Abstract
Proper neural progenitor behavior in conjunction with orderly vasculature formation is fundamental to the development of the neocortex. However, the mechanisms coordinating neural progenitor behavior and vessel growth remain largely elusive. Here we show that robust metabolic production of lactate by radial glial progenitors (RGPs) co-regulates vascular development and RGP division behavior in the developing mouse neocortex. RGPs undergo a highly organized lineage progression program to produce diverse neural progeny. Systematic single-cell metabolic state analysis revealed that RGPs and their progeny exhibit distinct metabolic features associated with specific cell types and lineage progression statuses. Symmetrically dividing, proliferative RGPs preferentially express a cohort of genes that support glucose uptake and anaerobic glycolysis. Consequently, they consume glucose in anaerobic metabolism and produce a high level of lactate, which promotes vessel growth. Moreover, lactate production enhances RGP proliferation by maintaining mitochondrial length. Together, these results suggest that specific metabolic states and metabolites coordinately regulate vasculature formation and progenitor behavior in neocortical development.







Similar content being viewed by others
Data availability
The datasets generated and/or analyzed in this study are available upon reasonable request. Bulk RNA sequencing data are deposited at the Gene Expression Omnibus under accession number GSE201991. The matrix of metabolic gene expression used for single-cell metabolic state analysis is provided in Supplementary Data 1. Source data are provided with this paper.
Code availability
No code or model was generated in this study.
References
Paredes, I., Himmels, P. & de Almodovar, C. R. Neurovascular communication during CNS development. Dev. Cell 45, 10–32 (2018).
Andreone, B. J., Lacoste, B. & Gu, C. Neuronal and vascular interactions. Annu. Rev. Neurosci. 38, 25–46 (2015).
Ma, S., Kwon, H. J., Johng, H., Zang, K. & Huang, Z. Radial glial neural progenitors regulate nascent brain vascular network stabilization via inhibition of Wnt signaling. PLoS Biol. 11, e1001469 (2013).
Stubbs, D. et al. Neurovascular congruence during cerebral cortical development. Cereb. Cortex 19, i32–i41 (2009).
Gao, P. et al. Deterministic progenitor behavior and unitary production of neurons in the neocortex. Cell 159, 775–788 (2014).
Kriegstein, A. & Alvarez-Buylla, A. The glial nature of embryonic and adult neural stem cells. Annu. Rev. Neurosci. 32, 149–184 (2009).
Gotz, M. & Huttner, W. B. The cell biology of neurogenesis. Nat. Rev. Mol. Cell Biol. 6, 777–788 (2005).
Homem, C. C., Repic, M. & Knoblich, J. A. Proliferation control in neural stem and progenitor cells. Nat. Rev. Neurosci. 16, 647–659 (2015).
Lin, Y. et al. Behavior and lineage progression of neural progenitors in the mammalian cortex. Curr. Opin. Neurobiol. 66, 144–157 (2021).
Angevine, J. B. & Sidman, R. L. Autoradiographic study of cell migration during histogenesis of cerebral cortex in mouse. Nature 192, 766–768 (1961).
Shen, Z. et al. Distinct progenitor behavior underlying neocortical gliogenesis related to tumorigenesis. Cell Rep. 34, 108853 (2021).
Clavreul, S. et al. Cortical astrocytes develop in a plastic manner at both clonal and cellular levels. Nat. Commun. 10, 4884 (2019).
Berg, D. A. et al. A common embryonic origin of stem cells drives developmental and adult neurogenesis. Cell 177, 654–668 (2019).
Vasudevan, A., Long, J. E., Crandall, J. E., Rubenstein, J. L. R. & Bhide, P. G. Compartment-specific transcription factors orchestrate angiogenesis gradients in the embryonic brain. Nat. Neurosci. 11, 429–439 (2008).
Tata, M., Ruhrberg, C. & Fantin, A. Vascularisation of the central nervous system. Mech. Dev. 138, 26–36 (2015).
