Abstract
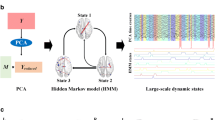
Functional connectivity measured from resting state fMRI (R-fMRI) data has been widely used to examine the brain’s functional activities and has been recently used to characterize and differentiate brain conditions. However, the dynamical transition patterns of the brain’s functional states have been less explored. In this work, we propose a novel computational framework to quantitatively characterize the brain state dynamics via hidden Markov models (HMMs) learned from the observations of temporally dynamic functional connectomics, denoted as functional connectome states. The framework has been applied to the R-fMRI dataset including 44 post-traumatic stress disorder (PTSD) patients and 51 normal control (NC) subjects. Experimental results show that both PTSD and NC brains were undergoing remarkable changes in resting state and mainly transiting amongst a few brain states. Interestingly, further prediction with the best-matched HMM demonstrates that PTSD would enter into, but could not disengage from, a negative mood state. Importantly, 84 % of PTSD patients and 86 % of NC subjects are successfully classified via multiple HMMs using majority voting.










Similar content being viewed by others
References
Allen EA, Damaraju E, Plis SM, Erhardt EB, Eichele T, Calhoun VD (2012) Tracking Whole-Brain Connectivity Dynamics in the Resting State. Cereb Cortex
Bassett DS, Wymbs NF, Porter MA, Mucha PJ, Carlson JM, Grafton ST (2011) Dynamic reconfiguration of human brain networks during learning. Proc Natl Acad Sci USA 108(18):7641–7646
Binnewijzend MA, Schoonheim MM, Sanz-Arigita E, Wink AM, van der Flier WM, Tolboom N, Adriaanse SM, Damoiseaux JS, Scheltens P, van Berckel BN, Barkhof F (2012) Resting-state fMRI changes in Alzheimer’s disease and mild cognitive impairment. Neurobiol Aging 33(9):2018–2028
Biswal BB (2012) Resting state fMRI: a personal history. Neuroimage 62(2):938–944
Biswal BB, Mennes M, Zuo XN, Gohel S, Kelly C, Smith SM, Beckmann CF, Adelstein JS, Buckner RL, Colcombe S, Dogonowski AM, Ernst M, Fair D, Hampson M, Hoptman MJ, Hyde JS, Kiviniemi VJ, Kotter R, Li SJ, Lin CP, Lowe MJ, Mackay C, Madden DJ, Madsen KH, Margulies DS, Mayberg HS, McMahon K, Monk CS, Mostofsky SH, Nagel BJ, Pekar JJ, Peltier SJ, Petersen SE, Riedl V, Rombouts SA, Rypma B, Schlaggar BL, Schmidt S, Seidler RD, Siegle GJ, Sorg C, Teng GJ, Veijola J, Villringer A, Walter M, Wang L, Weng XC, Whitfield-Gabrieli S, Williamson P, Windischberger C, Zang YF, Zhang HY, Castellanos FX, Milham MP (2010) Toward discovery science of human brain function. Proc Natl Acad Sci USA 107(10):4734–4739
Chang C, Glover GH (2010) Time-frequency dynamics of resting-state brain connectivity measured with fMRI. Neuroimage 50(1):81–98
Cordes D, Haughton V, Carew JD, Arfanakis K, Maravilla K (2002) Hierarchical clustering to measure connectivity in fMRI resting-state data. Magn Reson Imaging 20(4):305–317
Dickerson BC, Sperling RA (2009) Large-scale functional brain network abnormalities in Alzheimer’s disease: insights from functional neuroimaging. Behav Neurol 21(1):63–75
Dijk KRAV, Hedden T, Venkataraman A, Evans KC, Lazar SW, Buckner RL (2010) Intrinsic functional connectivity as a tool for human connectomics: theory, properties, and optimization. J Neurophysiol 103(1):297–321
Eavani H, Satterthwaite T, Gur R, Gur R, Davatzikos C (2013) Unsupervised Learning of Functional Network Dynamics in Resting State fMRI. In: Gee J, Joshi S, Pohl K, Wells W, Zöllei L (eds) Information Processing in Medical Imaging, vol 7917., Lecture notes in computer scienceSpringer, Berlin Heidelberg, pp 426–437
Ekman M, Derrfuss J, Tittgemeyer M, Fiebach CJ (2012) Predicting errors from reconfiguration patterns in human brain networks. Proc Natl Acad Sci USA 109(41):16714–16719
Fox MD, Raichle ME (2007) Spontaneous fluctuations in brain activity observed with functional magnetic resonance imaging. Nat Rev Neurosci 8(9):700–711
Francati V, Vermetten E, Bremner JD (2007) Functional neuroimaging studies in posttraumatic stress disorder: review of current methods and findings. Depress Anxiety 24(3):202–218
Gilbert CD, Sigman M (2007) Brain states: top-down influences in sensory processing. Neuron 54(5):677–696
Hagmann P, Cammoun L, Gigandet X, Gerhard S, Ellen Grant P, Wedeen V, Meuli R, Thiran J-P, Honey CJ, Sporns O (2010) MR connectomics: principles and challenges. J Neurosci Methods 194(1):34–45
Holtzheimer PE, Mayberg HS (2011) Stuck in a rut: rethinking depression and its treatment. Trends Neurosci 34(1):1–9
Jain AK, Murty MN, Flynn PJ (1999) Data Clustering: a Review. ACM Comput Surv 31(3):264–323
