Abstract
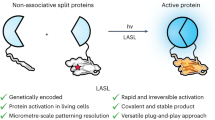
The conditional assembly of split-protein pairs to modulate biological activity is commonly achieved by fusing split-protein fragments to dimerizing components that bring inactive pairs into close proximity in response to an exogenous trigger. However, current methods lack full spatial and temporal control over reconstitution, require sustained activation and lack specificity. Here light-activated SpyLigation (LASL), based on the photoregulation of the covalent SpyTag (ST)/SpyCatcher (SC) peptide–protein reaction, assembles nonfunctional split fragment pairs rapidly and irreversibly in solution, in engineered biomaterials and intracellularly. LASL introduces an ortho-nitrobenzyl(oNB)-caged lysine into SC’s reactive site to generate a photoactivatable SC (pSC). Split-protein pairs of interest fused to pSC and ST are conditionally assembled via near-ultraviolet or pulsed near-infrared irradiation, as the uncaged SC can react with ST to ligate appended fragments. We describe procedures for the efficient synthesis of the photocaged amino acid that is incorporated within pSC (<5 days) as well as the design and cloning of LASL plasmids (1–4 days) for recombinant protein expression in either Escherichia coli (5–6 days) or mammalian cells (4–6 days), which require some prior expertise in protein engineering. We provide a chemoenzymatic scheme for appending bioorthogonal reactive handles onto E. coli-purified pSC protein (<4 days) that permits LASL component incorporation and patterned protein activation within many common biomaterial platforms. Given that LASL is irreversible, the photolithographic patterning procedures are fast and do not require sustained light exposure. Overall, LASL can be used to interrogate and modulate cell signaling in various settings.
Key points
-
The procedures cover a synthetic route to generate the photocaged lysine for photocaged SpyCatcher expression, the design of light-activated SpyLigation split proteins and plasmid vector design for expression in bacterial or mammalian cells.
-
The light-activated SpyLigation recombinant proteins are expressed and tested in solution, tethered into engineered biomaterials by site-specifically appending biorthogonal reactive handles on photocaged SpyCatcher through a sortase-mediated reaction, and transfected into mammalian cells for intracellular protein localization/activation via photolithographic patterning.









Similar content being viewed by others
Data availability
References
Shekhawat, S. S. & Ghosh, I. Split-protein systems: beyond binary protein–protein interactions. Curr. Opin. Chem. Biol. 15, 789–797 (2011).
Michnick, S. W., Ear, P. H., Manderson, E. N., Remy, I. & Stefan, E. Universal strategies in research and drug discovery based on protein-fragment complementation assays. Nat. Rev. Drug Discov. 6, 569–582 (2007).
Fink, T. et al. Design of fast proteolysis-based signaling and logic circuits in mammalian cells. Nat. Chem. Biol. 15, 115–122 (2019).
Rihtar, E. et al. Chemically inducible split protein regulators for mammalian cells. Nat. Chem. Biol. 19, 64–71 (2023).
Gao, Y. et al. Complex transcriptional modulation with orthogonal and inducible dCas9 regulators. Nat. Methods 13, 1043–1049 (2016).
Farrar, M. A., Alberola-lla, J. & Perlmutter, R. M. Activation of the Raf-1 kinase cascade by coumermycin-induced dimerization. Nature 383, 178–181 (1996).
Spencer, D. M., Wandless, T. J., Schreiber, S. L. & Crabtree, G. R. Controlling signal transduction with synthetic ligands. Science 262, 1019–1024 (1993).
Kawano, F., Suzuki, H., Furuya, A. & Sato, M. Engineered pairs of distinct photoswitches for optogenetic control of cellular proteins. Nat. Commun. 6, 6256 (2015).
Levskaya, A., Weiner, O. D., Lim, W. A. & Voigt, C. A. Spatiotemporal control of cell signalling using a light-switchable protein interaction. Nature 461, 997–1001 (2009).
Kennedy, M. J. et al. Rapid blue-light–mediated induction of protein interactions in living cells. Nat. Methods 7, 973–975 (2010).
Ruskowitz, E. R. et al. Spatiotemporal functional assembly of split protein pairs through a light-activated SpyLigation. Nat. Chem. 15, 694–704 (2023).
Zakeri, B. et al. Peptide tag forming a rapid covalent bond to a protein, through engineering a bacterial adhesin. Proc. Natl Acad. Sci. USA 109, E690–7 (2012).
Keeble, A. H. & Howarth, M. Power to the protein: enhancing and combining activities using the Spy toolbox. Chem. Sci. 11, 7281–7291 (2020).
Keeble, A. H. & Howarth, M. in Methods in Enzymology vol. 617 443–461 (Elsevier, 2019).
Hoorens, M. W. H. & Szymanski, W. Reversible, spatial and temporal control over protein activity using light. Trends Biochem. Sci. 43, 567–575 (2018).