Namba, T., Nardelli, J., Gressens, P. & Huttner, W. B. Metabolic regulation of neocortical expansion in development and evolution. Neuron 109, 408–419 (2021).
Xiao, Z., Dai, Z. & Locasale, J. W. Metabolic landscape of the tumor microenvironment at single cell resolution. Nat. Commun. 10, 3763 (2019).
Noctor, S. C., Martinez-Cerdeno, V., Ivic, L. & Kriegstein, A. R. Cortical neurons arise in symmetric and asymmetric division zones and migrate through specific phases. Nat. Neurosci. 7, 136–144 (2004).
Anthony, T. E., Klein, C., Fishell, G. & Heintz, N. Radial glia serve as neuronal progenitors in all regions of the central nervous system. Neuron 41, 881–890 (2004).
Malatesta, P., Hartfuss, E. & Gotz, M. Isolation of radial glial cells by fluorescent-activated cell sorting reveals a neuronal lineage. Development 127, 5253–5263 (2000).
Noctor, S. C., Flint, A. C., Weissman, T. A., Dammerman, R. S. & Kriegstein, A. R. Neurons derived from radial glial cells establish radial units in neocortex. Nature 409, 714–720 (2001).
Englund, C. et al. Pax6, Tbr2, and Tbr1 are expressed sequentially by radial glia, intermediate progenitor cells, and postmitotic neurons in developing neocortex. J. Neurosci. 25, 247–251 (2005).
Martynoga, B., Drechsel, D. & Guillemot, F. Molecular control of neurogenesis: a view from the mammalian cerebral cortex. Cold Spring Harb. Perspect. Biol. 4, a008359 (2012).
Telley, L. et al. Temporal patterning of apical progenitors and their daughter neurons in the developing neocortex. Science 364, eaav2522 (2019).
Pollen, A. A. et al. Low-coverage single-cell mRNA sequencing reveals cellular heterogeneity and activated signaling pathways in developing cerebral cortex. Nat. Biotechnol. 32, 1053–1058 (2014).
Johnson, M. B. et al. Single-cell analysis reveals transcriptional heterogeneity of neural progenitors in human cortex. Nat. Neurosci. 18, 637–646 (2015).
Di Bella, D. J. et al. Molecular logic of cellular diversification in the mouse cerebral cortex. Nature 595, 554–55 (2021).
Fan, X. et al. Single-cell transcriptome analysis reveals cell lineage specification in temporal-spatial patterns in human cortical development. Sci. Adv. 6, eaaz2978 (2020).
Lunt, S. Y. & Vander Heiden, M. G. Aerobic glycolysis: meeting the metabolic requirements of cell proliferation. Annu. Rev. Cell Dev. Biol. 27, 441–464 (2011).
Shyh-Chang, N. & Ng, H. H. The metabolic programming of stem cells. Genes Dev. 31, 336–346 (2017).
Porporato, P. E., Dhup, S., Dadhich, R. K., Copetti, T. & Sonveaux, P. Anticancer targets in the glycolytic metabolism of tumors: a comprehensive review. Front. Pharmacol. 2, 49 (2011).
Kim, J. W., Tchernyshyov, I., Semenza, G. L. & Dang, C. V. HIF-1-mediated expression of pyruvate dehydrogenase kinase: a metabolic switch required for cellular adaptation to hypoxia. Cell Metab. 3, 177–185 (2006).
Ito, K. & Suda, T. Metabolic requirements for the maintenance of self-renewing stem cells. Nat. Rev. Mol. Cell Biol. 15, 243–256 (2014).
Patra, K. C. & Hay, N. The pentose phosphate pathway and cancer. Trends Biochem. Sci. 39, 347–354 (2014).
Felipe Barros, L. Metabolic signaling by lactate in the brain. Trends Neurosci. 36, 396–404 (2013).
Poetzsch, A. et al. L-lactate exerts a pro-proliferative effect on adult hippocampal precursor cells in vitro. iScience 24, 102126 (2021).
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575–57 (2019).