Jinli O, Li X, Peng W, Xiang L, Dajiang Z, Rongxin J, Yufeng W, Yaowu C, Jing Z, Tianming L Modeling brain functional dynamics via hidden Markov models. In: Neural Engineering (NER), 2013 6th International IEEE/EMBS Conference on 6–8 Nov. 2013. pp 569–572
Kennedy D (2010) Making Connections in the Connectome Era. Neuroinformatics 8(2):61–62
Kittler J, Hatef M, Duin RPW, Matas J (1998) On combining classifiers. IEEE Trans Pattern Anal Mach Intell 20(3):226–239
Li K, Guo L, Nie J, Li G, Liu T (2009) Review of methods for functional brain connectivity detection using fMRI. Comput Med Imaging Graph 33(2):131–139
Li K, Zhu D, Guo L, Li Z, Lynch ME, Coles C, Hu X, Liu T (2013a) Connectomics signatures of prenatal cocaine exposure affected adolescent brains. Hum Brain Mapp 34(10):2494–2510
Li X, Lim C, Li K, Guo L, Liu T (2013b) Detecting brain state changes via fiber-centered functional connectivity analysis. Neuroinformatics 11(2):193–210
Li X, Zhu D, Jiang X, Jin C, Zhang X, Guo L, Zhang J, Hu X, Li L, Liu T (2014) Dynamic functional connectomics signatures for characterization and differentiation of PTSD patients. Hum Brain Mapp 35(4):1761–1778
Liang P, Wang Z, Yang Y, Jia X, Li K (2011) Functional disconnection and compensation in mild cognitive impairment: evidence from DLPFC connectivity using resting-state fMRI. PLoS ONE 6(7):e22153
Lynall M-E, Bassett DS, Kerwin R, McKenna PJ, Kitzbichler M, Muller U, Bullmore E (2010) Functional connectivity and brain networks in schizophrenia. J Neurosci 30(28):9477–9487
Majeed W, Magnuson M, Hasenkamp W, Schwarb H, Schumacher EH, Barsalou L, Keilholz SD (2011) Spatiotemporal dynamics of low frequency BOLD fluctuations in rats and humans. Neuroimage 54(2):1140–1150
Murtagh F (1983) A survey of recent advances in hierarchical clustering algorithms. Comput J 26(4):354–359
Nianjun Liu, Davis Rial BC, Kootsookos PJ (2004) Effect of initial HMM choices in multiple sequence training for gesture recognition. International Conference on Information Technology: Coding and Computing 1:608–613
Passingham RE, Stephan KE, Kotter R (2002) The anatomical basis of functional localization in the cortex. Nat Rev Neurosci 3(8):606–616
Rabiner LR (1989) A tutorial on hidden Markov models and selected applications in speech recognition. Proc IEEE 77(2):257–286
Richiardi J, Eryilmaz H, Schwartz S, Vuilleumier P, Van De Ville D (2011) Decoding brain states from fMRI connectivity graphs. Neuroimage 56(2):616–626
Ruta D, Gabrys B (2005) Classifier selection for majority voting. Information Fusion 6(1):63–81
Schwarz G (1978) Estimating the dimension of a model. Ann Stat 6(2):461–464
Smith SM, Miller KL, Moeller S, Xu J, Auerbach EJ, Woolrich MW, Beckmann CF, Jenkinson M, Andersson J, Glasser MF, Van Essen DC, Feinberg DA, Yacoub ES, Ugurbil K (2012) Temporally-independent functional modes of spontaneous brain activity. Proc Natl Acad Sci USA 109(8):3131–3136
Sporns O, Tononi G, Kötter R (2005) The human connectome: a structural description of the human brain. PLoS Comput Biol 1(4):e42
Stebbins GT, Murphy CM (2009) Diffusion tensor imaging in Alzheimer’s disease and mild cognitive impairment. Behav Neurol 21(1):39–49
Stratonovich RL (1960) Conditional Markov Processes. Theory Prob Appl 5(2):156–178
Williams R (2010) The human connectome: just another ‘ome? Lancet Neurol 9(3):238–239
Zhang X, Guo L, Li X, Zhang T, Zhu D, Li K, Chen H, Lv J, Jin C, Zhao Q, Li L, Liu T (2013) Characterization of task-free and task-performance brain states via functional connectome patterns. Med Image Anal 17(8):1106–1122
Zhu D, Li K, Guo L, Jiang X, Zhang T, Zhang D, Chen H, Deng F, Faraco C, Jin C, Wee C-Y, Yuan Y, Lv P, Yin Y, Hu X, Duan L, Hu X, Han J, Wang L, Shen D, Miller LS, Li L, Liu T (2013) DICCCOL: dense individualized and common connectivity-based cortical landmarks. Cereb Cortex 23(4):786–800
Acknowledgments
T Liu was supported by NIH R01 DA033393, NIH R01 AG-042599, NSF CBET-1302089 and NSF CAREER Award IIS-1149260. Lingjiang Li was supported by The National Natural Science Foundation of China (30830046) and The National 973 Program of China (2009 CB918303). J Zhang was supported by start-up funding and Sesseel Award from Yale University. The authors would like to thank the anonymous reviewers for their constructive comments and thank Elliott Chung for his proofreading.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Ou, J., Xie, L., Jin, C. et al. Characterizing and Differentiating Brain State Dynamics via Hidden Markov Models. Brain Topogr 28, 666–679 (2015). https://doi.org/10.1007/s10548-014-0406-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10548-014-0406-2