Ruskowitz, E. R. & DeForest, C. A. Proteome-wide analysis of cellular response to ultraviolet light for biomaterial synthesis and modification. ACS Biomater. Sci. Eng. 5, 2111–2116 (2019).
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Chen, P. R. et al. A facile system for encoding unnatural amino acids in mammalian cells. Angew. Chem. Int. Ed. 48, 4052–4055 (2009).
Young, T. S., Ahmad, I., Yin, J. A. & Schultz, P. G. An enhanced system for unnatural amino acid mutagenesis in E. coli. J. Mol. Biol. 395, 361–374 (2010).
Yanagisawa, T. et al. Multistep engineering of Pyrrolysyl–tRNA synthetase to genetically encode Nɛ-(o-Azidobenzyloxycarbonyl) lysine for site-specific protein modification. Chem. Biol. 15, 1187–1197 (2008).
Serfling, R. et al. Designer tRNAs for efficient incorporation of non-canonical amino acids by the pyrrolysine system in mammalian cells. Nucleic Acids Res. 46, 1–10 (2018).
To, T.-L., Zhang, Q. & Shu, X. Structure-guided design of a reversible fluorogenic reporter of protein-protein interactions: design of a reversible fluorogenic reporter of PPIs. Protein Sci. 25, 748–753 (2016).
Hall, M. P. et al. Engineered luciferase reporter from a deep sea shrimp utilizing a novel imidazopyrazinone substrate. ACS Chem. Biol. 7, 1848–1857 (2012).
Guimaraes, C. P. et al. Site-specific C-terminal and internal loop labeling of proteins using sortase-mediated reactions. Nat. Protoc. 8, 1787–1799 (2013).
Mao, H., Hart, S. A., Schink, A. & Pollok, B. A. Sortase-mediated protein ligation: a new method for protein engineering. J. Am. Chem. Soc. 126, 2670–2671 (2004).
Shadish, J. A., Benuska, G. M. & DeForest, C. A. Bioactive site-specifically modified proteins for 4D patterning of gel biomaterials. Nat. Mater. 18, 1005–1014 (2019).
Gawade, P. M., Shadish, J. A., Badeau, B. A. & DeForest, C. A. Logic‐based delivery of site‐ specifically modified proteins from environmentally responsive hydrogel biomaterials. Adv. Mater. 31, 1902462 (2019).
Batalov, I., Stevens, K. R. & DeForest, C. A. Photopatterned biomolecule immobilization to guide three-dimensional cell fate in natural protein-based hydrogels. Proc. Natl Acad. Sci. USA 118, e2014194118 (2021).
Agard, N. J., Prescher, J. A. & Bertozzi, C. R. A strain-promoted [3 + 2] azide−alkyne cycloaddition for covalent modification of biomolecules in living systems. J. Am. Chem. Soc. 126, 15046–15047 (2004).
DeForest, C. A., Polizzotti, B. D. & Anseth, K. S. Sequential click reactions for synthesizing and patterning three-dimensional cell microenvironments. Nat. Mater. 8, 659–664 (2009).
Arakawa, C. K., Badeau, B. A., Zheng, Y. & DeForest, C. A. Multicellular vascularized engineered tissues through user‐programmable biomaterial photodegradation. Adv. Mater. 29, 1703156 (2017).
Badeau, B. A., Comerford, M. P., Arakawa, C. K., Shadish, J. A. & DeForest, C. A. Engineered modular biomaterial logic gates for environmentally triggered therapeutic delivery. Nat. Chem. 10, 251–258 (2018).
Szymczak, A. L. et al. Correction of multi-gene deficiency in vivo using a single ‘self-cleaving’ 2A peptide–based retroviral vector. Nat. Biotechnol. 22, 589–594 (2004).
Roberts, P. J. et al. Rho family GTPase modification and dependence on CAAX motif-signaled posttranslational modification. J. Biol. Chem. 283, 25150–25163 (2008).
Keeble, A. H. et al. Approaching infinite affinity through engineering of peptide–protein interaction. Proc. Natl Acad. Sci. USA 116, 26523–26533 (2019).
Keeble, A. H. et al. DogCatcher allows loop-friendly protein–protein ligation. Cell Chem. Biol. 29, 339–350.e10 (2022).
Wu, X.-L., Liu, Y., Liu, D., Sun, F. & Zhang, W.-B. An intrinsically disordered peptide-peptide stapler for highly efficient protein ligation both in vivo and in vitro. J. Am. Chem. Soc. 140, 17474–17483 (2018).
Fierer, J. O., Veggiani, G. & Howarth, M. SpyLigase peptide–peptide ligation polymerizes affibodies to enhance magnetic cancer cell capture. Proc. Natl Acad. Sci. USA 111, E1176–E1181 (2014).
Buldun, C. M., Jean, J. X., Bedford, M. R. & Howarth, M. Snoopligase catalyzes peptide– peptide locking and enables solid-phase conjugate isolation. J. Am. Chem. Soc. 140, 3008–3018 (2018).