Daw, C. C. et al. Lactate elicits ER-mitochondrial Mg2+ dynamics to integrate cellular metabolism. Cell 183, 474–489 (2020).
Magistretti, P. J. & Allaman, I. Lactate in the brain: from metabolic end-product to signalling molecule. Nat. Rev. Neurosci. 19, 235–249 (2018).
Laughton, J. D. et al. Differential messenger RNA distribution of lactate dehydrogenase LDH-1 and LDH-5 isoforms in the rat brain. Neuroscience 96, 619–625 (2000).
Vegran, F., Boidot, R., Michiels, C., Sonveaux, P. & Feron, O. Lactate influx through the endothelial cell monocarboxylate transporter MCT1 supports an NF-κ B/IL-8 pathway that drives tumor angiogenesis. Cancer Res. 71, 2550–2560 (2011).
DeBerardinis, R. J. & Chandel, N. S. We need to talk about the Warburg effect. Nat. Metab. 2, 127–129 (2020).
Liberti, M. V. & Locasale, J. W. The Warburg effect: how does it benefit cancer cells? Trends Biochem. Sci. 41, 211–218 (2016).
Li, L. et al. Visualizing the distribution of synapses from individual neurons in the mouse brain. PLoS ONE 5, e11503 (2010).
Bultje, R. S. et al. Mammalian Par3 regulates progenitor cell asymmetric division via notch signaling in the developing neocortex. Neuron 63, 189–202 (2009).
Iwata, R., Casimir, P. & Vanderhaeghen, P. Mitochondrial dynamics in postmitotic cells regulate neurogenesis. Science 369, 858–85 (2020).
Khacho, M. et al. Mitochondrial dynamics impacts stem cell identity and fate decisions by regulating a nuclear transcriptional program. Cell Stem Cell 19, 232–247 (2016).
Pfeiffer, T., Schuster, S. & Bonhoeffer, S. Cooperation and competition in the evolution of ATP-producing pathways. Science 292, 504–507 (2001).
Heiden, M. G. V., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Wang, J. et al. Brain endothelial cells maintain lactate homeostasis and control adult hippocampal neurogenesis. Cell Stem Cell 25, 754–75 (2019).
Lev-Vachnish, Y. et al. L-lactate promotes adult hippocampal neurogenesis. Front. Neurosci. 13, 403 (2019).
Carrard, A. et al. Role of adult hippocampal neurogenesis in the antidepressant actions of lactate. Mol. Psychiatry 26, 6723–6735 (2021).
Javaherian, A. & Kriegstein, A. A stem cell niche for intermediate progenitor cells of the embryonic cortex. Cereb. Cortex 19, i70–i77 (2009).
Homem, C. C. F. et al. Ecdysone and mediator change energy metabolism to terminate proliferation in Drosophila neural stem cells. Cell 158, 874–888 (2014).
Khacho, M., Harris, R. & Slack, R. S. Mitochondria as central regulators of neural stem cell fate and cognitive function. Nat. Rev. Neurosci. 20, 34–48 (2019).
Peng, M. et al. Aerobic glycolysis promotes T helper 1 cell differentiation through an epigenetic mechanism. Science 354, 481–484 (2016).
Insolera, R., Bazzi, H., Shao, W., Anderson, K. V. & Shi, S.-H. Cortical neurogenesis in the absence of centrioles. Nat. Neurosci. 17, 1528–1535 (2014).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Cillo, A. R. et al. Immune landscape of viral- and carcinogen-driven head and neck cancer. Immunity 52, 183–199 (2020).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Van den Berge, K. et al. Trajectory-based differential expression analysis for single-cell sequencing data. Nat. Commun. 11, 1201 (2020).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the \(2^{{{-\Delta\Delta}{c}}_{\rm{T}}}\) method. Methods 25, 402–408 (2001).
Wang, X. W. & Seed, B. A PCR primer bank for quantitative gene expression analysis. Nucleic Acids Res. 31, e154 (2003).
