Abstract
This study aimed to overexpress an inulinase gene in Kluyveromyces marxianus to achieve the inulinase overproduction and preparation of ultra-high-fructose syrup. First, the inulinase gene (INU1 gene) was overexpressed through codon optimization and selection of a suitable promoter. Then, the inulinase was overproduced by high-cell-density fermentation. Finally, ultra-high-fructose syrup was prepared. It was found that optimization of the codons of the native INU1 gene encoding inulinase from Kluyveromyces marxianus KM-0 made a recombinant strain KM-N70 carrying the optimized INU1N gene produce 251.4 U/mL of the inulinase activity. Furthermore, inulinase activity produced by a recombinant KM-KN16 strain carrying the optimized INU1N gene directed by the native TPS1 promoter from K. marxianus KM-0 reached 338.5 U/mL and expression level of the optimized INU1N gene in the recombinant KM-KN16 strain was also greatly enhanced. During a 10-L fermentation, the recombinant KM-KN16 strain could produce 374.3 U/mL of inulinase activity within 24 h, while during a high-cell-density fed-batch fermentation, the recombinant KM-KN16 strain could produce 896.1 U/mL of inulinase activity and OD600nm value of its culture reached 108. The crude inulinase preparation obtained in this study had an inulinase activity of 18,699.8 ± 736.4 U/g of the crude preparation. It was found that 90.3% of 332.4 g/L of inulin was hydrolyzed to produce 41.0 g/L of glucose and 256.0 g/L of fructose and 91.1% of 328.2 g/L of inulin in the extract of the tubers of Jerusalem artichoke was hydrolyzed to produce 48.3 g/L of glucose and 250.5 g/L of fructose by the crude inulinase preparation (75 U/g of the substrate) within 8 h. The hydrolysates contained major monosaccharides and a trace amount of trisaccharides and the monosaccharides were composed of around 85% fructose and 15% glucose. So far, any other yeasts available have produced only up to 120 U/mL of inulinase activity. Together, this made the recombinant KM-KN16 strain be the best inulinase producer at this moment. The inulinase activity of 18,699.8 ± 736.4 U/g of the crude preparation and the ultra-high-fructose syrup with 41.0 g/L of glucose and 256.0 g/L of fructose were obtained. The inulinase activity obtained in this study was the highest among all the inulinase activities produced by yeast, fungal, and bacterial strains obtained so far.







Similar content being viewed by others
References
Angov E (2011) Codon usage: nature’s roadmap to expression and folding of proteins. Biotechnol J 6:650–659
Chi ZM, Wang F, Chi Z, Yue LX, Liu GL, Zhang T (2009b) Bioproducts from Aureobasidium pullulans, a biotechnologically important yeast. Appl Microbiol Biotechnol 82:793–804
Chi ZM, Chi Z, Liu GL, Wang F, Ju L, Zhang T (2009a) Saccharomycopsis fibuligera and its applications in biotechnology. Biotechnol Adv 27:423–431
Chi ZM, Zhang T, Cao TS, Liu XY, Cui W, Zhao CH (2011) Biotechnological potential of inulin for bioprocesses. Bioresour Technol 102:4295–4303
Cui W, Wang Q, Zhang F, Zhang SC, Chi ZM, Madzak C (2011) Direct conversion of inulin into single cell protein by the engineered Yarrowia lipolytica carrying inulinase gene. Process Biochem 46:1442–1448
Gong F, Sheng J, Chi ZM (2007) Inulinase production by a marine yeast Pichia guilliermondii and inulin hydrolysis by the crude inulinase. J Ind Microbiol Biotechnol 34:179–185
