Abstract
Genome-wide association studies (GWAS) have identified hundreds of loci associated with kidney-related traits such as glomerular filtration rate, albuminuria, hypertension, electrolyte and metabolite levels. However, these impressive, large-scale mapping approaches have not always translated into an improved understanding of disease or development of novel therapeutics. GWAS have several important limitations. Nearly all disease-associated risk loci are located in the non-coding region of the genome and therefore, their target genes, affected cell types and regulatory mechanisms remain unknown. Genome-scale approaches can be used to identify associations between DNA sequence variants and changes in gene expression (quantified through bulk and single-cell methods), gene regulation and other molecular quantitative trait studies, such as chromatin accessibility, DNA methylation, protein expression and metabolite levels. Data obtained through these approaches, used in combination with robust computational methods, can deliver robust mechanistic inferences for translational exploitation. Understanding the genetic basis of common kidney diseases means having a comprehensive picture of the genes that have a causal role in disease development and progression, of the cells, tissues and organs in which these genes act to affect the disease, of the cellular pathways and mechanisms that drive disease, and of potential targets for disease prevention, detection and therapy.
Key points
-
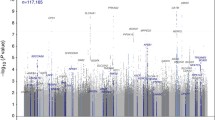
Genome-wide association studies (GWAS) have identified a large number of nucleotide variations that show strong and reproducible association with kidney-related traits such as serum and urine metabolites, estimated glomerular filtration rate, albuminuria and hypertension.
-
More than 95% of disease-associated GWAS signals are located in non-coding regions of the genome, which can be identified by analysing chromatin accessibility and conformation, DNA methylation and transcription factor binding.
-
Disease-associated genetic variants seem to be located in cell type-specific gene regulatory regions, where they modulate disease risk by quantitatively altering gene transcript levels.
-
The use of advanced computational approaches to combine datasets orthogonal to GWAS data, including molecular quantitative trait studies, single-cell transcriptomics and epigenetic information, is necessary for the prioritization of GWAS variants.



Similar content being viewed by others
References
Warejko, J. K. et al. Whole exome sequencing of patients with steroid-resistant nephrotic syndrome. Clin. J. Am. Soc. Nephrol. 13, 53–62 (2018).
Mallawaarachchi, A. C. et al. Whole-genome sequencing overcomes pseudogene homology to diagnose autosomal dominant polycystic kidney disease. Eur. J. Hum. Genet. 24, 1584–1590 (2016).
Boucher, C. & Sandford, R. Autosomal dominant polycystic kidney disease (ADPKD, MIM 173900, PKD1 and PKD2 genes, protein products known as polycystin-1 and polycystin-2). Eur. J. Hum. Genet. 12, 347–354 (2004).
Dempfle, A. et al. Gene-environment interactions for complex traits: definitions, methodological requirements and challenges. Eur. J. Hum. Genet. 16, 1164–1172 (2008).
Bomba, L., Walter, K. & Soranzo, N. The impact of rare and low-frequency genetic variants in common disease. Genome Biol. 18, 77 (2017).
Stringer, S. et al. Underestimated effect sizes in GWAS: fundamental limitations of single SNP analysis for dichotomous phenotypes. PLoS ONE 6, e27964 (2011).
Visscher, P. M. et al. Five years of GWAS discovery. Am. J. Hum. Genet. 90, 7–24 (2012).
Wuttke, M. et al. A catalog of genetic loci associated with kidney function from analyses of a million individuals. Nat. Genet. 51, 957–972 (2019).
Morris, A. P. et al. Trans-ethnic kidney function association study reveals putative causal genes and effects on kidney-specific disease aetiologies. Nat. Commun. 10, 29 (2019).
Hellwege, J. N. et al. Mapping eGFR loci to the renal transcriptome and phenome in the VA Million Veteran Program. Nat. Commun. 10, 3842 (2019).
Kanai, M. et al. Genetic analysis of quantitative traits in the Japanese population links cell types to complex human diseases. Nat. Genet. 50, 390–400 (2018).
Kottgen, A. et al. Multiple loci associated with indices of renal function and chronic kidney disease. Nat. Genet. 41, 712–717 (2009).
Teumer, A. et al. Genome-wide association studies identify genetic loci associated with albuminuria in diabetes. Diabetes 65, 803–817 (2016).
Ehret, G. B. et al. The genetics of blood pressure regulation and its target organs from association studies in 342,415 individuals. Nat. Genet. 48, 1171–1184 (2016).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012). This study shows that disease-causing variants localize to gene regulatory regions.
Schaub, M. A. et al. Linking disease associations with regulatory information in the human genome. Genome Res. 22, 1748–1759 (2012).
Schaid, D. J., Chen, W. & Larson, N. B. From genome-wide associations to candidate causal variants by statistical fine-mapping. Nat. Rev. Genet. 19, 491–504 (2018). This study highlights the method of fine-mapping to identify causal variants.
