
Carroll D (2014) Genome engineering with targetable nucleases. Annu Rev Biochem 83:409–439
Cong L, Ran AF, Cox D, Lin S, Barretto R, Habib N, Zhang F (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823
Kim S, Kim D, Cho SW, Kim J, Kim J-S (2014) Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res 24:1012–1019
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR-Cas9 for genome engineering. Cell 157:1262–1278
Koonin EV, Makarova KS, Zhang F (2017) Diversity, classification and evolution of CRISPR-Cas systems. Curr Opin Microbiol 37:67–78
Mali P, Esvelt KM, Church GM (2013) Cas9 as a versatile tool for engineering biology. Nat Methods 10:957–963
Jinek M, Chylinski K, Fonfara I et al (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821
Cohen J (2020) A cut above: pair that developed CRISPR earns historic award. Science 370:271–272
Adli M (2018) The CRISPR tool kit for genome editing and beyond. Nat Commun 9:1911
Long C, McAnally JR, Shelton JM, Mireault AA, Bassel-Duby R, Olson EN (2015) Prevention of muscular dystrophy in mice by CRISPR/Cas9–mediated editing of germline DNA. Science 345:1184–1188
Jo DH, Song DW, Cho CS et al (2019) CRISPR-Cas9-mediated therapeutic editing of Rpe65 ameliorates the disease phenotypes in a mouse model of Leber congenital amaurosis. Sci Adv 5:eaax1210
Erwood S, Laselva O, Bily TMI, Brewer RA, Rutherford AH, Bear CE, Ivakine EA (2020) Allele-specific prevention of nonsense-mediated decay in cystic fibrosis using homology-independent genome editing. Mol Ther Methods Clin Dev 17:1118–1128
Zhang F (2015) CRISPR/Cas9: prospects and challenges. Hum Gene Ther 26:409–410
Islam A (2017) Therapeutic suppression of nonsense mutation: an emerging target in multiple diseases and thrombotic disorders. Curr Pharm Des 23:1598–1609
Linde L, Boelz S, Nissim-Rafinia M, Kerem B (2007) Nonsense-mediated mRNA decay affects nonsense transcript levels and governs response of cystic fibrosis patients to gentamicin. J Clin Invest 117:683–692
Turner KA, Choy FYM (2015) Treatment of nonsense mutations using stop codon read-through therapeutics and creation of animal models using CRISPR-Cas9. J Mol Genet Med 9:1000172
Pérez B, Rodríguez-Pombo P, Ugarte M, Desviat LR (2012) Readthrough strategies for therapeutic suppression of nonsense mutations in inherited metabolic disease. Mol Syndr 3:230–236
Usuki F, Yamashita A, Higuchi I, Ohnishi T, Shiraishi T, Osame M, Ohno S (2004) Inhibition of nonsense-mediated mRNA decay rescues the phenotype in Ullrich’s disease. Ann Neurol 55:740–744
Lentini L, Melfi R, Cancemi P, Pibiri I, Leonardo AD (2019) Caffeine boosts Ataluren’s readthrough activity. Heliyon 5:e01963
Usuki F, Yamashita A, Kashima I, Higuchi I, Osame M, Ohno S (2006) Specific inhibition of nonsense-mediated mRNA decay components, SMG-1 or Upf1, rescues the phenotype of Ullrich disease fibroblasts. Mol Ther 14:351–360
Usuki F, Yamashita A, Shiraishi T, Shiga A, Onodera O, Higuchi I, Ohno S (2013) Inhibition of SMG-8, a subunit of SMG-1 kinase, ameliorates nonsense-mediated mRNA decay-exacerbated mutant phenotypes without cytotoxicity. Proc Natl Acad Sci USA 110:15037–15042
Gong Q, Stump MR, Zhou Z (2011) Inhibition of nonsense-mediated mRNA decay by antisense morpholino oligonucleotides restores functional expression of hERG nonsense and frameshift mutations in long QT syndrome. J Mol Cell Cardiol 50:223–229
Vicente-Crespo M, Palacios IM (2012) Nonsense-mediated mRNA decay and development: shoot the messenger to survive? Biochem Soc Trans 38:1500–1505
Keeling KM, Wang D, Dai Y, Murugesan S, Chenna B, Clark J, Bedwell DM (2013) Attenuation of nonsense-mediated mRNA decay enhances in vivo nonsense suppression. PLoS ONE 8:e60478