Buescher, J. M. et al. A roadmap for interpreting 13C metabolite labeling patterns from cells. Curr. Opin. Biotechnol. 34, 189–201 (2015).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Shen, Q., Zhong, W. M., Jan, Y. N. & Temple, S. Asymmetric Numb distribution is critical for asymmetric cell division of mouse cerebral cortical stem cells and neuroblasts. Development 129, 4843–4853 (2002).
Ratz, M. et al. Clonal relations in the mouse brain revealed by single-cell and spatial transcriptomics. Nat. Neurosci. 25, 285–28 (2022).
Acknowledgements
We thank M. O. Li (Memorial Sloan Kettering Cancer Center) and H. Zheng (Tsinghua University) for kindly providing Ldhafl/fl mice and lentivirus package plasmids, respectively; X. Liu, X. Wang and Y. Wang from the Metabolomics Core at the Technology Center of Protein Research at Tsinghua University for metabolic assays; and Shi laboratory members for insightful discussions and inputs. This work was supported by the Ministry of Science and Technology of China (2021ZD0202300), the National Natural Science Foundation of China (32021002), the Beijing Outstanding Young Scientist Program (BJJWZYJH01201910003012) and the Beijing Municipal Science & Technology Commission (Z20111000530000 and Z211100003321001). The funders had no role in the study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
X.D. and S.-H.S. conceived the project. X.D. performed most experiments, with help from D.W. and Jiameng Ma for mitochondrial morphology analysis and from Jian Ma for mouse genetics. Q.Z. and X.Y. collected and analyzed the single-cell gene expression dataset. X.D. and S.-H.S. wrote the paper, with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Sama Sleiman and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Single cell transcriptomic analysis of the developing mouse neocortex at E9.5-E17.5.
(a) UMAP plot of the gene expression profiles of individual cells indicating their cell type identities (up) and developmental stages (down). NE, neuroepithelia; RGP, radial glial progenitor; IP, intermediate progenitor; EN, excitatory neuron; IN, interneuron; OPC, oligodendrocyte precursor cell; EpiC, epithelial cell; EndoC, endothelial cell; MGC, microglial cell; BC, blood cell; MC, meningeal cell. (b) Violin plots of the well-established marker gene expression for individual cell types. (c) UMAP plot of the expression profiles of the dorsal RGP lineage cells regarding cell types (up) or developmental stages (down). (d) UMAP plot of gene expression profiles of individual cells from a previously published scRNA-seq dataset of the developing mouse neocortices31 indicating cell type identities (left) and developmental stages (middle), and UMAP plot of the expression profiles of metabolic genes in individual RGPs, IPs, and ENs of the developing mouse neocortex that reveals 5 metabolic states (S1-S5, right). (e) Heatmap of the metabolic pathway enrichment analysis for each metabolic state. The enrichment score is scaled to its maximum value across states (red, maximum value; blue, minimum value). (f) UMAP plot of the expression profiles of metabolic genes in individual cells from a previously published scRNA-seq dataset of the developing human neocortices from 7 weeks (W) to 28 W32 that reveals 4 distinct metabolic states (S) 1–4. RGs, RGPs; vRGs, ventricular RGs; oRGs, outer subventricular zone RGs. (g) Pie charts showing the temporal/developmental stage composition of the cells in S4 and S5 in our dataset. (h) Visualization of all RGPs coloured by the developmental stage (top) or pseudo-time (bottom) in our dataset.
Extended Data Fig. 2 Systematic expression of glycolytic and TCA cycle genes at different metabolite states.
