Abstract
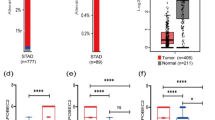
APOBEC3B belongs to the cytidine deaminase apolipoprotein B mRNA-editing enzyme catalytic polypeptide-like (APOBEC3) family of enzymes and induces C to T transitions of target DNA by cytidine deamination. Recently, several mutations in various cancers have been linked to APOBEC3B, suggesting a crucial role for this protein in carcinogenesis and cancer development. However, the significance of APOBEC3B in esophageal squamous cell carcinoma (ESCC) remains uncertain. In addition, the APOBEC3B immunoreactivity in cancer tissues is uncertain. Recently, we have demonstrated that PIK3CA mutation and the methylation level of long interspersed nucleotide element 1 (LINE-1) (a surrogate marker of global DNA methylation level) are prognostic markers and have crucial role on malignancy in ESCC patients. This study aims to clarify the impact of APOBEC3B on the clinical, pathological, and molecular features of ESCC. We evaluated APOBEC3B expression in ESCC and investigated the relationships among the immunoreactivity of APOBEC3B, clinical and pathological features, and the molecular features of ESCC (PIK3CA mutation, p53 expression, and LINE-1 methylation level). The immunoreactivity and mRNA level of APOBEC3B were significantly higher in cancer tissues than in noncancerous esophageal mucosae (P = 0.050). APOBEC3B expression was significantly correlated with PIK3CA mutation (P = 0.013), particularly with C to T transitions of PIK3CA (P = 0.041). Moreover, a high expression of APOBEC3B was significantly associated with LINE-1 hypomethylation (P = 0.027). Given the crucial roles of PIK3CA mutation and LINE-1 methylation levels, our findings might provide new insights into the biological mechanisms of ESCC tumorigenesis and progression.



Similar content being viewed by others
Abbreviations
- APOBEC:
-
Apolipoprotein B mRNA-editing enzyme catalytic polypeptide-like
- ESCC:
-
Esophageal squamous cell carcinoma
- LINE-1:
-
Long interspersed nucleotide element-1
- PCR:
-
Polymerase chain reaction
References
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136(5):E359–86.
Enzinger PC, Mayer RJ. Esophageal cancer. N Engl J Med. 2003;349(23):2241–52.
Allum WH, Stenning SP, Bancewicz J, Clark PI, Langley RE. Long-term results of a randomized trial of surgery with or without preoperative chemotherapy in esophageal cancer. J Clin Oncol. 2009;27(30):5062–7.
Rizk NP, Ishwaran H, Rice TW, Chen LQ, Schipper PH, Kesler KA, et al. Optimum lymphadenectomy for esophageal cancer. Ann Surg. 2010;251(1):46–50.
Thallinger CM, Raderer M, Hejna M. Esophageal cancer: a critical evaluation of systemic second-line therapy. J Clin Oncol. 2011;29(35):4709–14.
Shigaki H, Baba Y, Watanabe M, Murata A, Ishimoto T, Iwatsuki M, et al. PIK3CA mutation is associated with a favorable prognosis among patients with curatively resected esophageal squamous cell carcinoma. Clin Cancer Res. 2013;19(9):2451–9.
Murata A, Baba Y, Watanabe M, Shigaki H, Miyake K, Karashima R, et al. p53 immunohistochemical expression and patient prognosis in esophageal squamous cell carcinoma. Med Oncol. 2013;30(4):728.
Iwagami S, Baba Y, Watanabe M, Shigaki H, Miyake K, Ishimoto T, et al. LINE-1 hypomethylation is associated with a poor prognosis among patients with curatively resected esophageal squamous cell carcinoma. Ann Surg. 2013;257(3):449–55.
Gaudet F, Hodgson JG, Eden A, Jackson-Grusby L, Dausman J, Gray JW, et al. Induction of tumors in mice by genomic hypomethylation. Science. 2003;300(5618):489–92.
Jones PA, Baylin SB. The fundamental role of epigenetic events in cancer. Nat Rev Genet. 2002;3(6):415–28.
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490(7418):61–70.
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SA, Behjati S, Biankin AV, et al. Signatures of mutational processes in human cancer. Nature. 2013;500(7463):415–21.
Nik-Zainal S, Alexandrov LB, Wedge DC, Van Loo P, Greenman CD, Raine K, et al. Mutational processes molding the genomes of 21 breast cancers. Cell. 2012;149(5):979–93.
Roberts SA, Lawrence MS, Klimczak LJ, Grimm SA, Fargo D, Stojanov P, et al. An APOBEC cytidine deaminase mutagenesis pattern is widespread in human cancers. Nat Genet. 2013;45(9):970–6.
Burns MB, Temiz NA, Harris RS. Evidence for APOBEC3B mutagenesis in multiple human cancers. Nat Genet. 2013;45(9):977–83.
Conticello SG. The AID/APOBEC family of nucleic acid mutators. Genome Biol. 2008;9(6):229.
Burns MB, Lackey L, Carpenter MA, Rathore A, Land AM, Leonard B, et al. APOBEC3B is an enzymatic source of mutation in breast cancer. Nature. 2013;494(7437):366–70.
Taylor BJ, Nik-Zainal S, Wu YL, Stebbings LA, Raine K, Campbell PJ, et al. DNA deaminases induce break-associated mutation showers with implication of APOBEC3B and 3A in breast cancer kataegis. Elife. 2013;2:e00534.
