Abstract
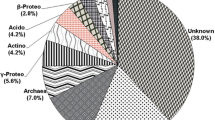
In this study we performed the phylogenetic analysis of non-cultivable bacteria from anthropogenically disturbed soil using partial sequences of the 16S rRNA (16S rDNA) and the heavy-metal resistance genes. This soil sample contained high concentrations of nickel (2,109 mg/kg), cobalt (355 mg/kg) and zinc (177 mg/kg), smaller concentrations of iron (35.75 mg/kg) and copper (32.2 mg/kg), and also a trace amount of cadmium (<0.25 mg/kg). The 16S rDNA sequences from a total of 74 bacterial clones were distributed into four broad taxonomic groups, Acidobacteria, Actinobacteria, Bacteroidetes and Gemmatimonadetes, and some of them were unidentified. Comparing our clone sequences with those from the GenBank database, only 9 clones displayed high similarity to known bacteria belongig to actinomycetes; others were identified as uncultured ones. Among clones evidently Actinobacteria predominated. Sixteen clones from soil sample carried only the nccA-like heavy-metal-resistance genes and all sequences showed too low similarity to known proteins encoded by these genes. However, our results suggested that the heavy-metal-contaminated soil is able to present very important reservoir of the new and until now unknown partly bacteria, partly heavy-metal-resistance determinants and their products. Bacteria and nccA-like genes identified in this study could represent the objects of interest as bioremediation agents because they can be potentially used in different transformation and immobilization processes.
Similar content being viewed by others
Abbreviations
- bp:
-
base pair
- czcA:
-
heavy-metal-resistance determinant
- nccA:
-
heavy-metal-resistance determinant
- PCR:
-
polymerase chain reaction
References
Ashraf R. & Ali T.A. 2007. Effect of heavy metals on soil microbial community and mung beans seed germination. Pak. J. Bot. 39: 629–636.
Bamborough L. & Cummings S.P. 2009. The impact of increasing heavy metal stress on the diversity and structure of the bacterial and actinobacterial communities of metallophytic grassland soil. Biol. Fertil. Soils 45: 273–280.
Becker J.M., Parkin T., Nakatsu C.H., Wilbur J.D. & Konopka A. 2006. Bacterial activity, community structure, and centimetre-scale spatial heterogeneity in contaminated soil. Microb. Ecol. 51: 220–231.
Bogdanova E.S., Bass I.A., Minakhin L.S., Petrova M.A., Mindlin S.Z., Volodin A.A., Kalyaeva E.S., Tiedje J.M., Hobman J.L., Brown N.L. & Nikiforov V.G. 1998. Horizontal spread of mer operons among Gram-positive bacteria in natural environments. Microbiology 144: 609–620.
Branco R., Chung A.P., Verissimo A. & Morais P.V. 2005. Impact of chromium-contaminated wastewaters on the microbial community of a river. FEMS Microbiol. Ecol. 54: 35–46.
Brim H., Heyndrickx M., de Vos P., Wilmotte A., Springael D., Schlegel H.G. & Mergeay M. 1999. Amplified rDNA restriction analysis and further genotypic characterisation of metalresistant soil bacteria and related facultative hydrogenotrophs. Syst. Appl. Microbiol. 22: 258–268.
Chien C., Kuo Y., Chen C., Hung C., Yeh C. & Yeh W. 2008. Microbial diversity of soil bacteria in agricultural field contaminated with heavy metals. J. Environ. Sci. 20: 359–363.
Eichorst S.A., Breznak J.A., Thomas M. & Schmidt T.M. 2007. Isolation and characterization of soil bacteria that define Terriglobus gen. nov., in the phylum Acidobacteria. Appl. Environ. Microbiol. 73: 2708–2717.
Ellis R.J., Morgan P., Weightman A.J. & Fry J.C 2003. Cultivation-dependent and -independent approaches for determining bacterial diversity in heavy-metal-contaminated soil. Appl. Environ. Microbiol. 69: 3223–3230.
Erbe J.L., Taylor K.B. & Hall L.M. 1995. Metalloregulation of the cyanobacterial smt locus: identification of SmtB binding sites and direct interaction with metals. Nucleic Acids Res. 23: 2472–2478.
Frey B., Stemmer M., Widmer F., Luster J.C. & Sperisen C. 2006. Microbial activity and community structure of a soil after heavy metal contamination in a model forest ecosystem. Soil Biol. Biochem. 38: 1745–1756.
Gremion F., Chatzinotas A. & Harms H. 2003. Comparative 16S rDNA and 16S rRNA sequence analysis indicates that Actinobacteria might be a dominant part of the metabolically active bacteria in heavy metal-contaminated bulk and rhizosphere soil. Environ. Microbiol. 5: 896–907.