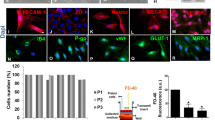
(a) The expression level of each gene is scaled to the maximum value across different metabolic states (yellow, maximum value; purple, minimum value). Red arrows indicate the well-known genes for different cell types or lineage progression statuses. (b, c) Heatmap of the glycolytic (b) and TCA cycle (c) genes expressed in specific metabolic states. Each gene is scaled to its maximum value across different states (red, maximum value; blue, minimum value). (d, e) Western blot assays showing the protein level of SLC2A1, HK2, ALDOA, GAPDH, ENO1, LDHA and PDK1 at the stages of E10.5, E13.5 and E15.5 (d), and quantification of the protein level normalized to ACTIN at different developmental stages (e) (n = 3 independent biological replicates at each stage). The statistical significance of the difference between E10.5 and E13.5 (purple), E10.5 and E15.5(yellow), and E13.5 and E15.5(red) is shown at the top. Data are shown as mean ± s.e.m. Two-tailed unpaired student’s t-test was used for statistical analysis.
Extended Data Fig. 3 Progressive vessel formation in the VZ of the developing neocortex.
(a) Representative images of E10.5, E13.5 or E15.5 brain sections immunostained for CD31(red), an endothelial cell marker, PAX6 (green), and counterstained with DAPI (blue). High magnification images (broken line rectangles) are shown at the bottom. Note the progressive formation of vessels in the VZ (broken lines). Scale bars: 200 μm and 50 μm. (b, c) Quantification of the percentage of vessel coverage in the dorsal neocortex (b) or in the VZ (c) at different developmental stages (n = 4 brains for each stage). (d) Representative images of E10.5, E13.5, or E15.5 brain sections immunostained for CD31 (red), PAX6 (green), and counterstained with DAPI (blue). The nucleus of endothelial cell was marked by broken circle. Scale bar: 20 μm. (e) Quantification of the numbers of endothelial cell and RGP in the VZ of the dorsal neocortex at different developmental stages (n = 4 brains for each stage). (f) Representative images of oxidized thiols staining in the developing neocortices at E10.5, E13.5, and E15.5. Colour bar indicates the intensity of oxidized thiols (white, maximum value; green, minimum value). Scale bar: 50 μm. Data are shown as mean ± s.e.m. Two-tailed unpaired student’s t-test was used for statistical analysis.
Extended Data Fig. 4 Metabolic flux of 13C6-glucose in E10.5 and E13.5 RGPs.
The schematic diagram of the glycolysis and TCA cycle pathway is shown at the centre. Relative metabolite abundance of E10.5 and E13.5 RGPs in the glycolysis (a, b) and TCA cycle (c-h) pathway revealed by 13C6-glucose tracing (n = 3 independent biological replicates). Note that 13C-labelled anaerobic glycolysis metabolites F-1,6-BP (a) and lactate (b) are more abundant in E10.5 RGPs than in E13.5 RGPs, whereas 13C-labelled TCA cycle metabolites acetyl-CoA (c), citrate (d), succinate (e), malate (f), 2-hydroxyglutarate (g) and aspartate (h) are more abundant in E13.5 RGPs than in E10.5 RGPs. Data are shown as mean ± s.e.m. Two-tailed unpaired student’s t-test was used for statistical analysis.
Extended Data Fig. 5 Proliferative RGPs at E10.5 metabolize glucose to generate a high level of nucleotide intermediates.
The schematic diagram of the purine and pyrimidine metabolism pathway is shown at the centre. Relative abundance of the metabolites in the purine and pyrimidine metabolism pathway in E10.5 or E13.5 RGPs revealed by 13C6-glucose tracing (n = 3 independent biological replicates). Note that 13C-labelled intermediate nucleotides ribose-5P, IMP, AMP, ADP, GDP, UMP, UDP and CDP are more abundant in E10.5 RGPs than in E13.5 RGPs, whereas 13C-labelled ATP, UTP and CTP are more abundant in E13.5 RGPs than in E10.5 RGPs. Data are shown as mean ± s.e.m. Two-tailed unpaired student’s t-test was used for statistical analysis.
Extended Data Fig. 6 Inhibition of lactate production and secretion disrupts vessel growth.
