Abstract
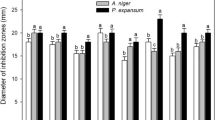
The aim of the study was a detail evaluation of genetic diversity among the lactic acid bacteria (LAB) strains having an advantage of a starter culture in order to select genotypically diverse strains with enhanced antimicrobial effect on some harmfull and pathogenic microorganisms. Antimicrobial activity of LAB was performed by the agar well diffusion method and was examined against the reference strains and foodborne isolates of Bacillus cereus, Listeria monocytogenes, Escherichia coli, Staphylococcus aureus and Salmonella Typhimurium. Antifungal activity was tested against the foodborne isolates of Candida parapsilosis, Debaromyces hansenii, Kluyveromyces marxianus, Pichia guilliermondii, Yarowia lipolytica, Aspergillus brasiliensis, Aspergillus versicolor, Cladosporium herbarum, Penicillium chrysogenum and Scopulariopsis brevicaulis.
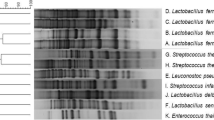
A total 40 LAB strains representing Lactobacillus (23 strains), Lactococcus (13 strains) and Streptococcus spp. (4 strains) were characterised by repetitive sequence based polymerase chain reaction fingerprinting which generated highly discriminatory profiles, confirmed the identity and revealed high genotypic heterogeneity among the strains. Many of tested LAB demonstrated strong antimicrobial activity specialised against one or few indicator strains. Twelve LAB strains were superior in suppressing growth of the whole complex of pathogenic bacteria and fungi. These results demonstrated that separate taxonomic units offered different possibilities of selection for novel LAB strains could be used as starter cultures enhancing food preservation.

Similar content being viewed by others
References
Cakir I (2010) Antibacterial and antifungal activities of some lactic acid bacteria isolated from naturally fermented herbs. J Food Agric Environ 8:223–226
Cizeikiene D, Juodeikiene G, Paskevicius A, Bartkiene E (2013) Antimicrobial activity of lactic acid bacteria against pathogenic and spoilage microorganism isolated from food and their control in wheat bread. Food Control 31:539–545
Claesson MJ, van Sinderen D, O'Toole PW (2008) Lactobacillus phylogenomics – towards a reclassification of the genus. Int J Syst Evol Micr 58:2945–2954
Domsch KH, Gams W, Anderson TH (1980) Compendium of soil fungi. Academic, London
Durlu-Özkaya F, Karabicak N, Kayali R, Esen B (2005) Inhibition of yeasts isolated from traditional Turkish cheeses by Lactobacillus spp. Int J Dairy Technol 58:111–114
Fhoula I, Najjari A, Turki Y, Jaballah S, Boudabous A, Ouzari H (2013) Diversity and Antimicrobial Properties of Lactic Acid Bacteria Isolated from Rhizosphere of Olive Trees and Desert Truffles of Tunisia. Biomed Res Int. http://dx.doi.org/10.1155/2013/405708 [12 September 2012].
Garcia-Vallve S, Palau J (1999) Romeu A, Horizontal gene transfer in glycosyl hydrolases inferred from codon usage in Escherichia coli and Bacillus subtilis. Mol Biol Evol 16:1125–1134
Gerez CL, Torino MI, Rollan G, Font de Valdez G (2009) Prevention of bread mould spoilage by using lactic acid bacteria with antifungal properties. Food Control 20:144–148
Gevers D, Huys G, Swings J (2001) Applicability of rep-PCR fingerprinting for identification of Lactobacillus species. FEMS Microbiol Lett 205:31–36
Kurtzman CP, Fell JW, Boekhout T, Robert V (2011) Methods for isolation, phenotypic characterization and maintenance of yeasts. In: Kurtzman CP, Fell JW, Boekhout T (eds) The Yeasts, a Taxonomic Study, 5th edn. Elsevier, Amsterdam, pp 87–110
Magnusson J, Ström K, Roos S, Sjögren J, Schnürer J (2003) Broad and complex antifungal activity among environmental is olates of lactic acid bacteria. FEMS Microbiol Lett 219:129–135
Melvydas V, Šakalytė J, Paškevičius A, Gedminienė G (2007) Search for biological control agents against Candida yeasts and other dermatomycetes. Biologija 18:45–49
Mezaini A, Bouras AD (2013) Antibacterial activity and probiotic properties of some lactic acid bacteria isolated from dairy products. Afr J Biotechnol 12:2949–2956
Mori S, Mori K, Suzuki I, Kasumi T (2004) Phylogenetic analysis of Lactococcus lactis subspecies based on decoding the sequence of the pepT tripeptidase gene, the pepV dipeptidase gene and 16S rRNA. Syst Appl Microbiol 27:414–422
Neyriz-Nagadehi M, Razavi-Rohani SM, Karim G, Zeynali A (2012) The Effects of Monolaurinand Lactic Acid Bacteria Starter Culture on Growth of Vegetative Cells of Bacillus cereus in Iranian White Fresh Cheese. Iran J Sci Technol 4:75–84
Petros A, Maragkoudakis PA, Zoumpopoulou G, Miaris C, Kalantzopoulos G, Pot B, Tsakalidou E (2006) Probiotic potential of Lactobacillus strains isolated from dairy products. Int Dairy J 16:189–199