He M, Wu D, Wu J, Chen J (2014) Enhanced expression of endoinulinase from Aspergillus niger by codon optimization in Pichia pastoris and its application in inulooligosaccharide production. J Ind Microbiol Biotechnol 41:105–114
Hensing M, Vrouwenvelder H, Hellinga C, Baartmans R, van Dijken H (1994) Production of the extracellular inulinase in high-cell density fed-batch cultivation of the yeast Kluyveromyces marxianus CBS 6556. Appl Microbiol Biotechnol 42:516–521
Kudl G, Murray AW, Tollervey D, Plotkin JB (2009) Coding-sequence determinants of gene expression in Escherichia coli. Science 324:255–258
Li Q, Zhao XQ, Chang AK, Zhang QM, Bai FW (2012) Ethanol-induced yeast flocculation directed by the promoter of TPS1 encoding trehalose-6-phosphatesynthase1 for efficient ethanol production. Metab Eng 14:1–8
Lim SH, Lee HW, Sok DE, Choi ES (2010) Recombinant production of an inulinase in a Saccharomyces cerevisiae gal80 strain. J Microbiol Biotechnol 20:1529–1533
Liu GL, Wang DS, Wang LF, Zhao SF, Chi ZM (2011) Mig1 is involved in mycelial formation and expression of the genes encoding extracellular enzymes in Saccharomycopsis fibuligera A11. Fungal Genet Biol 48:904–913
Liu GL, Chi Z, Chi ZM (2013) Molecular characterization and expression of microbial inulinase genes. Crit Rev Microbiol 39(2):152–165
Liu GL, Fu GY, Chi Z, Chi ZM (2014) Enhanced expression of the codon-optimized exo-inulinase gene from the yeast Meyerozyma guilliermondii in Saccharomyces sp. W0 and bioethanol production from inulin. Appl Microbiol Biotechnol 98:9129–9138
Liu X, Wu D, Wu J, Chen J (2012) Optimization of the production of Aspergillus niger α-glucosidase expressed in Pichia pastoris. World J Microbiol Biotechnol 29:533–540
Ma ZC, Liu NN, Chi Z, Liu GL, Chi ZM (2015) Genetic modification of the marine-isolated yeast Aureobasidium melanogenum P16 for efficient pullulan production from inulin. Mar Biotechnol 17:511–522
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual.2nd ed. Beijing: Cold Spring Harbor Laboratory Press, pp: 367–70 [Chinesetranslating ed.]
Saunders R, Deane CM (2010) Synonymous codon usage influences the local protein structure observed. Nucleic Acids Res 38:6719–6728
Singh PK, Joseph J, Goyal S, Grover A, Shukla P (2016) Functional analysis of the binding model of microbial inulinases using docking and molecular dynamics simulation. J Mol Model 22:69–76
Singh PK, Shukla P (2012) Molecular modeling and docking of microbial inulinases towards perceptive enzyme-substrate interactions. Indian J Microbiol 52(3):373–380
Singh PK, Kumar V, Yadav R, Shukla P (2017) Bioengineering for microbial inulinases: trends and applications. Curr Protein Pept Sci 18:966–972
Singh RS, Dhaliwal R, Puri M (2006) Production of inulinase from Kluyveromyces marxianus YS-1 using root extract of Asparagus racemosus. Process Biochem 41:1703–1707
Spiro RG (1966) Analysis of sugars found in glycoproteins. Methods Enzymol 8:3–26
Wang JM, Zhang T, Chi Z, Liu GL, Chi ZM (2011) 18S rDNAintegration of the exo-inulinase gene into chromosomes of the high ethanol producing yeast Saccharomyces sp. W0 for direct conversion of inulin to bioethanol. Biom Bioeng 35:3032–3039
Wang LF, Wang ZP, Liu XY, Chi ZM (2013) Citric acid production from extract of Jerusalem artichoke tubers by the genetically engineered yeast Yarrowia lipolytica strain 30 and purification of citric acid. Bioprocess Biosyst Eng 36:1759–1766
