Abstract
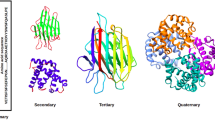
The biological function of protein depends mainly on its tertiary structure which is determined by its amino acid sequence via the process of protein folding. Prediction of protein structure from its amino acid sequence is one of the most prominent problems in computational biology. Two basic methodologies on protein structure prediction are combined: ab initio method (3-D space lattice) and fold recognition method (hidden Markov model). The primary structure of proteins and 3-D coordinates of amino acid residues are put together in one hidden Markov model to learn the path of amino acid residues in 3-D space from the first atom to the last atom of each protein of each fold. Therefore, each model has the information of 3-D path of amino acids of each fold. The proposed method is compared to fold recognition methods which have hidden Markov model as a base of their algorithms having approaches on only amino acid sequence or secondary structure. To validate the proposed method, the models are assessed with three datasets. Results show that the proposed models outperform 7-HMM and 3-HMM in the same dataset. The face-centered cubic lattice which is the most compacted 3-D lattice reached the maximum classification accuracy in all experiments in comparison with the performance of the most effective version of optimized 3-HMM as well as the performance of the latest version of SAM 3.5. Results show that 3-D coordinates of atoms of amino acids in proteins have an important role in prediction. It also has great hidden information as compared to secondary structure of proteins in fold classification.









Similar content being viewed by others
References
Bahamish HAA, Abdullah R, Salam RA (2009) Protein tertiary structure prediction using artificial bee colony algorithm. In: Third Asia international conference on modelling & simulation, pp 258–263
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H et al (2000) The Protein Data Bank. Nucl Acids Res 28:235–242
Bidargaddi NP, Chetty M, Kamruzzaman J (2009) Combining segmental semi-Markov models with neural networks for protein secondary structure prediction. Neurocomputing 72:3943–3950
Camproux AC, Tufféry P (2005) Hidden Markov Model-derived structural alphabet for proteins: the learning of protein local shapes captures sequence specificity. Biochem Biophys Acta 1724:394–403
Caoa H, Ihma Y, Wangb C-Z, Morrisb JR, Sua M, Dobbsc D et al (2004) Three-dimensional threading approach to protein structure recognition. Polymer 45:687–697
Chandonia JM, Hon G, Walker NS, Lo Conte L, Koehl P, Levitt M et al (2004) The ASTRAL Compendium in 2004. Nucleic Acids Res 32:D189–D192
Chmielnicki W, Stapor K (2012) A hybrid discriminative/generative approach to protein fold recognition,”. Neurocomputing 75:194–198
Deschavanne P, Tufféry P (2009) Enhanced protein fold recognition using a structural alphabet. Proteins 76:129–137
Dorn M, Silva MB, Buriol LS, Lamb LC (2014) Three-dimensional protein structure prediction: methods and computational strategies. Comput Biol Chem 53:251–276
Dotu I, Cebrian M, Van Hentenryck P, Clote P (2011) On lattice protein structure prediction revisited. IEEE/ACM Trans Comput Biol Bioinform 8:1620–1632
Elofsson A, Hargbo J (1999) Hidden Markov models that use predicted secondary structures for fold recognition. Proteins 36:68–76
Finn RD, Clements J, Eddy SR (2011) HMMER web server: interactive sequence similarity searching. Nucl Acids Res 39:W29–W37
Fox NK, Brenner SE, Chandonia JM (2015) The value of protein structure classification information-Surveying the scientific literature. Proteins Struct Funct Bioinform 83:2025–2038
Gheraibia Y, Moussaoui A (2012) Prediction of 3D protein structure using a genetic algorithm and a K nearest neighbour classifier. In: Biomedical engineering international conference BIOMEIC’12, Algeria
Karchin R, Cline M, Mandel-Gutfreund Y, Karplus K (2003) Hidden Markov models that use predicted local structure for fold recognition: alphabets of backbone geometry. Proteins 51:504–514
Karplus K, Sjölander K, Barrett C, Cline M, Haussler D, Hughey R et al (1997) Predicting protein structure using hidden Markov models. Proteins Struct Funct Bioinform 29:134–139
Karplus K, Karchin R, Shackelford G, Hughey R (2005) Calibrating E-values for hidden Markov models using reverse-sequence null models. Bioinformatics 21:4107–4115