Holmes A.J., Bowyer J., Holley M.P., O’Donoghue M., Montgomery M. & Gillings M.R. 2000. Diverse, yet-to-be-cultured members of the Rubrobacter subdivision of the Actinobacteria are widespread in Australian arid soils. FEMS Microbiol. Ecol. 33: 111–120.
Hugenholtz P. 2002. Exploring prokaryotic diversity in the genomic era. Genome Biol. 3: reviewsS0003.
Ji G. & Silver S. 1992. Regulation and expression of the arsenic resistance operon from Staphylococcus aureus plasmid pI258. J. Bacteriol. 174: 3684–3694.
Kandeler E., Tscherko D., Bruce K.D., Stemmer M., Hobbs P.J., Bardgett R.D. & Amelung W. 2000. Structure and function of the soil microbial community in microhabitats of a heavy metal polluted soil. Biol. Fertil. Soils 32: 390–400.
Karelová E., Harichová J., Stojnev T., Pangallo D. & Ferianc P. 2011. The isolation of heavy-metal resistant culturable bacteria and resistance determinants from a heavy-metalcontaminated site. Biologia 66: 18–26.
Kersters K., de Vos P., Gillis M., Swings J., Vandamme P. & Stackebrandt E. 2006. Introduction to the Proteobacteria, pp. 3–37. In: Dwarkin M., Falkow S., Rosenberg E., Schleifer K.H. & Stackebrandt E. (eds), The Prokaryotes, 3rd Edn, vol. 5, Springer, New York.
Khan S., Hesham A.E.L., Qiao M., Rehman S. & He J.Z. 2010. Effect of Cd and Pb on soil microbial community structure and activities. Environ. Sci. Polut. Res. 17: 288–296.
Kimura M. 1980. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16: 111–120.
Kozdroj J. & van Elsas J.D. 2001. Structural diversity of microorganisms in chemically perturbed soil assessed by molecular and cytochemical approaches. J. Microbiol. Methods 43: 197–212.
Lane D.J. 1991. 16S/23S rRNA sequencing, pp. 115–148. In: Stackebrandt E. & Goodfellow M. (eds), Nucleic Acid Techniques in Bacterial Systematics, John Wiley & Sons, New York.
Liu H.Y., Probs A. & Liao B. 2005. Metal contamination of soils and crops affected by the Chenzhou lead/zinc mine spill (Hunan, China). Sci. Total Environ. 339: 153–166.
Mergeay M., Monchy S., Vallaeys T., Auquier V., Benotmane A., Bertin P., Taghavi S., Dunn J., Van Der Lelie D. & Wattiez R. 2003. Ralstonia metallidurans, a bacterium specifically adapted to toxic metals: towards a catalogue of metal-responsive genes. FEMS Microbiol. Rev. 27: 385–410.
Mobley H.L., Chen C.M., Silver S. & Rosen B.P. 1983. Cloning and expression of R-factor mediated arsenate resistance in Escherichia coli. Mol. Gen. Genet. 191: 421–426.
Nascimento A.M.A. & Chartone-Souza E. 2003. Operon mer: bacterial resistance to mercury and potential for bioremediation of contaminated environments. Genet. Mol. Res. 2: 92–101.
Nies D.H. 1999. Microbial heavy-metal resistance. Appl. Microbiol. Biotechnol. 51: 730–750.
Ogilvie L.A. & Grant A. 2008. Linking pollution induced community tolerance (PICT) and microbial community structure in chronically metal polluted estuarine sediments. Mar. Environ. Res. 65: 187–198.
Page R.D.M. 1996. TreeView: an application to display phylogenetic trees on personal computers. Comput. Applic. Biosci. 12: 357–358.
Pazirandeh M., Wells B. & Ryan R.L. 1998. Development of bacterium-based heavy metal biosorbents: enhanced uptake of cadmium and mercury by Escherichia coli expressing a metal binding motif. Appl. Environ. Microbiol. 64: 4068–4072.
Pechrada J., Sajjaphan K. & Sadowsky M.J. 2010. Structure and diversity of arsenic-resistant bacteria in an old tin mine area of Thailand. J. Microbiol. Biotechnol. 20: 169–178.
Ranjard L., Brothier E. & Nazaret S. 2000. Sequencing bands of ribosomal intergenic spacer analysis fingerprints for characterization and microscale distribution of soil bacterium populations responding to mercury spiking. Appl. Environ. Microbiol. 66: 5334–5339.
Ranjard L., Lignier L. & Chaussod R. 2006. Cumulative effect of short-term polymetal contamination on soil bacterial community structure. Appl. Environ. Microbiol. 72: 1684–1687.
Rosenstein R., Peschel A., Wieland B. & Götz F. 1992. Expression and regulation of the antimonite, arsenite, and arsenate resistance operon of Staphylococcus xylosus plasmid pSX267. J. Bacteriol. 174: 3676–3683.