Lawrence MS, Stojanov P, Polak P, Kryukov GV, Cibulskis K, Sivachenko A, et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature. 2013;499(7457):214–8.
Chan K, Roberts SA, Klimczak LJ, Sterling JF, Saini N, Malc EP, et al. An APOBEC3A hypermutation signature is distinguishable from the signature of background mutagenesis by APOBEC3B in human cancers. Nat Genet. 2015;47(9):1067–72.
Walker BA, Wardell CP, Murison A, Boyle EM, Begum DB, Dahir NM, et al. APOBEC family mutational signatures are associated with poor prognosis translocations in multiple myeloma. Nat Commun. 2015;6:6997.
Swanton C, McGranahan N, Starrett GJ, Harris RS. APOBEC enzymes: mutagenic fuel for cancer evolution and heterogeneity. Cancer Discov. 2015;5(7):704–12.
Siriwardena SU, Guruge TA, Bhagwat AS. Characterization of the catalytic domain of human APOBEC3B and the critical structural role for a conserved methionine. J Mol Biol. 2015;427(19):3042–55.
Leonard B, Hart SN, Burns MB, Carpenter MA, Temiz NA, Rathore A, et al. APOBEC3B upregulation and genomic mutation patterns in serous ovarian carcinoma. Cancer Res. 2013;73(24):7222–31.
Yang Z, Zhuang L, Yu Y, Zhou W, Lu Y, Xu Q, et al. Overexpression of APOBEC3F in tumor tissues is potentially predictive for poor recurrence-free survival from HBV-related hepatocellular carcinoma. Discov Med. 2015;20(112):349–56.
Zhang Y, Delahanty R, Guo X, Zheng W, Long J. Integrative genomic analysis reveals functional diversification of APOBEC gene family in breast cancer. Hum Genomics. 2015;9(1):34.
Land AM, Wang J, Law EK, Aberle R, Kirmaier A, Krupp A, et al. Degradation of the cancer genomic DNA deaminase APOBEC3B by SIV Vif. Oncotarget. 2015;6(37):39969–79.
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM, et al. Reporting recommendations for tumor marker prognostic studies (REMARK). J Natl Cancer Inst. 2005;97(16):1180–4.
Ogino S, Kawasaki T, Nosho K, Ohnishi M, Suemoto Y, Kirkner GJ, et al. LINE-1 hypomethylation is inversely associated with microsatellite instability and CpG island methylator phenotype in colorectal cancer. Int J Cancer. 2008;122(12):2767–73.
Ogino S, Nosho K, Kirkner GJ, Kawasaki T, Chan AT, Schernhammer ES, et al. A cohort study of tumoral LINE-1 hypomethylation and prognosis in colon cancer. J Natl Cancer Inst. 2008;100(23):1734–8.
Baba Y, Huttenhower C, Nosho K, Tanaka N, Shima K, Hazra A, et al. Epigenomic diversity of colorectal cancer indicated by LINE-1 methylation in a database of 869 tumors. Mol Cancer. 2010;9:125.
Macduff DA, Harris RS. Directed DNA deamination by AID/APOBEC3 in immunity. Curr Biol. 2006;16(6):R186–9.
Jarmuz A, Chester A, Bayliss J, Gisbourne J, Dunham I, Scott J, et al. An anthropoid-specific locus of orphan C to U RNA-editing enzymes on chromosome 22. Genomics. 2002;79(3):285–96.
Goila-Gaur R, Strebel K. HIV-1 Vif, APOBEC, and intrinsic immunity. Retrovirology. 2008;5:51.
Henderson S, Chakravarthy A, Su X, Boshoff C, Fenton TR. APOBEC-mediated cytosine deamination links PIK3CA helical domain mutations to human papillomavirus-driven tumor development. Cell Rep. 2014;7(6):1833–41.
Cescon DW, Haibe-Kains B, Mak TW. APOBEC3B expression in breast cancer reflects cellular proliferation, while a deletion polymorphism is associated with immune activation. Proc Natl Acad Sci USA. 2015;112(9):2841–6.
Sieuwerts AM, Willis S, Burns MB, Look MP, Meijer-Van Gelder ME, Schlicker A, et al. Elevated APOBEC3B correlates with poor outcomes for estrogen-receptor-positive breast cancers. Horm Cancer. 2014;5(6):405–13.
Periyasamy M, Patel H, Lai CF, Nguyen VT, Nevedomskaya E, Harrod A, et al. APOBEC3B-mediated cytidine deamination is required for estrogen receptor action in breast cancer. Cell Rep. 2015;13(1):108–21.
Xu L, Chang Y, An H, Zhu Y, Yang Y, Xu J. High APOBEC3B expression is a predictor of recurrence in patients with low-risk clear cell renal cell carcinoma. Urol Oncol. 2015;33(8):340e1–18.
Zhang J, Wei W, Jin HC, Ying RC, Zhu AK, Zhang FJ. The roles of APOBEC3B in gastric cancer. Int J Clin Exp Pathol. 2015;8(5):5089–96.
Acknowledgments
No funding was received for this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None.
Rights and permissions
About this article
Cite this article
Kosumi, K., Baba, Y., Ishimoto, T. et al. APOBEC3B is an enzymatic source of molecular alterations in esophageal squamous cell carcinoma. Med Oncol 33, 26 (2016). https://doi.org/10.1007/s12032-016-0739-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-016-0739-7