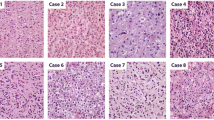
Abstract
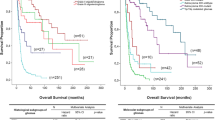
It has become increasingly evident that morphologically similar gliomas may have distinct clinical phenotypes arising from diverse genetic signatures. To date, glial tumours from the Tunisian population have not been investigated. To address this, we correlated the clinico-pathology with molecular data of 110 gliomas by a combination of HM450K array, MLPA and TMA-IHC. PTEN loss and EGFR amplification were distributed in different glioma histological groups. However, 1p19q co-deletion and KIAA1549:BRAF fusion were, respectively, restricted to Oligodendroglioma and Pilocytic Astrocytoma. CDKN2A loss and EGFR overexpression were more common within high-grade gliomas. Furthermore, survival statistical correlations led us to identify Glioblastoma (GB) prognosis subtypes. In fact, significant lower overall survival (OS) was detected within GB that overexpressed EGFR and Cox2. In addition, IDH1R132H mutation seemed to provide a markedly survival advantage. Interestingly, the association of IDHR132H mutation and EGFR normal status, as well as the association of differentiation markers, defined GB subtypes with good prognosis. By contrast, poor survival GB subtypes were defined by the combination of PTEN loss with PDGFRa expression and/or EGFR amplification. Additionally, GB presenting p53-negative staining associated with CDKN2A loss or p21 positivity represented a subtype with short survival. Thus, distinct molecular subtypes with individualised prognosis were identified. Interestingly, we found a unique histone mutation in a poor survival young adult GB case. This tumour exceptionally associated the H3F3A G34R mutation and MYCN amplification as well as 1p36 loss and 10q loss. Furthermore, by exhibiting a remarkable methylation profile, it emphasised the oncogenic power of G34R mutation connecting gliomagenesis and chromatin regulation.





Similar content being viewed by others
References
Ostrom QT, Bauchet L, Davis FG, Deltour I, Fisher JL, Langer CE, Pekmezci M, Schwartzbaum JA et al (2014) The epidemiology of glioma in adults: a “state of the science” review. Neuro-Oncology 16(7):896–913. doi:10.1093/neuonc/nou087
Crocetti E, Trama A, Stiller C, Caldarella A, Soffietti R, Jaal J, Weber DC, Ricardi U et al (2012) Epidemiology of glial and non-glial brain tumours in Europe. Eur J Cancer 48(10):1532–1542. doi:10.1016/j.ejca.2011.12.013
Ohgaki H, Kleihues P (2005) Population-based studies on incidence, survival rates, and genetic alterations in astrocytic and oligodendroglial gliomas. J Neuropathol Exp Neurol 64(6):479–489
Trabelsi S, Brahim DH, Ladib M, Mama N, Harrabi I, Tlili K, Yacoubi MT, Krifa H et al (2014) Glioma epidemiology in the central Tunisian population: 1993-2012. Asian Pac J Cancer Prev 15(20):8753–8757
Ohgaki H, Kleihues P (2005) Epidemiology and etiology of gliomas. Acta Neuropathol 109(1):93–108. doi:10.1007/s00401-005-0991-y
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, Scheithauer BW, Kleihues P (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114(2):97–109. doi:10.1007/s00401-007-0243-4
Figarella-Branger D, Labrousse F, Mohktari K, Societe franc aise de n, Reseau de neuro-oncologie p (2012) Guidelines for adult diffuse gliomas WHO grade II, III and IV: pathology and biology. Societe franc aise de neuropathologie. Reseau de neuro-oncologie pathologique. Ann Pathol 32(5):318–327. doi:10.1016/j.annpat.2012.09.228