(a) Quantification of the lactate concentration in E11.5 VZ tissues cultured in vitro treated with LDHA/B inhibitor GSK2837808A (GSK) or DMSO as control for 4 hours (n = 3 independent biological replicates). (b, c) Quantification of the extracellular (b) and intracellular (c) lactate concentration in E10.5 RGPs cultured in vitro treated with GSK or DMSO for ~10 hours (n = 3 independent biological replicates). (d) Representative images of E11 brain sections treated with 10 mM GSK (right) or DMSO (left) as a control for ~5 hours, immunostained for cleaved CASPASE-3 (green), PAX6 (red) and counterstained with DAPI (blue). High magnification images of the dorsal neocortex (NCX, broken line rectangles) are shown on the right. Scale bar: 50 μm. (e) Quantification of cleaved CASPASE-3+ cells in the dorsal NCX (DMSO, n = 4 DMSO injected brains; GSK, n = 5 GSK injected brains). (f) Representative images of E11 brain sections treated with MCT inhibitor SR13800 (right) or DMSO (left) as a control for ~5 hours in vivo and immunostained for CD31 (red), and counterstained with DAPI (blue). Broken lines indicate the dorsal NCX. High magnification images of the NCX (broken line rectangles) are shown on the right. Note that the density of vessels in the SR13800-treated NCX is greatly reduced compared with the control. Scale bars: 200 μm, 100 μm, and 50 μm. (g) Quantification of the percentage of vessel coverage in the dorsal NCX. (DMSO, n = 4 DMSO injected brains; SR13800, n = 3 SR13800 injected brains). Data are shown as mean ± s.e.m. Two-tailed unpaired student’s t-test was used for statistical analysis.
Extended Data Fig. 7 Lactate stimulates vessel growth in the VZ via the functional murine IL-8 homologue CXCL1 signalling pathway.
(a) Represent images of E11 cortices immunostained for CXCL1 (left) or CXCL2 (right) (green) and CD31 (red), and counterstained with DAPI (blue). High magnification images (broken line rectangles) are shown to the right. Note that CXCL1, but not CXCL2, is prominently expressed in endothelial cells in the developing neocortex. Scale bar: 20 μm and 10 μm. (b) Representative images of E11 cortices treated with DMSO (top, left), LDH inhibitor GSK2837808A (GSK; top, right), Lactate (bottom, left), and GSK + Lactate (bottom, right) for ~5 hours in vivo and immunostained for CXCL1 (green) and CD31 (red), and counterstained with DAPI (blue). High magnification images (broken line rectangles) are shown at the right. Note the reduced expression of CXCL1 upon GSK treatment, which can be partially rescued by exogenous lactate supplement. Scale bar: 20 μm. (c) Quantification of CXCL1 staining intensity in endothelial cells. Each dot represents an endothelial cell (DMSO, n = 29 from 3 brains; GSK, n = 17 from 3 brains; Lactate, n = 16 from 3 brains; GSK + Lactate, n = 20 from 3 brains; a.u., arbitrary unit). (d) Representative images of E11 brain sections treated with 25 mM CXCL1/CXCR1 inhibitor Reparixin (right) or DMSO as a control (left) in vivo for ~5 hours, and immunostained for CD31 (red) and PAX6 (green), and counterstained with DAPI (blue). Note that the vessel coverage in the VZ is greatly reduced upon Reparixin treatment. Scale bars: 200 μm, 100 μm, and 50 μm. (e) Quantification of the percentage of vessel coverage in the dorsal NCX (DMSO, n = 4 brains; Reparixin, n = 5 brains). Data are shown as mean ± s.e.m. Two-way ANOVA test was used for statistical analysis.
Extended Data Fig. 8 Suppression of lactate production systematically upregulates neuronal differentiation genes expression and downregulates progenitor proliferation genes expression.
