Abstract
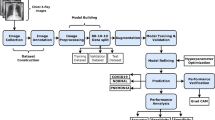
In the context of the COVID-19 pandemic, early diagnosis and treatment are crucial to avoid a drastic increase in the number of new infections. Chest imaging plays an important role in the early detection of the disease since it can be used to identify the initial phase lung infection caused by the SARS-CoV-2 virus. Recently, some researchers have begun to explore Convolutional Neural Networks (CNNs) to detect COVID-19 cases from chest X-ray images. CNN is a category of deep artificial neural networks that has demonstrated great success in computer vision applications, such as video and image analysis. However, this type of network is still affected by abnormal real-world factors such as unbalanced data, presence of noise, blurring, or other quality degradation that can diminish their overall performance when they are not properly handled. This work introduces a methodology to explore and compare the overall accuracy achieved by an Xception-based CNN model when it is combined with benchmark techniques such as unsharp masking, batch balance, and data augmentation aiming at enhancing image details (such as the bone structure and lung) and handling unbalanced datasets. Experiments are done referring to the COVIDx dataset. Preliminary results demonstrate that the proposed methodology leads to higher accuracy when implementing both image enhancement and batch balancing.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Alom, M.Z., Rahman, M.M.S., Nasrin, M.S., Taha, T.M., Asari, V.K.: COVID\(_{M}\)TNet: COVID-19 detection with multi-task deep learning approaches, April 2020. http://arxiv.org/abs/2004.03747
Apostolopoulos, I.D., Mpesiana, T.A.: Covid-19: automatic detection from X-ray images utilizing transfer learning with convolutional neural networks. Phys. Eng. Sci. Med. 43(2), 635–640 (2020). https://doi.org/10.1007/s13246-020-00865-4
Ayan, E., Unver, H.M.: Diagnosis of pneumonia from chest X-Ray images using deep learning. In: 2019 Scientific Meeting on Electrical-Electronics & Biomedical Engineering and Computer Science (EBBT), pp. 1–5. IEEE, April 2019. https://doi.org/10.1109/EBBT.2019.8741582, https://ieeexplore.ieee.org/document/8741582/
Chollet, F.: Xception: deep learning with depthwise separable convolutions, October 2016. http://arxiv.org/abs/1610.02357
Cohen, J.P., Morrison, P., Dao, L.: Covid-19 image data collection. arXiv 2003.11597 (2020). https://github.com/ieee8023/covid-chestxray-dataset
Frid-Adar, M., Diamant, I., Klang, E., Amitai, M., Goldberger, J., Greenspan, H.: GAN-based synthetic medical image augmentation for in- creased CNN performance in liver lesion classification. Neurocomputing 321, 321–331 (2018). https://doi.org/10.1016/j.neucom.2018.09.013. https://linkinghub.elsevier.com/retrieve/pii/S0925231218310749
Han, C., Murao, K., Satoh, S., Nakayama, H.: Learning more with less: GAN-based medical image augmentation, March 2019. http://arxiv.org/abs/1904.00838
Kassani, S.H., Kassani, P.H., Wesolowski, M.J., Schneider, K.A., Deters, R.: Breast cancer diagnosis with transfer learning and global pooling, September 2019. http://arxiv.org/abs/1909.11839
Liang, G., Hong, H., Xie, W., Zheng, L.: Combining convolutional neural network with recursive neural network for blood cell image classification. IEEE Access 6, 36188–36197 (2018). https://doi.org/10.1109/ACCESS.2018.2846685. https://ieeexplore.ieee.org/document/8402091/
Mikołajczyk, A., Grochowski, M.: Data augmentation for improving deep learning in image classification problem. In: 2018 International Interdisciplinary PhD Workshop, pp. 117–122, May 2018. https://doi.org/10.1109/IIPHDW.2018.8388338
Ozturk, T., Talo, M., Yildirim, E.A., Baloglu, U.B., Yildirim, O., Rajendra Acharya, U.: Automated detection of COVID-19 cases using deep neural networks with X-ray images. Comput. Biol. Med. 121, 103792 (2020). https://doi.org/10.1016/j.compbiomed.2020.103792. https://linkinghub.elsevier.com/retrieve/pii/S0010482520301621
Redmon, J., Farhadi, A.: YOLOv3: an incremental improvement, April 2018. http://arxiv.org/abs/1804.02767
Sandfort, V., Yan, K., Pickhardt, P.J., Summers, R.M.: Data augmentation using generative adversarial networks (CycleGAN) to improve generalizability in CT segmentation tasks. Sci. Rep. 9(1), 16884 (2019). https://doi.org/10.1038/s41598-019-52737-x. http://www.nature.com/articles/s41598-019-52737-x
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J., Wojna, Z.: Rethinking the inception architecture for computer vision, December 2016. http://arxiv.org/abs/1512.00567
Wang, L., Wong, A.: Covid-net: a tailored deep convolutional neural network design for detection of covid-19 cases from chest radiography images (2020)
Yan, Q., et al.: COVID-19 chest CT image segmentation - a deep convolutional neural network solution, April 2020. http://arxiv.org/abs/2004.10987
Zou, K.H., O’Malley, A.J., Mauri, L.: Receiver-operating characteristic analysis for evaluating diagnostic tests and predictive models. Circulation 115(5), 654–657 (2007). https://doi.org/10.1161/CIRCULATIONAHA.105.594929. https://www.ahajournals.org/doi/10.1161/CIRCULATIONAHA.105.594929
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this paper
Cite this paper
Mejia, H., Guzman, F., Bustamante-Orellana, C., Guachi-Guachi, L. (2020). Pre-processing and Handling Unbalanced Data in CNN for Improving Automated Detection of COVID-19 Cases: Preliminary Results. In: Rodriguez Morales, G., Fonseca C., E.R., Salgado, J.P., Pérez-Gosende, P., Orellana Cordero, M., Berrezueta, S. (eds) Information and Communication Technologies. TICEC 2020. Communications in Computer and Information Science, vol 1307. Springer, Cham. https://doi.org/10.1007/978-3-030-62833-8_11
Download citation
DOI: https://doi.org/10.1007/978-3-030-62833-8_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-62832-1
Online ISBN: 978-3-030-62833-8
eBook Packages: Computer ScienceComputer Science (R0)