Saitou N. & Nei M. 1987. The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4: 406–425.
Saltikov C.W. & Olson B.H. 2002. Homology of Escherichia coli R773 arsA, arsB, and arsC genes in arsenic-resistant bacteria isolated from raw sewage and arsenic-enriched creek waters. Appl. Environ. Microbiol. 68: 280–288.
Sandaa R.A., Torsvik V. & Enger Ø. 2001. Influence of long-term heavy-metal contamination on microbial communities in soil. Soil Biol. Biochem. 33: 287–295.
Sandaa R.A., Torsvik V., Enger Ø., Daae F.L., Castberg T. & Hahn D. 2006. Analysis of bacterial communities in heavy metal-contaminated soils at different levels of resolution. FEMS Microbiol. Ecol. 30: 237–251.
Sandrin T.R., Chech A.M. & Maier R.M. 2000. A rhamnolipid biosurfactant reduces cadmium toxicity during biodegradation of naphthalene. Appl. Environ. Microbiol. 66: 4585–4588.
Schmidt T. & Schlegel H.G. 1994. Combined nickel-cobaltcadmium resistance encoded by the ncc locus of Alcaligenes xylosoxidans 31A. J. Bacteriol. 176: 7045–7054.
Silver S. & Phung L.T. 1996. Bacterial heavy metal resistance: new surprises. Annu. Rev. Microbiol. 50: 753–789.
Sobolev D. & Begonia M.F.T. 2008. Effects of heavy metal contamination upon soil microbes: lead-induced changes in general and denitrifying microbial communities as evidenced by molecular markers. Int. J. Environ. Res. Public Health 5: 450–456.
Sun L.N., Zhang Y.F., He L.Y., Chen Z.J., Wang Q.Y., Qian M. & Sheng X.F. 2010. Genetic diversity and characterization of heavy metal-resistant-endophytic bacteria from two copper-tolerant plant species on copper mine wasteland. Biores. Technol. 101: 501–509.
Štursa P., Uhlík O., Kurzawová V., Koubek J., Ionescu M., Strohalm M., Lovecká P., Macek T. & Macková M. 2009. Approaches for diversity analysis of cultivable and noncultivable bacteria in real soil. Plant Soil Environ. 55: 389–396.
Takaichi S., Maoka T., Takasaki K. & Hanada S. 2010. Carotenoids of Gemmatimonas aurantiaca (Gemmatimonadetes): identification of a novel carotenoid, deoxyoscillol 2-rhamnoside, and proposed biosynthetic pathway of oscillol 2,29-dirhamnoside. Microbiology 156: 757–763.
Thiyagarajan V., Lau S., Tsoi M., Zhang W. & Qian P.Y. 2010. Monitoring bacterial biodiversity in surface sediment using terminal restriction fragment length polymorphism analysis (T-RFLP): application to coastal environment, pp. 151–163. In: Ishimatsu A. & Lie H.J. (eds), Coastal Environmental and Ecosystem Issues of the East China Sea, Terrapub & Nagasaki University.
Vivas A., Moreno B., del Val C., Macci C., Masciandaroc G. & Benitez E. 2008. Metabolic and bacterial diversity in soils historically contaminated by heavy metals and hydrocarbons. J. Environ. Monit. 10: 1287–1296.
Wu H., Zhao H., Wen C., Guo Y., Guo J., Xu M. & Li X. 2012. A comparative study of bacterial community structures in the sediments from brominated flame retardants contaminated river and non-contaminated reservoir. Afr. J. Microbiol. Res. 6: 3248–3260.
Wuertz S. & Mergeay M. 1997. The impact of heavy metals on soil microbial communities and their activities, pp. 607–642. In: Van Elsas J.D., Trevors J.T. & Wellington E.M.H. (eds), Modern Soil Microbiology, Marcel Dekker, New York.
Zhang H., Sekiguchi Y., Hanada S., Hugenholtz P., Kim H., Kamagata Y. & Nakamura K. 2003. Gemmatimonas aurantiaca gen. nov., sp. nov., a Gram-negative, aerobic, polyphosphate-accumulating micro-organism, the first cultured representative of the new bacterial phylum Gemmatimonadetes phyl. nov. Int. J. Syst. Evol. Microbiol. 53: 1155–1163.
Zhang H.B., Yang M.X., Shi W., Zheng Y., Tao T. & Zhao Z.W. 2007. Bacterial diversity in mine tailings compared by cultivation and cultivation-independent methods and their resistance to lead and cadmium. Microbial. Ecol. 54: 705–712.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Harichová, J., Karelová, E., Pangallo, D. et al. Structure analysis of bacterial community and their heavy-metal resistance determinants in the heavy-metal-contaminated soil sample. Biologia 67, 1038–1048 (2012). https://doi.org/10.2478/s11756-012-0123-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11756-012-0123-9