Baradaran-Heravi A, Balgi AD (2016) Novel small molecules potentiate premature termination codon readthrough by aminoglycosides. Nucleic Acids Res 44:6583–6598
Rabea SM, Baradaran-Heravi A, Balgi AD, Krause A, Farahabadi SH, Roberge M, Grierson DS (2019) 2-Aminothiazole-4-carboxamides enhance readthrough of premature termination codons by aminoglycosides. ACS Med Chem Lett 10:726–731
Teles Siefers AI, Condinho M, Custódio S, Pereira BF, Fernandes R, Gonçalves V, Costa PJ, Lacerda R, Marques AR, Romão L (2018) Genetics of personalized medicine: cancer and rare diseases. Cell Oncol 41:335–341
Barton-Davis ER, Cordier L, Shoturma DI, Leland SE, Sweeney LH (1999) Aminoglycoside antibiotics restore dystrophin function to skeletal muscles of mdx mice. J Clin Invest 104:375–381
Yu H, Liu X, Huang J, Zhang Y, Hu R, Pu J (2013) Comparison of read-through effects of aminoglycosides and PTC124 on rescuing nonsense mutations of HERG gene associated with long QT syndrome. Int J Mol Med 33:729–735
Yang C, Feng J, Sommer SS (2007) A mouse model for nonsense mutation bypass therapy shows a dramatic multiday response to geneticin. Proc Natl Acad Sci USA 104:15394–15399
Du M, Bedwell DM (2002) Aminoglycoside suppression of a premature stop mutation in a Cftr−/− mouse carrying a human CFTR-G542X transgene. J Mol Med 80:595–604
Kuschal C, DiGiovanna JJ, Khan SG, Gatti RA, Kraemer KH (2013) Repair of UV photolesions in xeroderma pigmentosum group C cells induced by translational readthrough of premature termination codons. Proc Natl Acad Sci USA 110:19483–19488
James PD, Raut S, Rivard GE, Poon M-C, Warner M, McKenna S, Leggo J, Lillicrap D (2005) Aminoglycoside suppression of nonsense mutations in severe hemophilia. Blood 106:3043–3048
Kellermayer R, Szigeti R, Keeling KM, Bedekovics T, Bedwell DM (2006) Aminoglycosides as potential pharmacogenetic agents in the treatment of Hailey–Hailey disease. J Invest Dermatol 126:229–231
Wilschanski M, Kerem E (2003) Gentamicin-induced correction of CFTR function in patients with cystic fibrosis and CFTR stop mutations. N Engl J Med 349:1433–1441
Clancy JP, Bebök Z, Ruiz F et al (2001) Evidence that systemic gentamicin suppresses premature stop mutations in patients with cystic fibrosis. Am J Respir Crit Care Med 163:1683–1692
Fazzari M, Frasca A, Bifari F, Landsberger N (2019) Aminoglycoside drugs induce efficient read-through of CDKL5 nonsense mutations, slightly restoring its kinase activity. RNA Biol 16:1414–1423
Lincoln V, Cogan J, Hou Y, Hirsch M, Chen M (2018) Gentamicin induces LAMB3 nonsense mutation readthrough and restores functional laminin 332 in junctional epidermolysis bullosa. Proc Natl Acad Sci USA 115:e6536–e6545
Azimov R, Abuladze N, Sassani P, Newman D, Kao L, Liu W, Orozco N, Ruchala P, Pushkin A, Kurtz I (2008) G418-mediated ribosomal read-through of a nonsense mutation causing autosomal recessive proximal renal tubular acidosis. Am J Physiol Renal Physiol 295:F633–F641
Zhang M, Heldin A, Palomar-Siles M, Öhlin S, Bykov VJN, Wiman KG (2018) Synergistic rescue of nonsense mutant tumor suppressor p53 by combination treatment with aminoglycosides and Mdm2 inhibitors. Front Oncol 7:323
Bukowy-Bieryllo Z, Dabrowski M, Witt M, Zietkiewicz E (2016) Aminoglycoside-stimulated readthrough of premature termination codons in selected genes involved in primary ciliary dyskinesia. RNA Biol 13:1041–1050
Finkel RS (2013) Readthrough strategies for suppression of nonsense mutations in Duchenne/Becker muscular dystrophy: aminoglycosides and Ataluren (PTC124). J Child Neurol 25:1158–1164
