Abstract
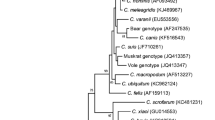
A total of 124 fecal specimens were collected from four deer farms in Zhengzhou City, China and examined for Cryptosporidium by Sheather’s sugar flotation technique. Cryptosporidim oocysts were detected in two 1-year-old sika deer, and one of the two specimens was genotyped by sequence and phylogenetic analyses of the small subunit ribosomal RNA (rRNA) (18S rRNA), 70-kDa heat shock protein (HSP70), actin, and Cryptosporidium oocyst wall protein (COWP) genes. Results obtained suggested that the Cryptosporidium studied belonged to Cryptosporidium cervine genotype, although slight sequence differences were noticed at the three loci. The similarities between this isolate and other Cryptosporidium cervine genotype isolates were 99.1–99.8%, 9.8%, 99.7%, and 100% at the 18S rRNA, HSP70, actin, and COWP loci, respectively. This study is the first report of Cryptosporidium infection in sika deer in China.

Similar content being viewed by others
References
Blackburn BG, Mazurek JM, Hlavsa M, Park J, Tillapaw M, Parrish M, Salehi E, Franks W, Koch E, Smith F, Xiao L, Arrowood M, Hill V, da Silva A, Johnston S, Jones JL (2006) Cryptosporidiosis associated with ozonated apple cider. Emerg Infect Dis 12:684–686
da Silva AJ, Cacciò S, Williams C, Won KY, Nace EK, Whittier C, Pieniazek NJ, Eberhard ML (2003) Molecular and morphologic characterization of a Cryptosporidium genotype identified in lemurs. Vet Parasitol 111:297–307
Felsenstein J (1989) PHYLIP: phylogeny inference package (version 3.2). Cladistics 5:164–166
Feltus DC, Giddings CW, Schneck BL, Monson T, Warshauer D, McEvoy JM (2006) Evidence supporting zoonotic transmission of Cryptosporidium spp. in Wisconsin. J Clin Microbiol 44:4303–4308
Feng Y, Alderisio KA, Yang W, Blancero LA, Kuhne WG, Nadareski CA, Reid M, Xiao L (2007) Cryptosporidium genotypes in wildlife from a New York watershed. Appl Environ Microbiol 73:6475–6483
Jiang J, Alderisio KA, Xiao L (2005) Distribution of Cryptosporidium genotypes in storm event water samples from three watersheds in New York. Appl Environ Microbio 171:4446–4454
Karanis P, Plutzer J, Halim NA, Igori K, Nagasawa H, Ongerth J, Liqing M (2007) Molecular characterization of Cryptosporidium from animal sources in Qinghai province of China. Parasitol Res 101:1575–1580
Leoni F, Amar C, Nichols G, Pedraza-Diaz S, McLauchlin J (2006) Genetic analysis of Cryptosporidium from 2414 humans with diarrhoea in England between 1985 and 2000. J Med Microbiol 55:703–707
Nichols GL, Chalmers RM, Sopwith W, Regan M, Hunter CA, Grenfell P, Harrison F, Lane C (2006) Cryptosporidiosis: a report on the surveillance and epidemiology of Cryptosporidium infection in England and Wales. Drinking Water Directorate Contract Number DWI 70/2/201. Drinking Water Inspectorate, UK, 142
Ong CS, Eisler DL, Alikhani A, Fung VW, Tomblin J, Bowie WR, Isaac-Renton JL (2002) Novel Cryptosporidium genotypes in sporadic cryptosporidiosis cases: first report of human infections with a cervine genotype. Emerg Infect Dis 8:263–268
Perz JF, Le Blancq SM (2001) Cryptosporidium parvum infection involving novel genotypes in wildlife from lower New York State. Appl Environ Microbiol 67:1154–1162
Ruecker NJ, Braithwaite SL, Topp E, Edge T, Lapen DR, Wilkes G, Robertson W, Medeiros D, Sensen CW, Neumann NF (2007) Tracking host sources of Cryptosporidium spp. in raw water for improved health risk assessment. Appl Environ Microbiol 73:3945–3957
Ryan UM, Xiao L, Read C, Zhou L, Lal AA, Pavlasek I (2003) Identification of novel Cryptosporidium genotypes from the Czech Republic. Appl Environ Microbiol 69:4302–4307
Ryan U, Bath C, Robertson I, Read C, Elliot A, McInnes L, Traub R, Besier B (2005) Sheep may not be an important zoonotic reservoir for Cryptosporidium and Giardia parasites. Appl Environ Microbiol 71:4992–4997
Santin M, Trout JM, Fayer R (2007) Prevalence and molecular characterization of Cryptosporidium and Giardia species and genotypes in sheep in Maryland. Vet Parasitol 146:17–24
Soba B, Petrovec M, Mioc V, Logar J (2006) Molecular characterisation of Cryptosporidium isolates from humans in Slovenia. Clin Microbiol Infect 12:918–921
Trotz-Williams LA, Martin DS, Gatei W, Cama V, Peregrine AS, Martin SW, Nydam DV, Jamieson F, Xiao L (2006) Genotype and subtype analyses of Cryptosporidium isolates from dairy calves and humans in Ontario. Parasitol Res 99:346–352
Xiao L, Escalante L, Yang C, Sulaiman I, Escalante AA, Montali RJ, Fayer R, Lal AA (1999) Phylogenetic analysis of Cryptosporidium parasites based on the small-subunit rRNA gene locus. Appl Environ Microbiol 65:1578–1583
Xiao L, Aldeeisio K, Limor J, Royer M, Lal AA (2000) Identification of species and sources of Cryptopsporidium oocysts in storm waters with a small-subunit rRNA-based diagnostic and genotyping tool. Appl Environ Microbiol 66:5492–5498
Xiao LH, Fayer R, Ryan U, Upton SJ (2004) Cryptosporidium taxonomy: recent advances and implications for public health. Clin Microbiol 17:72–97
Xiao L, Alderisio K, Singh A (2006a) Development and standardization of a Cryptosporidium genotyping tool for water samples. Awwa Research Foundation, Denver, CO
Xiao L, Alderisio KA, Jiang J (2006b) Detection of cryptosporidium oocysts in water: effect of the number of samples and analytic replicates on test results. Appl Environ Microbiol 72:5942–5947
Zhou L, Singh A, Jiang J, Xiao L (2003) Molecular surveillance of Cryptosporidium spp. in raw wastewater in Milwaukee: implications for understanding outbreak occurrence and transmission dynamics. J Clin Microbiol 41:5254–5257
Ziegler PE, Wade SE, Schaaf SL, Chang YF, Mohammed HO (2007) Cryptosporidium spp. from small mammals in the New York city watershed. J Wildl Dis 43:586–596
Acknowledgments
This study was supported in part by Henan Innovation Project for Prominent University Research Talents (number 2004KYCX002).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Wang, R., Wang, J., Sun, M. et al. Molecular characterization of the Cryptosporidium cervine genotype from a sika deer (Cervus nippon Temminck) in Zhengzhou, China and literature review. Parasitol Res 103, 865–869 (2008). https://doi.org/10.1007/s00436-008-1069-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-008-1069-2