Abstract
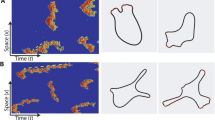
Two multiscale (hybrid) stochastic reaction–diffusion models of actin dynamics in a filopodium are investigated. Both hybrid algorithms combine compartment-based and molecular-based stochastic reaction–diffusion models. The first hybrid model is based on the models previously developed in the literature. The second hybrid model is based on the application of a recently developed two-regime method (TRM) to a fully molecular-based model, which is also developed in this paper. The results of hybrid models are compared with the results of the molecular-based model. It is shown that both approaches give comparable results, although the TRM model better agrees quantitatively with the molecular-based model.






Similar content being viewed by others
References
Andrews, S. (2005). Serial rebinding of ligands to clustered receptors as exemplified by bacterial chemotaxis. Phys. Biol., 2, 111–122.
Andrews, S., & Bray, D. (2004). Stochastic simulation of chemical reactions with spatial resolution and single molecule detail. Phys. Biol., 1, 137–151.
Arjunan, S., & Tomita, M. (2010). A new multicompartmental reaction–diffusion modeling method links transient membrane attachment of E. coli MinE to E-ring formation. Syst. Synth. Biol., 4(1), 35–53.
Cao, Y., Li, H., & Petzold, L. (2004). Efficient formulation of the stochastic simulation algorithm for chemically reacting systems. J. Chem. Phys., 121(9), 4059–4067.
Engblom, S., Ferm, L., Hellander, A., & Lötstedt, P. (2009). Simulation of stochastic reaction–diffusion processes on unstructured meshes. SIAM J. Sci. Comput., 31, 1774–1797.
Erban, R., & Chapman, S. J. (2007). Reactive boundary conditions for stochastic simulations of reaction–diffusion processes. Phys. Biol., 4(1), 16–28.
Erban, R., & Chapman, S. J. (2009). Stochastic modeling of reaction–diffusion processes: algorithms for bimolecular reactions. Phys. Biol., 6(4), 046001.
Erban, R., Chapman, S. J., & Maini, P. (2007). A practical guide to stochastic simulations of reaction–diffusion processes, 35 pages, arXiv:0704.1908.
Ferm, L., Hellander, A., & Lötstedt, P. (2010). An adaptive algorithm for simulation of stochastic reaction–diffusion processes. J. Comput. Phys., 229, 343–360.
Flegg, M., Chapman, J., & Erban, R. (2012). The two-regime method for optimizing stochastic reaction–diffusion simulations. J. R. Soc. Interface, 9(70), 859–868.
Flegg, M., Rüdiger, S., & Erban, R. (2013). Diffusive spatio-temporal noise in a first-passage time model for intracellular calcium release. J. Chem. Phys. doi:10.1063/1.4796417.
Flekkøy, E., Feder, J., & Wagner, G. (2001). Coupling particles and fields in a diffusive hybrid model. Phys. Rev. E, 64, 066302.
Franz, B., Flegg, M., Chapman, J., & Erban, R. (2013, to appear). Multiscale reaction–diffusion algorithms: PDE-assisted Brownian dynamics. SIAM J. Appl. Math. arXiv:1206.5860.
Gibson, M., & Bruck, J. (2000). Efficient exact stochastic simulation of chemical systems with many species and many channels. J. Phys. Chem. A, 104, 1876–1889.
Gillespie, D. (1977). Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem., 81(25), 2340–2361.
Hattne, J., Fange, D., & Elf, J. (2005). Stochastic reaction–diffusion simulation with MesoRD. Bioinformatics, 21(12), 2923–2924.
Hepburn, I., Chen, W., Wils, S., & De Schutter, E. (2012). STEPS: efficient simulation of stochastic reaction–diffusion models in realistic morphologies. BMC Syst. Biol., 6, 36.
Ho, C.-P. (2012). Multi-scale reaction diffusion simulations in biology. M.Sc. Thesis, University of Oxford.
