Abstract
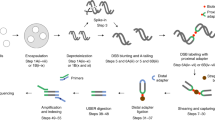
Cells have evolved DNA repair mechanisms to maintain their genetic information unaltered and a DNA damage response pathway that coordinates DNA repair with several cellular events. Despite a clear role for DNA damage in the form of DNA double-strand breaks (DSBs) in several cellular processes, the most commonly used methods to detect DNA lesions are indirect, and rely on antibody-based recognition of DNA damage-associated factors, leaving several important questions unanswered. Differently, here we describe DNA damage In situ ligation followed by Proximity Ligation Assay (DI-PLA), that allows sensitive detection of physical DSBs in fixed cells, through direct labeling of the DSBs with biotinylated oligonucleotides, and subsequent signal amplification by PLA between biotin and a partner protein in the proximity of the DNA break.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Change history
18 April 2019
Work in Fabrizio d’Adda di Fagagna’s laboratory is supported by the Associazione Italiana per la Ricerca sul Cancro, AIRC (application 12971), Cariplo Foundation (grant 2010.0818 and 2014-0812), Fondazione Telethon (GGP12059 and GGP17111), Association for International Cancer Research (AICR-Worldwide Cancer Research Rif. N. 14-1331).
References
Jackson SP, Bartek J (2009) The DNA-damage response in human biology and disease. Nature 461:1071–1078. https://doi.org/10.1038/nature08467
Hoeijmakers JH (2009) DNA damage, aging, and cancer. N Engl J Med 361:1475–1485. https://doi.org/10.1056/NEJMra0804615
Schneider L, Pellegatta S, Favaro R et al (2013) DNA damage in mammalian neural stem cells leads to astrocytic differentiation mediated by BMP2 signaling through JAK-STAT. Stem Cell Reports 1:123–138. https://doi.org/10.1016/j.stemcr.2013.06.004
Mosteiro L, Pantoja C, Alcazar N et al (2016) Tissue damage and senescence provide critical signals for cellular reprogramming in vivo. Science 354(80):aaf4445. https://doi.org/10.1126/science.aaf4445
Vitelli V, Galbiati A, Iannelli F et al (2017) Recent advancements in DNA damage transcription crosstalk and high-resolution mapping of DNA breaks. Annu Rev Genomics Hum Genet 18:87–113
Kim JA, Kruhlak M, Dotiwala F et al (2007) Heterochromatin is refractory to γH2AX modification in yeast and mammals. J Cell Biol 178:209–218. https://doi.org/10.1083/jcb.200612031
Schneider L, Fumagalli M, d’Adda di Fagagna F (2012) Terminally differentiated astrocytes lack DNA damage response signaling and are radioresistant but retain DNA repair proficiency. Cell Death Differ 19:582–591. https://doi.org/10.1038/cdd.2011.129
Soutoglou E, Misteli T (2008) Activation of the cellular DNA damage response in the absence of DNA lesions. Science 320(80):1507–1510. https://doi.org/10.1126/science.1159051.Activation
Shmuel A (1992) Identification of programmed cell death in situ. Cell 119:493–501. https://doi.org/10.1083/jcb.119.3.493
Olive PL, Wlodek D, Banáth JP (1991) DNA double-strand breaks measured in individual cells subjected to gel electrophoresis. Cancer Res 51(17):4671–4676
Olive PL, Banath JP (2006) The comet assay: a method to measure DNA damage in individual cells. Nat Protoc 1:23–29
Söderberg O, Gullberg M, Jarvius M et al (2006) Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat Methods 3:995–1000. https://doi.org/10.1038/nmeth947
Galbiati A, Beausèjour C, d’Adda di Fagagna F (2017) A novel single-cell method provides direct evidence of persistent DNA damage in senescent cells and aged mammalian tissues. Aging Cell 16:422–427. https://doi.org/10.1111/acel.12573
Carpenter AE, Jones TR, Lamprecht MR et al (2006) CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol 7:R100. https://doi.org/10.1186/gb-2006-7-10-r100
Acknowledgments
Work in Fabrizio d’Adda di Fagagna’s laboratory is supported by the Associazione Italiana per la Ricerca sul Cancro, AIRC (application 12971), Cariplo Foundation (grant 2010.0818 and 2014-0812), Fondazione Telethon (GGP12059 and GGP17111), Association for International Cancer Research (AICR-Worldwide Cancer Research Rif. N. 14-1331), the Italian Ministry of Education Universities and Research EPIGEN Project, a European Research Council advanced grant (322726), AriSLA (project “DDRNA and ALS”), AIRC Special Program 5 per mille metastases Project n 21091, AMANDA project Accordo Quadro Regione Lombardia–CNR, and flagship progetto InterOmics.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Galbiati, A., d’Adda di Fagagna, F. (2019). DNA Damage In Situ Ligation Followed by Proximity Ligation Assay (DI-PLA). In: Demaria, M. (eds) Cellular Senescence. Methods in Molecular Biology, vol 1896. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8931-7_2
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8931-7_2
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8930-0
Online ISBN: 978-1-4939-8931-7
eBook Packages: Springer Protocols