Kong L, Zhang L (2014) Novel structure-driven features for accurate prediction of protein structural class. Genomics 103:292–297
Lampros C, Papaloukas C, Exarchos TP, Goletsis Y, Fotiadis DI (2007a) Sequence-based protein structure prediction using a reduced state-space hidden Markov model. Comput Biol Med 37:1211–1224
Lampros C, Papaloukas C, Exarchos K (2007b) Improvement in fold recognition accuracy of a reduced-state-space hidden Markov model by using secondary structure information in scoring. In: 29th annual international conference of the IEEE EMBS, France
Lampros C, Papaloukas C, Exarchos K, Fotiadis DI, Tsalikakis D (2009) Improving the protein fold recognition accuracy of a reduced state-space hidden Markov model. Comput Biol Med 39:907–914
Lampros C, Simos T, Exarchos TP, Exarchos KP, Papaloukas C, Fotiadis DI (2014) Assessment of optimized Markov models in protein fold classification. J Bioinform Comput Biol 12(4):1450016. https://doi.org/10.1142/S0219720014500164
Lampros C, Papaloukas C, Exarchos T, Fotiadis DI (2017) HMMs in Protein Fold Classification. Hidden Markov Models Methods Mol Biol 1552:13–27
Lee J, Kim S-Y, Joo K, Kim I, Lee J (2004) Prediction of protein tertiary structure using PROFESY, a novel method based on fragment assembly and conformational space annealing. Proteins Struct Funct Bioinform 56:704–714
Lee SY, Lee JY, Jung KS, Ryu KH (2009) A 9-state hidden Markov model using protein secondary structure information for protein fold recognition. Comput Biol Med 39:527–534
Lin C-J, Su S-C (2011) Protein 3D HP model folding simulation using a hybrid of genetic algorithm and particle swarm optimization. Int J Fuzzy Syst 13:140–147
Márquez-Chamorro AE, Divina F, Aguilar-Ruiz JS, Bacardit J, Asencio-Cortés G, Santiesteban-Toca CE (2012) A NSGA-II algorithm for the residue-residue contact prediction. Springer, Berlin, pp 234–244
Murzin AG, Brenner SE, Hubbard T, Chothia C (1995) SCOP: a structural classification of proteins database for the investigation of sequences and structures. J Mol Biol 247:536–540
Nanni L, Brahnamc S, Lumini A (2014) Prediction of protein structure classes by incorporating different protein descriptors into general Chou’s pseudo amino acid composition. J Theor Biol 360:109–116
Pitteri M, Zanzotto G (1996) On the definition and classification of Bravais lattices. Acta Cryst A52:830–838
Rashid MA, Newton MAH, Hoque MT, Sattar A (2013a) Mixing energy models in genetic algorithms for on-lattice protein structure prediction. BioMed Res Int 27:37–52
Rashid MA, Newton MAH, Hoque MT, Sattar A (2013b) A local search embedded genetic algorithm for simplified protein structure prediction. 2013 IEEE congress on evolutionary computation. https://doi.org/10.1109/CEC.2013.6557688
Regad L, Guyon F, Maupetit J, Tufféry P, Camproux AC (2008) A Hidden Markov Model applied to the protein 3D structure analysis. Comput Stat Data Anal 52:3198–3207
Shi J-Y, Zhang Y-N (2010) Using hierarchical hidden Markov models to perform sequence-based classification of protein structure. In: IEEE 10th international conference on signal processing, Beijing, pp 1789–1792
Song NY, Yan H (2013) Autoregressive and iterative hidden Markov models for periodicity detection and solenoid structure recognition in protein sequences. IEEE J Biomed Health Inform 17:436–441
Stanfel LE (1996) A new approach to clustering the amino acids. J Theor Biol 183:195–205
Tan C-W, Jones DT (2008) Using neural networks and evolutionary information in decoy discrimination for protein tertiary structure prediction. BMC Bioinform 94:19–42
Valavanis I, Spyrou G, Nikita K (2010) A similarity network approach for the analysis and comparison of protein sequence/structure sets. J Biomed Inform 43:257–267
Yang Y, Faraggi E, Zhao H, Zhou Y (2011) Improving protein fold recognition and template-based modeling by employing probabilistic-based matching between predicted one-dimensional structural properties of query and corresponding native properties of templates. Struct Bioinform 27:2076–2082
Yoon B-J (2009) Hidden Markov models and their applications in biological sequence analysis. Curr Genom 10:402–415
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Peyravi, F., Latif, A. & Moshtaghioun, S.M. A Composite Approach to Protein Tertiary Structure Prediction: Hidden Markov Model Based on Lattice. Bull Math Biol 81, 899–918 (2019). https://doi.org/10.1007/s11538-018-00542-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-018-00542-4