Abstract
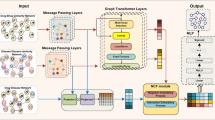
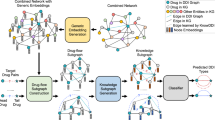
Drug–drug interactions (DDIs) for emerging drugs offer possibilities for treating and alleviating diseases, and accurately predicting these with computational methods can improve patient care and contribute to efficient drug development. However, many existing computational methods require large amounts of known DDI information, which is scarce for emerging drugs. Here we propose EmerGNN, a graph neural network that can effectively predict interactions for emerging drugs by leveraging the rich information in biomedical networks. EmerGNN learns pairwise representations of drugs by extracting the paths between drug pairs, propagating information from one drug to the other, and incorporating the relevant biomedical concepts on the paths. The edges of the biomedical network are weighted to indicate the relevance for the target DDI prediction. Overall, EmerGNN has higher accuracy than existing approaches in predicting interactions for emerging drugs and can identify the most relevant information on the biomedical network.





Similar content being viewed by others
Data availability
The resplit dataset35 in DrugBank, TWOSIDES and HetioNet for the S1 and S2 settings is publicly available at https://doi.org/10.5281/zenodo.10016715. Source data are provided with this paper.
Code availability
The code for EmerGNN36 is available at https://github.com/LARS-research/EmerGNN.
References
Su, X., Wang, H., Zhao, N., Wang, T. & Cui, Y. Trends in innovative drug development in China. Nat. Rev. Drug Discov. 21, 709–710 (2022).
Ledford, H. Hundreds of COVID trials could provide a deluge of new drugs. Nature 603, 25–27 (2022).
Percha, B. & Altman, R. B. Informatics confronts drug-drug interactions. Trends Pharmacol. Sci. 34, 178–184 (2013).
Vilar, S. et al. Similarity-based modeling in large-scale prediction of drug-drug interactions. Nat. Protoc. 9, 2147–2163 (2014).
Tanvir, F., Islam, M. I. K. & Akbas, E. Predicting drug-drug interactions using meta-path based similarities. In Proc. IEEE Conference on Computational Intelligence in Bioinformatics and Computational Biology (eds Hallinan, J. et al.) 1–8 (IEEE, 2021).
Yu, Y. et al. SumGNN: multi-typed drug interaction prediction via efficient knowledge graph summarization. Bioinformatics 37, 2988–2995 (2021).
Letinier, L. et al. Risk of drug–drug interactions in out-hospital drug dispensings in France: results from the drug–drug interaction prevalence study. Front. Pharmacol. 10, 265 (2019).
Jiang, H. et al. Adverse drug reactions and correlations with drug–drug interactions: a retrospective study of reports from 2011 to 2020. Front. Pharmacol. 13, 923939 (2022).
Rogers, D. & Hahn, M. Extended-connectivity fingerprints. J. Chem. Inf. Model. 50, 742–754 (2010).
Dewulf, P., Stock, M. & De Baets, B. Cold-start problems in data-driven prediction of drug-drug interaction effects. Pharmaceuticals 14, 429 (2021).
Liu, Z., Wang, X.-N., Yu, H., Shi, J.-Y. & Dong, W.-M. Predict multi-type drug-drug interactions in cold start scenario. BMC Bioinformatics 23, 75 (2022).
Yao, J., Sun, W., Jian, Z., Wu, Q. & Wang, X. Effective knowledge graph embeddings based on multidirectional semantics relations for polypharmacy side effects prediction. Bioinformatics 38, 2315–2322 (2022).
Zitnik, M., Agrawal, M. & Leskovec, J. Modeling polypharmacy side effects with graph convolutional networks. Bioinformatics 34, i457–i466 (2018).
Karim, M. R. et al. Drug–drug interaction prediction based on knowledge graph embeddings and convolutional-LSTM network. In Proc. 10th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics (eds Shi, X. & Buck, M.) 113–123 (Association for Computing Machinery, 2019).
Huang, K., Xiao, C., Glass, L. M., Zitnik, M. & Sun, J. SkipGNN: predicting molecular interactions with skip-graph networks. Sci. Rep. 10, 21092 (2020).
Lin, X., Quan, Z., Wang, Z.-J., Ma, T. & Zeng, X. KGNN: knowledge graph neural network for drug-drug interaction prediction. In Proc. Twenty-Ninth International Joint Conference on Artificial Intelligence (ed. Bessiere, C.) 2739–2745 (IJCAI, 2020).
Ren, Z.-H. et al. A biomedical knowledge graph-based method for drug-drug interactions prediction through combining local and global features with deep neural networks. Brief. Bioinformatics 23, bbac363 (2022).
Himmelstein, D. S. et al. Systematic integration of biomedical knowledge prioritizes drugs for repurposing. eLife 6, e26726 (2017).