(a) Heatmap of progenitor proliferation and neuronal differentiation genes expression in GSK2837808A (GSK) treated RGPs compared with DMSO treated control (n = 3 independent biological replicates). (b) Gene Ontology (GO) term enrichment analysis of upregulated (up) and downregulated (down) genes in GSK treated RGPs compared with DMSO treated RGPs. (c) Schematic diagram of the glycerol-3 phosphate (glycerol-3P) metabolism pathway. (d-f) Relative metabolite abundance of lactate (d), acetyl-CoA (e) and glycerol-3 phosphate (f) in E10.5 RGPs treated with GSK revealed by 13C6-glucose tracing assay (n = 3 independent biological replicates). Note that 13C-labelled lactate were dramatically decreased, whereas 13C-labelled acetyl-CoA and glycerol-3P were significantly increased in GSK treated RGPs. (g, h) Relative metabolite abundance of lactate (g), and glycerol-3P (h) in E10.5 and E13.5 RGPs and P14 neurons revealed by 13C6-glucose tracing assay (n = 3 independent biological replicates). Note that 13C-labelled lactate was dramatically decreased, whereas glycerol-3P production was strongly increased, in P14 neurons compared with proliferative RGPs at E10.5. Data are shown as mean ± s.e.m. Two-tailed unpaired student’s t-test was used for statistical analysis.
Extended Data Fig. 9 Removal of Ldha and Ldhb reduces lactate production and promotes neuron differentiation.
(a) Western blot assay showing the efficacy of LDHA removal in Ldha cKO using Foxg1-Cre. (b) Quantification of lactate concentration in the VZ of the WT and Ldha cKO neocortices at E11.5 (n = 3 independent biological replicates). Note that there is no obvious difference in lactate concentration between WT and Ldha cKO. (c) Western blot assays showing the efficacy of Ldhb shRNAs in suppressing LDHB protein expression. (d) Representative images of E10.5 neocortical cultures from wildtype mice infected with lentiviruses carrying EGFP/control shRNA (top) or EGFP/Ldhb shRNA-b (bottom) for 48 hours, immunostained for EGFP (green) and PAX6 (red) or TUJ1 (white), and counterstained with DAPI (blue). Scale bar: 20 μm. (e) Quantification of lactate concentration in the EGFP+ cell from wildtype mice infected with lentiviruses carrying EGFP/control shRNA or EGFP/Ldhb shRNA-b in vitro culture for 48 hours (n = 3 independent biological replicates). Note that no obvious difference in lactate concentration between WT and Ldhb knockdown. (f) Quantification of the percentage of EGFP+ cells that are PAX6+ or TUJ1+ (n = 3 independent biological replicates). (g) Diagram of removal of Ldha and Ldhb by combining Ldha cKO and Ldhb shRNA knockdown. (h) Quantification of lactate concentration in the EGFP + cell from wildtype or Ldha cKO mice infected with lentiviruses carrying EGFP/control shRNA (top) or EGFP/Ldhb shRNA (bottom) in vitro culture for 48 hours, respectively. Note that the obvious decrease in lactate concentration between WT and Ldha cKO;Ldhb shRNA-b (n = 3 independent biological replicates). (i) Representative images of E10.5 neocortical cultures from wildtype (WT, top) or Ldha cKO (bottom) mice infected with lentiviruses carrying EGFP/control shRNA (top) or EGFP/Ldhb shRNA (bottom) for 72 hours, immunostained for EGFP (green) and PAX6 (red) or TUJ1 (white), and counterstained with DAPI (blue). Scale bar: 25 μm. (j) Quantification of the percentage of EGFP+ cells that are PAX6+ or TUJ1+ (n = 3 independent biological replicates). Data are shown as mean ± s.e.m. Two-tailed unpaired student’s t-test was used for statistical analysis.
Extended Data Fig. 10 Regulation of mitochondrial length in RGPs by lactate.