Pitt JI (2007) The genus Penicillium and its teleomorphic states Eupenicillium and Talaromyces. Academic, London and New York
Prodelalova J, Spanova A, Rittich B (2005) Application of PCR, rep-PCR and RAPD techniques for typing of Lactococcus lactis strains. Folia Microbiol 50:150–154
Rademaker JLW, Herbet H, Starrenburg MJC, Sabri M, Naser SN, Gevers D, Kelly WJ, Hugenholtz J, Swings J, van Hylckama Vlieg JE (2007) Diversity Analysis of Dairy and Nondairy Lactococcus lactis Isolates, Using a Novel Multilocus Sequence Analysis Scheme and (GTG)5-PCR Fingerprinting. Appl Environ Microbiol 73:7128–7137
Rossland E, Andersen Borge GI, Langsrud T, Sorhaug T (2003) Inhibition of Bacillus cereus by strains of Lactobacillus and Lactococcus in milk. Int J Food Microbiol 89:205–212
Rukure G, Bester BH (2001) Survival and growth of Bacillus cereus during Gouda cheese manufacturing. Food Control 12:31–36
Samson RA, Hoekstra ES, Frisvad JC, Filtenborg O (2000) Introduction to food- and airborne fungi. CVS, Utrecht
Scheirlinck I, Van der Meulen R, Van Schoor A, Vancanneyt M, De Vuyst L, Vandamme P, Huys G (2007) Influence of geographical origin and flour type on diversity of lactic acid bacteria in traditional Belgian sourdoughs. Appl Environ Microbiol 73:6262–6269
Sip A, Wieckowicz M, Olejnik-Schmidt A, Grajek W (2012) Anti-Listeria activity of lactic acid bacteria isolated from golka, a regional cheese produced in Poland. Food Control 26:117–124
Smaoui S, Elleuch L, Bejar W, Karray-Rehai I, Ayadi I, Jaouadi B, Mathieu F, Chouayekh H, Bejar S, Mellouli L (2010) Inhibition of fungi and gram-negative bacteria by bacteriocin BacTN635 produced by Lactobacillus plantarum sp. TN635. Appl Biochem Biotechnol 162:1132–1146
Stoyanova LG, Ustyugova EA, Sultimova TD, Bilanenko EN, Fedorova GB, Khatrukha GS, Netrusov AI (2010) New Antifungal Bacteriocin-Synthesizing Strains of Lactococcus lactis ssp. lactis as the Perspective Biopreservatives for Protection of Raw Smoked Sausages. Am J Agric Biol Sci 5:477–485
Ström K, Sjögren J, Broberg A, Schnürer J (2002) Lactobacillus plantarum MiLAB 393 produces the antifungal cyclic dipeptides cyclo(L-Phe-L-Pro) and nL-Phe-trans-4-OH-L-Pro) and 3-henyllactic Acid. Appl Environ Microb 68:4322–4327
Suomalainen TH, Mayra-Makinen AM (1999) Propionic acid bacteria as protective cultures in fermented milks and breads. Lait 79:165–174
Svec P, Sedlacek I, Zackova L, Novakova D, Kukletova M (2009) Lactobacillus spp. associated with early childhood caries. Folia Microbiol 54:53–58
Svec P, Sedlacek I, Chrapava M, Vandamme P (2011) (GTG)5-PCR fingerprinting of lactobacilli isolated from cervix of healthy women. Folia Microbiol 56:80–83
Svec P, Novakova D, Zackova L, Kukletova M, Sedlacek I (2008) Evaluation of (GTG)5-PCR for rapid identification of Streptococcus mutants. Antonie Van Leeuwenhoek 94:573–579
Svec P, Sedlacek I (2008) Characterization of Lactococcus lactis subsp. lactis isolated from surface waters. Folia Microbiol 53:53–56
Tadesse G, Ephraim E, Ashenafi M (2008) Assesment of the antimicrobial activity of lactic acid bacteria isolated from Borde and Shamita, traditional Ethiopian fermented beverages, on some food-borne pathogens and effect of growth medium on the inhibitory activity. J Food Saf 5:13–20
Todorov SD (2008) Bacteriocin production by Lactobacillus plantarum AMA-K isolated from Amasi, a Zimbabwean fermented milk product and study of adsorption of bacteriocin AMA-K to Listeria spp. Braz J Microbiol 38:178–187
Valerio F, Favilla M, De Bellis P, Sisto A, De Candia S, Lavermicocca P (2009) Antifungal activity of strains of lactic acid bacteria isolated from semolina ecosystem against Penicillium roqueforti, Aspergillus niger and Endomyces fibuliger contaminating bakery products. Syst Appl Microbiol 32:438–448
Vescovo M, Scolari G, Zacconi C (2006) Inhibition of Listeria innocua growth by antimicrobial-producing lactic acid cultures in vacuum-packed cold-smoked salmon. Food Microbiol 23:689–693
Watanabe T (2002) Pictorial atlas of soil and seed fungi: morfphologies of cultured fungi and key to species. CRC Press, Washington DC
Yang E, Fan L, Jiang Y, Doucette C, Fillmore S (2012) Antimicrobial activity of bacteriocin-producing lactic acid bacteria isolated from cheeses and yogurts. AMB Express 2:1–12
Acknowledgments
We greatly acknowledge financial support from the Research Council of Lithuania, grant SVE-05/2012 PIENRUGBAKT
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Šalomskienė, J., Abraitienė, A., Jonkuvienė, D. et al. Selection of enhanced antimicrobial activity posing lactic acid bacteria characterised by (GTG)5-PCR fingerprinting. J Food Sci Technol 52, 4124–4134 (2015). https://doi.org/10.1007/s13197-014-1512-6
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13197-014-1512-6