Brennan C, Momota H, Hambardzumyan D, Ozawa T, Tandon A, Pedraza A, Holland E (2009) Glioblastoma subclasses can be defined by activity among signal transduction pathways and associated genomic alterations. PLoS One 4(11):e7752. doi:10.1371/journal.pone.0007752
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, Zheng S, Chakravarty D et al (2013) The somatic genomic landscape of glioblastoma. Cell 155(2):462–477. doi:10.1016/j.cell.2013.09.034
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L et al (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17(1):98–110. doi:10.1016/j.ccr.2009.12.020
Yip S, Butterfield YS, Morozova O, Chittaranjan S, Blough MD, An J, Birol I, Chesnelong C et al (2012) Concurrent CIC mutations, IDH mutations, and 1p/19q loss distinguish oligodendrogliomas from other cancers. J Pathol 226(1):7–16. doi:10.1002/path.2995
Zhu H, Acquaviva J, Ramachandran P, Boskovitz A, Woolfenden S, Pfannl R, Bronson RT, Chen JW et al (2009) Oncogenic EGFR signaling cooperates with loss of tumor suppressor gene functions in gliomagenesis. Proc Natl Acad Sci U S A 106(8):2712–2716. doi:10.1073/pnas.0813314106
Talasila KM, Soentgerath A, Euskirchen P, Rosland GV, Wang J, Huszthy PC, Prestegarden L, Skaftnesmo KO et al (2013) EGFR wild-type amplification and activation promote invasion and development of glioblastoma independent of angiogenesis. Acta Neuropathol 125(5):683–698. doi:10.1007/s00401-013-1101-1
Okada Y, Hurwitz EE, Esposito JM, Brower MA, Nutt CL, Louis DN (2003) Selection pressures of TP53 mutation and microenvironmental location influence epidermal growth factor receptor gene amplification in human glioblastomas. Cancer Res 63(2):413–416
Simon M, Koster G, Menon AG, Schramm J (1999) Functional evidence for a role of combined CDKN2A (p16-p14(ARF))/CDKN2B (p15) gene inactivation in malignant gliomas. Acta Neuropathol 98(5):444–452
Kraus JA, Glesmann N, Beck M, Krex D, Klockgether T, Schackert G, Schlegel U (2000) Molecular analysis of the PTEN, TP53 and CDKN2A tumor suppressor genes in long-term survivors of glioblastoma multiforme. J Neuro-Oncol 48(2):89–94
Ichimura K, Bolin MB, Goike HM, Schmidt EE, Moshref A, Collins VP (2000) Deregulation of the p14ARF/MDM2/p53 pathway is a prerequisite for human astrocytic gliomas with G1-S transition control gene abnormalities. Cancer Res 60(2):417–424
Zhao S, Lin Y, Xu W, Jiang W, Zha Z, Wang P, Yu W, Li Z et al (2009) Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1alpha. Science 324(5924):261–265. doi:10.1126/science.1170944
Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA, Yuan W, Kos I, Batinic-Haberle I et al (2009) IDH1 and IDH2 mutations in gliomas. N Engl J Med 360(8):765–773. doi:10.1056/NEJMoa0808710
Weller M, Felsberg J, Hartmann C, Berger H, Steinbach JP, Schramm J, Westphal M, Schackert G et al (2009) Molecular predictors of progression-free and overall survival in patients with newly diagnosed glioblastoma: a prospective translational study of the German Glioma Network. J Clin Oncol : Off J Am Soc Clin Oncol 27(34):5743–5750. doi:10.1200/JCO.2009.23.0805
Weller M, Berger H, Hartmann C, Schramm J, Westphal M, Simon M, Goldbrunner R, Krex D et al (2007) Combined 1p/19q loss in oligodendroglial tumors: predictive or prognostic biomarker? Clin Cancer Res 13(23):6933–6937. doi:10.1158/1078-0432.CCR-07-0573