(a) Representative images of mitochondrial morphology in RGPs immunostained for EGFP (green) and mito-dsRed (red), and counterstained with DAPI (blue). Scale bar: 5 μm. (b) Quantification of the mitochondrial length in RGPs (E10.5, n = 25 from 2 embryonic mice; E13.5, n = 31 from 3 embryonic mice). (c) Representative images of E10.5 or E13.5 RGPs cultured in vitro for 48 hours, immunostained for TOMM20 (white) and PAX6 (green), and counterstained with DAPI (blue). Scale bar: 5 μm. (d) Quantification of the mitochondrial length in RGPs (E10.5, n = 61 from 3 independent biological experiments; E13.5, n = 86 from 3 independent biological experiments). (e) Representative images of E10.5 or E13.5 neurons cultured in vitro for 48 hours, immunostained for TOMM20 (white) and TUJ1 (red), and counterstained with DAPI (blue). Scale bar: 5 μm. (f) Quantification of the mitochondrial length in neurons (E10.5, n = 54 from 3 independent biological experiments; E13.5, n = 69 from 3 independent biological experiments). (g) Representative images of E10.5 neocortical culture in vitro treated with LDH inhibitor GSK2837808A (GSK, bottom) or DMSO (top) for ~48 hours, immunostained for PAX6 (green) and TOMM20 (white), and counterstained with DAPI (blue). Scale bar: 5 μm. (h) Quantification of the mitochondrial length in RGPs (DMSO, n = 52 from 3 independent biological experiments; GSK, n = 30 from 3 independent biological experiments). (i) Representative images of E10.5 neocortical culture in vitro treated with GSK (bottom) or DMSO (top) for ~48 hours, immunostained for TUJ1 (red) and TOMM20 (white), and counterstained with DAPI (blue). Scale bar: 5 μm. (j) Quantification of mitochondrial length in neurons (DMSO, n = 31 from 3 independent biological experiments; GSK, n = 30 from 3 independent biological experiments). (k) Representative images of E10.5 neocortical cultures treated with GSK or DMSO for ~48 hours, immunostained for PAX6 (red) and TUJ1 (white), and counterstained with DAPI (blue). Scale bar: 20 μm. (l-n) Quantification of the total cell density (l) or the density of PAX6+ cells (m) per 100-μm2 area, or the percentage of PAX6+ or TUJ1+ cells (n) (n = 3 independent biological experiments). Data are shown as mean ± s.e.m. Two tailed unpaired student’s t-test was used for statistical analysis.
Supplementary information
Supplementary Information
Supplementary Fig. 1. FACS gating strategy. Representative gating strategy for all FACS panels: size selection, single-cell selection, live-cell selection and CD133-APC+ cell gating
Supplementary Table 1
Mouse information used in the experiments
Supplementary Data 1
The matrix of metabolic gene expression used for single-cell metabolic state analysis
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 2
Unprocessed Western Blots and Statistical Source Data
Source Data Extended Data Fig. 2
Unprocessed Western Blots and Statistical Source Data
Source Data Extended Data Fig. 2
Unprocessed Western Blots and Statistical Source Data
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 8
Statistical Source Data
Source Data Extended Data Fig. 9
Unprocessed Western Blots and Statistical Source Data
Source Data Extended Data Fig. 9
Unprocessed Western Blots and Statistical Source Data
Source Data Extended Data Fig. 10
Statistical Source Data
Rights and permissions
About this article
Cite this article
Dong, X., Zhang, Q., Yu, X. et al. Metabolic lactate production coordinates vasculature development and progenitor behavior in the developing mouse neocortex. Nat Neurosci 25, 865–875 (2022). https://doi.org/10.1038/s41593-022-01093-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-022-01093-7
- Springer Nature America, Inc.
This article is cited by
-
Foxp1 suppresses cortical angiogenesis and attenuates HIF-1alpha signaling to promote neural progenitor cell maintenance
EMBO Reports (2024)
-
Targeting adenosine A2A receptors for early intervention of retinopathy of prematurity
Purinergic Signalling (2024)
-
The role of lactate in cardiovascular diseases
Cell Communication and Signaling (2023)
-
A breakdown of metabolic reprogramming in microglia induced by CKLF1 exacerbates immune tolerance in ischemic stroke
Journal of Neuroinflammation (2023)
-
Neurogenic–angiogenic synchrony via lactate
Nature Neuroscience (2022)