Abstract
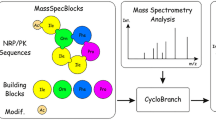
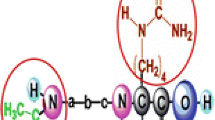
Nonribosomal peptides often possess pronounced bioactivity, and thus, they are often interesting hit compounds in natural product-based drug discovery programs. Their mass spectrometric characterization is difficult due to the predominant occurrence of non-proteinogenic monomers and, especially in the case of cyclic peptides, the complex fragmentation patterns observed. This makes nonribosomal peptide tandem mass spectra annotation challenging and time-consuming. To meet this challenge, software tools for this task have been developed. In this chapter, the workflow for using the software mMass for the annotation of experimentally obtained peptide tandem mass spectra is described. mMass is freely available (http://www.mmass.org), open-source, and the most advanced and user-friendly software tool for this purpose. The software enables the analyst to concisely annotate and interpret tandem mass spectra of linear and cyclic peptides. Thus, it is highly useful for accelerating the structure confirmation and elucidation of cyclic as well as linear peptides and depsipeptides.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Tiburzi F, Visca P, Imperi F (2007) Do nonribosomal peptide synthetases occur in higher eukaryotes? IUBMB Life 59:730–733
Caboche S, Leclère V, Pupin M, Kucherov G et al (2010) Diversity of monomers in nonribosomal peptides: towards the prediction of origin and biological activity. J Bacteriol 192:5143–5150
Tidgewell K, Clark BR, Gerwick WH (2010) The natural products chemistry of cyanobacteria. In: Mander L, Liu H-W (eds) Comprehensive natural products II: chemistry and biology. Elsevier, Oxford, pp 141–188
Niedermeyer T, Brönstrup M (2012) Natural-product drug discovery from microalgae. In: Posten C, Walter C (eds) Microalgal biotechnology: integration and economy. de Gruyter, Berlin, pp 169–200
Ziemert N, Ishida K, Liaimer A, Hertweck C et al (2008) Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angew Chemie Int Ed 47:7756–7759
Velásquez JE, van der Donk WA (2011) Genome mining for ribosomally synthesized natural products. Curr Opin Chem Biol 15:11–21
Marahiel MA (2009) Working outside the protein-synthesis rules: insights into non-ribosomal peptide synthesis. J Pept Sci 15: 799–807
Schwarzer D, Finking R, Marahiel MA (2003) Nonribosomal peptides: from genes to products. Nat Prod Rep 20:275–287
Strieker M, Tanović A, Marahiel MA (2010) Nonribosomal peptide synthetases: structures and dynamics. Curr Opin Struct Biol 20: 234–240
Finking R, Marahiel MA (2004) Biosynthesis of nonribosomal peptides. Annu Rev Microbiol 58:453–488
Tillett D, Dittmann E, Erhard M, von Döhren H et al (2000) Structural organization of microcystin biosynthesis in Microcystis aeruginosa PCC7806: an integrated peptide-polyketide synthetase system. Chem Biol 7: 753–764
Dittmann E, Neilan BA, Börner T (2001) Molecular biology of peptide and polyketide biosynthesis in cyanobacteria. Appl Microbiol Biotechnol 57:467–473
Christiansen G, Fastner J, Erhard M, Börner T et al (2003) Microcystin biosynthesis in planktothrix: genes, evolution, and manipulation. J Bacteriol 185:564–572
Caboche S, Pupin M, Leclère V et al (2008) NORINE: a database of nonribosomal peptides. Nucleic Acids Res 36:326–331
Dreyfuss M, Härri E, Hofmann H et al (1976) Cyclosporin A and C: new metabolites from Trichoderma polysporum. Microbiology 133: 125–133
Von Wartburg A, Traber R (1988) Cyclosporins, fungal metabolites with immunosuppressive activities. Prog Med Chem 25: 1–33
McCormick MH, Stark WM, Pittenger GE et al (1956) Vancomycin, a new antibiotic. I. Chemical and biologic properties. Antibiot Annu 3:606–611
Nagarajan R (1991) Antibacterial activities and modes of action of vancomycin and related glycopeptides. Antimicrob Agents Chemother 35:605–609
Rohr J (2006) Cryptophycin anticancer drugs revisited. ACS Chem Biol 1:747–750
Rusconi F (2009) massXpert 2: a cross-platform software environment for polymer chemistry modelling and simulation/analysis of mass spectrometric data. Bioinformatics 25:2741–2742
Jagannath S, Sabareesh V (2007) Peptide Fragment Ion Analyser (PFIA): a simple and versatile tool for the interpretation of tandem mass spectrometric data and de novo sequencing of peptides. Rapid Commun Mass Spectrom 21:3033–3038
Liu W, Ng J, Meluzzi D, Bandeira N et al (2009) The interpretation of tandem mass spectra obtained from cyclic non-ribosomal peptides. Anal Chem 81:4200–4209
Niedermeyer THJ, Strohalm M (2012) mMass as a software tool for the annotation of cyclic peptide tandem mass spectra. PLoS One 7:e44913
Acknowledgment
The author thanks R. Pozzi, T. Schafhauser and M. Strohalm for critically reading the manuscript and suggesting improvements.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Niedermeyer, T.H.J. (2016). Annotating and Interpreting Linear and Cyclic Peptide Tandem Mass Spectra. In: Evans, B. (eds) Nonribosomal Peptide and Polyketide Biosynthesis. Methods in Molecular Biology, vol 1401. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-3375-4_13
Download citation
DOI: https://doi.org/10.1007/978-1-4939-3375-4_13
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-3373-0
Online ISBN: 978-1-4939-3375-4
eBook Packages: Springer Protocols