Wang ZP, Fu WJ, Xu HM, Chi ZM (2014) Direct conversion of inulin into cell lipid by an inulinase-producing yeast Rhodosporidium toruloides 2F5. Bioresour Technol 161:131–136
Wei W, Zheng Z, Liu Y, Zhu X (1998) Optimizing the culture conditions for higher inulinase production by Kluyveromyces sp. Y-85 and scaling-up fermentation. J Ferment Bioeng 86:395–399
Yuan B, Wang SA, Li FL (2013a) Expression of exo-inulinase genes in Saccharomyces cerevisiae to improve ethanol production from inulin sources. Biotechnol Lett 35:1589–1592
Yuan WJ, Zhao X, Chen L, Bai F (2013b) Improved ethanol production in Jerusalem artichoke tubers by overexpression of inulinase gene in Kluyveromyces marxianus. Biotechnol Bioprocess Eng 18:721–727
Zhang T, Gong F, Peng Y, Chi ZM (2009) Optimization for high-level expression of the Pichia guilliermondii recombinant inulinase in Pichia pastoris and characterization of the recombinant inulinase. Process Biochem 44:1335–1339
Zhang T, Chi Z, Chi ZM, Parrou JL, Gong F (2010) Expression of the inulinase gene from the marine-derived Pichia guilliermondii in Saccharomyces sp. W0 and ethanol production from inulin. Microb Biotechnol 3:576–582
Zhang LL, Tan MJ, Liu GL, Chi Z, Wang GY, Chi ZM (2015) Cloning and characterization of an inulinase gene from the marine yeast Candida membranifaciens subsp. flavinogenie W14-3 and its expression in Saccharomyces sp. W0 for ethanol production. Mol Biotechnol 57:337–347
Zhang S, Yang F, Wang Q, Hua Y, Zhao ZK (2012) High-level secretory expression and characterization of the recombinant Kluyveromyces marxianus inulinase. Process Biochem 47:151–155
Zhang F, Wang ZP, Chi Z, Raoufi Z, Abdollahi S, Chi ZM (2013) The changes in Tps1 activity, trehalose content and expression of TPS1 gene in the psychrotolerant yeast Guehomyces pullulans 17-1 grown at different temperatures. Extremophiles 17:241–249
Zhou HX, Xin FH, Chi Z, Liu GL, Chi ZM (2014) Inulinase production by the yeast Kluyveromyces marxianus with the disrupted MIG1 gene and the over-expressed inulinase gene. Process Biochem 49:1867–1874
Funding
This research was supported by the National Natural Science Foundation of China and the Grant No. is 31500029.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary File 1
The primers used in this study (DOC 46 kb)
Supplementary File 2
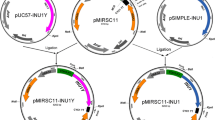
The contruction of the expression plasmid carrying the optimized INU1 gene (DOC 731 kb)
Supplementary File 3
Comparison of the native INU1 gene from K. marxianus KM-0 and the optimized INU1N gene obtained in this study. (DOC 1622 kb)
Supplementary File 4.
Thin-layer chromatogram of the hydrolysis products of inulin and the extract of the tubers of of Jerusalem artichoke by the crude inulinase preparation. Lane 1: glucose; Lane 2, Fructose; Lane 3, Sucrose; Lane 4, Raffinose; Lane 5, Inulin solution; Lane 6, The hydrolysate of inulin; Lane 7, The hysrolysate of the extract of the tubers of Jerusalem artichoke; Lane 8, The decoloured hydrolysate of the extract of the tubers of Jerusalem artichoke. (DOC 256 kb)
Rights and permissions
About this article
Cite this article
Zhang, Y., Li, YF., Chi, Z. et al. Inulinase hyperproduction by Kluyveromyces marxianus through codon optimization, selection of the promoter, and high-cell-density fermentation for efficient inulin hydrolysis. Ann Microbiol 69, 647–657 (2019). https://doi.org/10.1007/s13213-019-01457-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-019-01457-8