Lorand, L. & Graham, R. M. Transglutaminases: crosslinking enzymes with pleiotropic functions. Nat. Rev. Mol. Cell Biol. 4, 140–156 (2003).
Fegan, A., White, B., Carlson, J. C. T. & Wagner, C. R. Chemically controlled protein assembly: techniques and applications. Chem. Rev. 110, 3315–3336 (2010).
Sun, J. & Sadelain, M. The quest for spatio-temporal control of CAR T cells. Cell Res. 25, 1281–1282 (2015).
Barlow, A. D., Nicholson, M. L. & Herbert, T. P. Evidence for rapamycin toxicity in pancreatic β-cells and a review of the underlying molecular mechanisms. Diabetes 62, 2674–2682 (2013).
Hartzell, E. J., Terr, J. & Chen, W. Engineering a Blue Light Inducible SpyTag System (BLISS). J. Am. Chem. Soc. 143, 8572–8577 (2021).
Wong, S., Mosabbir, A. A. & Truong, K. An engineered split intein for photoactivated protein trans-splicing. PLoS ONE 10, e0135965 (2015).
Cadet, J., Sage, E. & Douki, T. Ultraviolet radiation-mediated damage to cellular DNA. Mutat. Res. Mol. Mech. Mutagen. 571, 3–17 (2005).
Lawrence, K. P. et al. The UV/visible radiation boundary region (385–405 nm) damages skin cells and induces “dark” cyclobutane pyrimidine dimers in human skin in vivo. Sci. Rep. 8, 12722 (2018).
Kozmin, S. et al. UVA radiation is highly mutagenic in cells that are unable to repair 7,8-dihydro-8-oxoguanine in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 102, 13538–13543 (2005).
Zhang, W. et al. Optogenetic control with a photocleavable protein, PhoCl. Nat. Methods 14, 391–394 (2017).
Coin, I., Beyermann, M. & Bienert, M. Solid-phase peptide synthesis: from standard procedures to the synthesis of difficult sequences. Nat. Protoc. 2, 3247–3256 (2007).
Kawada, Y., Kodama, T., Miyashita, K., Imanishi, T. & Obika, S. Synthesis and evaluation of novel caged DNA alkylating agents bearing 3,4-epoxypiperidine structure. Org. Biomol. Chem. 10, 5102 (2012).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
Dolberg, T. B. et al. Computation-guided optimization of split protein systems. Nat. Chem. Biol. 17, 531–539 (2021).
Chen, X., Zaro, J. L. & Shen, W.-C. Fusion protein linkers: property, design and functionality. Adv. Drug Deliv. Rev. 65, 1357–1369 (2013).
Phillips-Jones, M. K., Hill, L. S., Atkinson, J. & Martin, R. Context effects on misreading and suppression at UAG codons in human cells. Mol. Cell. Biol. 15, 6593–6600 (1995).
Beier, H. Misreading of termination codons in eukaryotes by natural nonsense suppressor tRNAs. Nucleic Acids Res. 29, 4767–4782 (2001).
Shadish, J. A. & DeForest, C. A. Site-selective protein modification: from functionalized proteins to functional biomaterials. Matter 2, 50–77 (2020).
Stoien, J. D. & Wang, R. J. Effect of near-ultraviolet and visible light on mammalian cells in culture. II. Formation of toxic photoproducts in tissue culture medium by blacklight. Proc. Natl Acad. Sci. USA 71, 3961–3965 (1974).
DeForest, C. A. & Tirrell, D. A. A photoreversible protein-patterning approach for guiding stem cell fate in three-dimensional gels. Nat. Mater. 14, 523–531 (2015).
Acknowledgements
The authors recognize C. Butcher, T. Rapp, R. Francis, I. Kopyeva, R. Bretherton, M. Ross and N. Gregorio for feedback on the paper, as well as E. Ruskowitz, A. Strange, C. Butcher, S. Kurniawan and J. Filteau who further contributed to the original LASL manuscript. This work was supported by a CAREER Award (DMR 1652141 to C.A.D.) and grants (DMR 1807398 and CBET 1803054 to C.A.D.) from the National Science Foundation, as well as a Maximizing Investigators’ Research Award (R35GM138036 to C.A.D.) from the National Institutes of Health. Student fellowship support was provided by the Institute for Stem Cell and Regenerative Medicine (to B.G.M.-R.)
Author information
Authors and Affiliations
Contributions
Each author contributed equally in the preparation of this Protocol.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Cecile Echalier, Tae-Young Yoon and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Ruskowitz, E. R. et al. Nat. Chem. 15, 694–704 (2023): https://doi.org/10.1038/s41557-023-01152-x
Shadish, J. A. et al. Nat. Mater. 18, 1005–1014 (2019): https://doi.org/10.1038/s41563-019-0367-7
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Munoz-Robles, B.G., DeForest, C.A. Irreversible light-activated SpyLigation mediates split-protein assembly in 4D. Nat Protoc 19, 1015–1052 (2024). https://doi.org/10.1038/s41596-023-00938-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00938-0
- Springer Nature Limited