Du M, Liu X, Welch EM, Hirawat S, Peltz SW, Bedwell DM (2008) PTC124 is an orally bioavailable compound that promotes suppression of the human CFTR-G542X nonsense allele in a CF mouse model. Proc Natl Acad Sci USA 105:2064–2069
Drake KM, Dunmore BJ, McNelly LN, Morrell NW, Aldred MA (2013) Correction of nonsense BMPR2 and SMAD9 mutations by ataluren in pulmonary arterial hypertension. Am J Respir Cell Mol Biol 49:403–409
Long L, Yang X, Southwood M, Moore S, Crosby A, Upton PD, Dunmore BJ, Morrell NW (2020) Targeting translational read-through of premature termination mutations in BMPR2 with PTC124 for pulmonary arterial hypertension. Pulm Circ 10:1–14
Liu X, Zhang Y, Zhang B, Gao H, Qiu C (2020) Nonsense suppression induced readthrough of a novel PAX6 mutation in patient-derived cells of congenital aniridia. Mol Genet Genomic Med 8:e1198
Bedwell D, Wang D, Welch E, Keeling KM (2015) The nonsense suppression drug PTC124 restored alpha-l-iduronidase activity and reduces glycosaminoglycan accumulation in MPS IH mice carrying the Idua-W402X mutation. Mol Genet Metab 114:S20
Kerem E, Hirawat S, Armoni S, Yaakov Y, Shoseyov D, Wilschanski M (2008) Effectiveness of PTC124 treatment of cystic fibrosis caused by nonsense mutations: a prospective phase II trial. Lancet 372:719–727
Konstan MW, VanDevanter DR, Rowe SM et al (2020) Efficacy and safety of ataluren in patients with nonsense-mutation cystic fibrosis not receiving chronic inhaled aminoglycosides: the international, randomized, double-blind, placebo-controlled Ataluren Confirmatory Trial in cystic fibrosis (ACT CF). J Cyst Fibros 19:595–601
Kerem E, Konstan MW, De Boeck K, Accurso FJ, Sermet-Gaudelus I, Wilschanski M, Elborn JS, Melotti P, Bronsveld I, Fajac I, Malfroot A, Rosenbluth DB, Walker PA, McColley SA, Knoop C, Quattrucci S, Rietschel E, Zeitlin PL, Barth J, Elfring GL, Welch EM, Branstrom A, Spiegel RJ, Peltz SW, Ajayi T, Rowe SM, Cystic Fibrosis Ataluren Study Group (2014) Ataluren for the treatment of nonsense-mutation cystic fibrosis: a randomised, double-blind, placebo-controlled phase 3 trial. Lancet Respir Med 2:539–547
Wilschanski M, Miller LL, Shoseyov D et al (2011) Chronic ataluren (PTC124) treatment of nonsense mutation cystic fibrosis. Eur Respir J 38:59–69
Finkel RS, Flanigan KM, Wong B et al (2013) Phase 2a study of Ataluren-mediated dystrophin production in patients with nonsense mutation Duchenne muscular dystrophy. PLoS ONE 8:e81302
Bushby K (2014) Ataluren treatment of patients with nonsense mutation dystrophinopathy. Muscle Nerve 50:477–487
McDonald CM, Campbell C, Torricelli RE, Finkel RS, Flanigan KM, Goemans N, Mercuri E (2017) Ataluren in patients with nonsense mutation Duchenne muscular dystrophy (ACT DMD): a multicentre, randomised, double-blind, placebo-controlled, phase 3 trial. Lancet 390:1489–1498
Caspi M, Firsow A, Rajkumar R, Skalka N, Moshkovitz I, Munitz A, Pasmanik-Chor M, Greif H, Megido D, Kariv R, Rosenberg DW, Rosin-Arbesfeld R (2016) A flow cytometry-based reporter assay identifies macrolide antibiotics as nonsense mutation read-through agents. J Mol Med 94:469–482
Acknowledgements
We thank Luka Clarke for critical reading and English editing of the manuscript.
Funding
This work was partially supported by UID/MULTI/04046/2019 Research Unit Grant (to BioISI) and by PTFC/BIM-MEC/3749/2014 research Grant (to LR) from Fundação para a Ciência e a Tecnologia, Portugal.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Martins-Dias, P., Romão, L. Nonsense suppression therapies in human genetic diseases. Cell. Mol. Life Sci. 78, 4677–4701 (2021). https://doi.org/10.1007/s00018-021-03809-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-021-03809-7