Hu, L., & Papoian, G. A. (2010). Mechano-chemical feedbacks regulate actin mesh growth in lamellipodial protrusions. Biophys. J., 98(8), 1375–1384.
Lan, Y., & Papoian, G. A. (2008). The stochastic dynamics of filopodial growth. Biophys. J., 94, 3839–3852.
Lipkova, J., Zygalakis, K., Chapman, J., & Erban, R. (2011). Analysis of Brownian dynamics simulations of reversible bimolecular reactions. SIAM J. Appl. Math., 71(3), 714–730.
Moro, E. (2004). Hybrid method for simulating front propagation in reaction–diffusion systems. Phys. Rev. E, 69, 060101.
Noselli, S. (2002). Drosophila, actin and videotape—new insights in wound healing. Nat. Cell Biol., 4, 251–253.
Opplestrup, T., Bulatov, V., Donev, A., Kalos, M., Gilmer, G., & Sadigh, B. (2009). First-passage kinetic Monte Carlo method. Phys. Rev. E, 80(6), 066701.
Schaus, T., Taylor, E., & Borisy, G. (2007). Self-organization of actin filament orientation in the dendritic-nucleation/array-treadmilling model. Proc. Natl. Acad. Sci. USA, 104, 7086–7091.
Stiles, J., & Bartol, T. (2001). Monte Carlo methods for simulating realistic synaptic microphysiology using MCell. In E. Schutter (Ed.), Computational neuroscience: realistic modeling for experimentalists (pp. 87–127). Boca Raton: CRC Press.
van Zon, J., & ten Wolde, P. (2005). Green’s-function reaction dynamics: a particle-based approach for simulating biochemical networks in time and space. J. Chem. Phys., 123, 234910.
Wagner, G., & Flekkøy, E. (2004). Hybrid computations with flux exchange. Philos. Trans. R. Soc. A, Math. Phys. Eng. Sci., 362, 1655–1665.
Yamazaki, D., Kurisu, S., & Takenawa, T. (2005). Regulation of cancer cell motility through actin reorganization. Cancer Sci., 96(7), 379–386.
Zhuravlev, P., & Papoian, G. (2009). Molecular noise of capping protein binding induces macroscopic instability in filopodial dynamics. Proc. Natl. Acad. Sci. USA, 106(28), 11570–11575.
Zhuravlev, P., & Papoian, G. A. (2011). Protein fluxes along the filopodium as a framework for understanding the growth-retraction dynamics: the interplay between diffusion and active transport. Cell Adhes. Migr., 5(5), 448–456.
Zhuravlev, P., Der, B., & Papoian, G. A. (2010). Design of active transport must be highly intricate: a possible role of myosin and ena/VASP for G-actin transport in filopodia. Biophys. J., 98, 1439–1448.
Zhuravlev, P., Lan, Y., Minakova, M., & Papoian, G. A. (2012). Theory of active transport in filopodia and stereocilia. Proc. Natl. Acad. Sci. USA, 109, 10849–10854.
Acknowledgements
The research leading to these results has received funding from the European Research Council under the European Community’s Seventh Framework Programme (FP7/2007–2013)/ ERC grant agreement No. 239870. Radek Erban would also like to thank Brasenose College, University of Oxford, for a Nicholas Kurti Junior Fellowship; the Royal Society for a University Research Fellowship; and the Leverhulme Trust for a Philip Leverhulme Prize. This prize money was used to support a research visit of Garegin Papoian in Oxford. Garegin Papoian was also supported by the National Science Foundation CAREER Award CHE-0846701.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Erban, R., Flegg, M.B. & Papoian, G.A. Multiscale Stochastic Reaction–Diffusion Modeling: Application to Actin Dynamics in Filopodia. Bull Math Biol 76, 799–818 (2014). https://doi.org/10.1007/s11538-013-9844-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-013-9844-3