Abstract
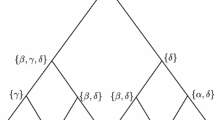
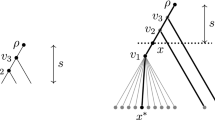
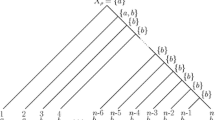
The accuracy of the Fitch method for reconstructing ancestral states on ultrametric phylogenetic trees is studied. Two recurrence relations for computing the accuracy are given here. Using these relations, we analyze the convergence of the accuracy of the Fitch method for reconstructing the root state on a complete binary tree of 2n leaves as n goes to infinity, present a closed-form formula for the accuracy on ultrametric comb trees, and provide a lower bound on the accuracy on arbitrary ultrametric phylogenetic trees.
Similar content being viewed by others
References
Baba, M.L., Goodman, M., Berger-Cohn, J., Demaille, J.G., Matsuda, G., 1984. The early adaptive evolution of calmodulin. Mol. Biol. Evol. 1, 442–455.
Bleher, P.M., Ruiz, J., Zagrebnov, V.A., 1995. On the purity of the limiting Gibbs state for the Ising model on the Bethe lattice. J. Stat. Phys. 79, 473–482.
Evans, W., Kenyon, C., Peres, Y., 2000. Broadcasting on trees and the Ising model. Ann. Appl. Probab. 10, 410–433.
Fischer, M., Thatte, B.D., 2009. Maximum parsimony on subsets of taxa. J. Theor. Biol. 260, 290–293.
Fitch, W.M., 1971. Toward defining the course of evolution: Minimum change for a specific tree topology. Syst. Zool. 20, 406–416.
Li, G.L., Steel, M., Zhang, L.X., 2008. More taxa are not necessarily better for the reconstruction of ancestral character states. Syst. Biol. 57, 647–653.
Li, G.L., Ma, J., Zhang, L.X., 2009. Greedy selection of species for ancestral state reconstruction on phylogenies: Elimination is better than insertion. PLOS One, in press.
Liberles, D.A. (Ed.), 2007. Ancestral Sequence Reconstruction. Oxford University Press, New York.
Maddison, W.P., 1995. Calculating the probability distributions of ancestral states reconstructed by parsimony on phylogenetic trees. Syst. Biol. 44, 474–481.
Mossel, E., 1998. Recursive reconstruction on periodic trees. Random Struct. Algorithms 16, 252–260.
Pauling, L., Zuckerkandl, E., 1963. Chemical paleogenetics: molecular restoration studies of extinct forms of lives. Acta Chem. Scand. 17, S9–S16.
Pachter, L., 2007. An introduction to reconstructing ancestral genomes. In: Proceedings of Symposia in Applied Mathematics, AMS Short Course Subseries, vol. 64, pp. 1–20.
Salisbury, B.A., Kim, J., 2001. Ancestral state estimation and taxon sampling density. Syst. Biol. 50, 557–564.
Steel, M., 1989. Distribution in bicoloured evolutionary trees. PhD thesis, Massey University, New Zealand.
Steel, M., Charleston, M., 1995. Five surprising properties of parsimoniously colored tree. Bull. Math. Biol. 57, 367–375.
Steel, M.A., Székely, L.A., 2007. Teasing apart two trees. Comb. Probab. Comput. 16, 903–922.
Thornton, J.W., 2004. Resurrecting ancient genes: Experimental analysis of extinct molecules. Nat. Rev. Genet. 5, 366–375.
Zhang, J., Nei, M., 1997. Accuracies of ancestral amino acid sequences inferred by parsimony, likelihood, and distance methods. J. Mol. Evol. 44, 139–146.
Author information
Authors and Affiliations
Corresponding author
Additional information
Work of L. Zhang was supported by ARF(R146-000-109-112).
Work of J. Shen was partially supported by NSF (CNS 0835834) and Texas Higher Education Coordinating Board (ARP 003615-0039-200).
Rights and permissions
About this article
Cite this article
Zhang, L., Shen, J., Yang, J. et al. Analyzing the Fitch Method for Reconstructing Ancestral States on Ultrametric Phylogenetic Trees. Bull. Math. Biol. 72, 1760–1782 (2010). https://doi.org/10.1007/s11538-010-9505-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-010-9505-8