Kipf, T. N. & Welling, M. Semi-supervised classification with graph convolutional networks. In Proc. 5th International Conference on Learning Representations https://openreview.net/forum?id=SJU4ayYgl (OpenReview.net, 2017).
Gilmer, J., Schoenholz, S. S., Riley, P. F., Vinyals, O. & Dahl, G. E. Neural message passing for quantum chemistry. In International Conference on Machine Learning (eds Precup, D. & Teh, Y. W.) 1263–1272 (Association for Computing Machinery, 2017).
Yu, H., Zhao, S. Y. & Shi, J. Y. STNN-DDI: a substructure-aware tensor neural network to predict drug-drug interactions. Brief. Bioinformatics 23, bbac209 (2022).
Wishart, D. S. et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 46, D1074–D1082 (2018).
Tatonetti, N. P., Ye, P. P., Daneshjou, R. & Altman, R. B. Data-driven prediction of drug effects and interactions. Sci. Transl. Med. 4, 125ra31–125ra31 (2012).
Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 20, 37–46 (1960).
Brown, D. G., Wobst, H. J., Kapoor, A., Kenna, L. A. & Southall, N. Clinical development times for innovative drugs. Nat. Rev. Drug Discov. 21, 793–794 (2021).
Liu, M. & Wittbrodt, E. Low-dose oral naloxone reverses opioid-induced constipation and analgesia. J. Pain Symptom Manag. 23, 48–53 (2002).
Estabrook, R. W. A passion for P450s (remembrances of the early history of research on cytochrome P450). Drug Metab. Dispos. 31, 1461–1473 (2003).
Vashishth, S., Sanyal, S., Nitin, V. & Talukdar, P. Composition-based multi-relational graph convolutional networks. In Proc. 8th International Conference on Learning Representations https://openreview.net/pdf?id=BylA_C4tPr (OpenReview.net, 2020).
Lao, N., Mitchell, T. & Cohen, W. Random walk inference and learning in a large scale knowledge base. In Proc. 2011 Conference on Empirical Methods in Natural Language Processing (eds Merlo, P. & Barzilay, R.) 529–539 (Association for Computing Machinery, 2011).
Xiong, W., Hoang, T. & Wang, W. Y. DeepPath: a reinforcement learning method for knowledge graph reasoning. In Proc. 2017 Conference on Empirical Methods in Natural Language Processing (eds Specia, L. et al.) 564–573 (Association for Computational Linguistics, 2017).
Zhang, M. & Chen, Y. Link prediction based on graph neural networks. In Proc. 32nd International Conference on Neural Information Processing Systems (eds Bengio, S. & Wallach, H. M.) 5171–5181 (Association for Computing Machinery, 2018).
Teru, K., Denis, E. & Hamilton, W. Inductive relation prediction by subgraph reasoning. In International Conference on Machine Learning (eds Daumé III, H. & Singh, A.) 9448–9457 (Association for Computing Machinery, 2020).
Nair, V. & Hinton, G. E. Rectified linear units improve restricted Boltzmann machines. In Proc. 27th International Conference on Machine Learning (eds Joachims, T. & Furnkranz, J.) 807–814 (Association for Computing Machinery, 2010).
Kingma, D. P & Ba, J. Adam: a method for stochastic optimization. In Proc. 3rd International Conference on Learning Representations (eds Bengio, Y. & LeCun, Y.) https://arxiv.org/pdf/1412.6980.pdf (2014).
Zhang, Y., Yue, L. & Yao, Q. EmerGNN_DDI_data. Zenodo https://doi.org/10.5281/zenodo.10016715 (2023).
Zhang, Y., Yue, L. & Yao, Q. LARS-research/EmerGNN: v1.0.0k. Zenodo https://doi.org/10.5281/zenodo.10017431 (2023).
Van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Acknowledgements
This project was supported by the National Natural Science Foundation of China (no. 92270106) and the CCF-Tencent Open Research Fund.
Author information
Authors and Affiliations
Contributions
Y. Zhang contributed to idea development, algorithm implementation, experimental design, results analysis and writing of the paper. Q.Y. contributed to idea development, experimental design, results analysis and writing of the paper. L.Y. contributed to algorithm implementation and results analysis. Y. Zheng contributed to results analysis and writing of the paper. All authors read, edited and approved the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Computational Science thanks Nguyen Quoc Khanh Le, Jian-Yu Shi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Kaitlin McCardle, in collaboration with the Nature Computational Science team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Sections 1 and 2, Tables 1–7 and Figs. 1–5.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Y., Yao, Q., Yue, L. et al. Emerging drug interaction prediction enabled by a flow-based graph neural network with biomedical network. Nat Comput Sci 3, 1023–1033 (2023). https://doi.org/10.1038/s43588-023-00558-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-023-00558-4
- Springer Nature America, Inc.