Smith JS, Perry A, Borell TJ, Lee HK, O’Fallon J, Hosek SM, Kimmel D, Yates A et al (2000) Alterations of chromosome arms 1p and 19q as predictors of survival in oligodendrogliomas, astrocytomas, and mixed oligoastrocytomas. J Clin Oncol : Off J Am Soc Clin Oncol 18(3):636–645
Trabelsi S, Mama N, Ladib M, Popov S, Burford A, Mokni M, Tlili K, Krifa H et al (2015) Adult recurrent pilocytic astrocytoma: clinical, histopathological and molecular study. Neuro-Chirurgie 61(6):392–397. doi:10.1016/j.neuchi.2015.07.002
Sievert AJ, Jackson EM, Gai X, Hakonarson H, Judkins AR, Resnick AC, Sutton LN, Storm PB et al (2009) Duplication of 7q34 in pediatric low-grade astrocytomas detected by high-density single-nucleotide polymorphism-based genotype arrays results in a novel BRAF fusion gene. Brain Pathol 19(3):449–458. doi:10.1111/j.1750-3639.2008.00225.x
Kaul A, Chen YH, Emnett RJ, Dahiya S, Gutmann DH (2012) Pediatric glioma-associated KIAA1549:BRAF expression regulates neuroglial cell growth in a cell type-specific and mTOR-dependent manner. Genes Dev 26(23):2561–2566. doi:10.1101/gad.200907.112
Jones DT, Kocialkowski S, Liu L, Pearson DM, Ichimura K, Collins VP (2009) Oncogenic RAF1 rearrangement and a novel BRAF mutation as alternatives to KIAA1549:BRAF fusion in activating the MAPK pathway in pilocytic astrocytoma. Oncogene 28(20):2119–2123. doi:10.1038/onc.2009.73
Hawkins C, Walker E, Mohamed N, Zhang C, Jacob K, Shirinian M, Alon N, Kahn D et al (2011) BRAF-KIAA1549 fusion predicts better clinical outcome in pediatric low-grade astrocytoma. Clin Cancer Res 17(14):4790–4798. doi:10.1158/1078-0432.CCR-11-0034
Cancer Genome Atlas Research N, Brat DJ, Verhaak RG, Aldape KD, Yung WK, Salama SR, Cooper LA, Rheinbay E et al (2015) Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N Engl J Med 372(26):2481–2498. doi:10.1056/NEJMoa1402121
Macaulay RJ (2015) Impending impact of molecular pathology on classifying adult diffuse gliomas. Cancer Control : J Moffitt Cancer Center 22(2):200–205
Wilhelmsson U, Eliasson C, Bjerkvig R, Pekny M (2003) Loss of GFAP expression in high-grade astrocytomas does not contribute to tumor development or progression. Oncogene 22(22):3407–3411. doi:10.1038/sj.onc.1206372
Lin A, Rodriguez FJ, Karajannis MA, Williams SC, Legault G, Zagzag D, Burger PC, Allen JC et al (2012) BRAF alterations in primary glial and glioneuronal neoplasms of the central nervous system with identification of 2 novel KIAA1549:BRAF fusion variants. J Neuropathol Exp Neurol 71(1):66–72. doi:10.1097/NEN.0b013e31823f2cb0
Jones DT, Kocialkowski S, Liu L, Pearson DM, Backlund LM, Ichimura K, Collins VP (2008) Tandem duplication producing a novel oncogenic BRAF fusion gene defines the majority of pilocytic astrocytomas. Cancer Res 68(21):8673–8677. doi:10.1158/0008-5472.CAN-08-2097
Smith JS, Alderete B, Minn Y, Borell TJ, Perry A, Mohapatra G, Hosek SM, Kimmel D et al (1999) Localization of common deletion regions on 1p and 19q in human gliomas and their association with histological subtype. Oncogene 18(28):4144–4152. doi:10.1038/sj.onc.1202759
Idbaih A, Marie Y, Pierron G, Brennetot C, Hoang-Xuan K, Kujas M, Mokhtari K, Sanson M et al (2005) Two types of chromosome 1p losses with opposite significance in gliomas. Ann Neurol 58(3):483–487. doi:10.1002/ana.20607
Jenkins RB, Blair H, Ballman KV, Giannini C, Arusell RM, Law M, Flynn H, Passe S et al (2006) A t(1;19)(q10;p10) mediates the combined deletions of 1p and 19q and predicts a better prognosis of patients with oligodendroglioma. Cancer Res 66(20):9852–9861. doi:10.1158/0008-5472.CAN-06-1796
Reuss DE, Sahm F, Schrimpf D, Wiestler B, Capper D, Koelsche C, Schweizer L, Korshunov A et al (2015) ATRX and IDH1-R132H immunohistochemistry with subsequent copy number analysis and IDH sequencing as a basis for an “integrated” diagnostic approach for adult astrocytoma, oligodendroglioma and glioblastoma. Acta Neuropathol 129(1):133–146. doi:10.1007/s00401-014-1370-3
Sturm D, Witt H, Hovestadt V, Khuong-Quang DA, Jones DT, Konermann C, Pfaff E, Tonjes M et al (2012) Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22(4):425–437. doi:10.1016/j.ccr.2012.08.024
Huang M, Weiss WA (2013) G34, another connection between MYCN and a pediatric tumor. Cancer Discov 3(5):484–486. doi:10.1158/2159-8290.CD-13-0126
Wagner EJ, Carpenter PB (2012) Understanding the language of Lys36 methylation at histone H3. Nat Rev Mol Cell Biol 13(2):115–126. doi:10.1038/nrm3274
Jones C, Baker SJ (2014) Unique genetic and epigenetic mechanisms driving paediatric diffuse high-grade glioma. Nature reviews Cancer 14(10). doi:10.1038/nrc3811
Cohen A, Sato M, Aldape K, Mason CC, Alfaro-Munoz K, Heathcock L, South ST, Abegglen LM et al (2015) DNA copy number analysis of grade II–III and grade IV gliomas reveals differences in molecular ontogeny including chromothripsis associated with IDH mutation status. Acta Neuropathol Commun 3:34. doi:10.1186/s40478-015-0213-3
Ohgaki H, Kleihues P (2010) Genetic profile of astrocytic and oligodendroglial gliomas. Brain Tumor Pathol 28(3):177–183. doi:10.1007/s10014-011-0029-1
Wu E, Palmer N, Tian Z, Moseman AP, Galdzicki M, Wang X, Berger B, Zhang H et al (2008) Comprehensive dissection of PDGF-PDGFR signaling pathways in PDGFR genetically defined cells. PLoS One 3(11):e3794. doi:10.1371/journal.pone.0003794
Maher EA, Furnari FB, Bachoo RM, Rowitch DH, Louis DN, Cavenee WK, DePinho RA (2001) Malignant glioma: genetics and biology of a grave matter. Genes Dev 15(11):1311–1333. doi:10.1101/gad.891601
McLendon R, Friedman A, Bigner D, Van Meir EG, Brat DJ, Mastrogianakis GM, Olson JJ, Mikkelsen T et al (2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455 (7216):1061-1068. doi:10.1038/nature07385
Acknowledgments
We would like to thank the patients for participating in this study. We would like to acknowledge the efforts of GETUC Tunisian group members ‘groupe d’étude des tumeurs cérébrales’. We also thank all team members in the Divisions of Molecular Pathology and Cancer Therapeutics at The Institute of Cancer Research in Sutton for their precious help in preparing this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
A total of 110 patients with gliomas were collected from departments of Neurosurgery and Histopathology in Sahloul and Farhat Hached University hospitals in Sousse with the ethical committee approval.
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Trabelsi, S., Chabchoub, I., Ksira, I. et al. Molecular Diagnostic and Prognostic Subtyping of Gliomas in Tunisian Population. Mol Neurobiol 54, 2381–2394 (2017). https://doi.org/10.1007/s12035-016-9805-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-016-9805-6