Dawn Teare, M. & Barrett, J. H. Genetic linkage studies. Lancet 366, 1036–1044 (2005).
Brown, C. R. et al. Linking stochastic fluctuations in chromatin structure and gene expression. PLoS Biol. 11, e1001621 (2013).
Pattaro, C. et al. Genetic associations at 53 loci highlight cell types and biological pathways relevant for kidney function. Nat. Commun. 7, 10023 (2016).
Park, J. et al. Single-cell transcriptomics of the mouse kidney reveals potential cellular targets of kidney disease. Science 360, 758–763 (2018). First study in the kidney to elucidate kidney disease target genes at the cellular level.
Qiu, C. et al. Renal compartment-specific genetic variation analyses identify new pathways in chronic kidney disease. Nat. Med. 24, 1721–1731 (2018). Study of human kidney compartment-specific eQTLs that identified and modelled target genes.
Uh, H. W. et al. How to deal with the early GWAS data when imputing and combining different arrays is necessary. Eur. J. Hum. Genet. 20, 572–576 (2012).
Flister, M. J. et al. Identifying multiple causative genes at a single GWAS locus. Genome Res. 23, 1996–2002 (2013).
Liu, B. & Montgomery, S. B. Identifying causal variants and genes using functional genomics in specialized cell types and contexts. Hum. Genet. 139, 95–102 (2020).
Sebastiani, P. et al. Genome-wide association studies and the genetic dissection of complex traits. Am. J. Hematol. 84, 504–515 (2009).
Pare, G., Mao, S. & Deng, W. Q. A machine-learning heuristic to improve gene score prediction of polygenic traits. Sci. Rep. 7, 12665 (2017).
Visscher, P. M. et al. 10 years of GWAS discovery: biology, function, and translation. Am. J. Hum. Genet. 101, 5–22 (2017).
Kiryluk, K. et al. GWAS for serum galactose-deficient IgA1 implicates critical genes of the O-glycosylation pathway. PLoS Genet. 13, e1006609 (2017).
Zhou, W. et al. Genome-wide prediction of DNase I hypersensitivity using gene expression. Nat. Commun. 8, 1038 (2017).
Buenrostro, J. D. et al. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Beckerman, P., Ko, Y. A. & Susztak, K. Epigenetics: a new way to look at kidney diseases. Nephrol. Dial. Transplant. 29, 1821–1827 (2014).
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Heintzman, N. D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009).
Kimura, H. Histone modifications for human epigenome analysis. J. Hum. Genet. 58, 439–445 (2013).
Jones, P. A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Luo, C., Hajkova, P. & Ecker, J. R. Dynamic DNA methylation: in the right place at the right time. Science 361, 1336–1340 (2018).
Wain, L. V. et al. Novel blood pressure locus and gene discovery using genome-wide association study and expression data sets from blood and the kidney. Hypertension 70, e4–e19 (2017).
Moran, S. & Esteller, M. Infinium DNA methylation microarrays on formalin-fixed, paraffin-embedded samples. Methods Mol. Biol. 1766, 83–107 (2018).
Mendizabal, I. et al. Cell type-specific epigenetic links to schizophrenia risk in the brain. Genome Biol. 20, 135 (2019).
Gaudreault, M. et al. Electrophoretic mobility shift assays for the analysis of DNA-protein interactions. Methods Mol. Biol. 543, 15–35 (2009).
Perna, A. & Alberi, L. A. TF-ChIP method for tissue-specific gene targets. Front. Cell Neurosci. 13, 95 (2019).
Vorontsov, I. E. et al. Genome-wide map of human and mouse transcription factor binding sites aggregated from ChIP-Seq data. BMC Res. Notes 11, 756 (2018).
Kidder, B. L., Hu, G. & Zhao, K. ChIP-Seq: technical considerations for obtaining high-quality data. Nat. Immunol. 12, 918–922 (2011).
Nishizaki, S. S. et al. Predicting the effects of SNPs on transcription factor binding affinity. Bioinformatics 36, 364–372 (2020).
Behera, V. et al. Exploiting genetic variation to uncover rules of transcription factor binding and chromatin accessibility. Nat. Commun. 9, 782 (2018).
Pennacchio, L. A. et al. Enhancers: five essential questions. Nat. Rev. Genet. 14, 288–295 (2013).
Wu, C. & Pan, W. Integration of enhancer-promoter interactions with GWAS summary results identifies novel schizophrenia-associated genes and pathways. Genetics 209, 699–709 (2018).
Dekker, J. et al. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechnol. 31, 1119–1125 (2013).
Li, G. et al. Chromatin interaction analysis with paired-end tag (ChIA-PET) sequencing technology and application. BMC Genomics 15 (Suppl 12), S11 (2014).
Li, G. et al. ChIA-PET tool for comprehensive chromatin interaction analysis with paired-end tag sequencing. Genome Biol. 11, R22 (2010).
Vanhille, L. et al. High-throughput and quantitative assessment of enhancer activity in mammals by CapStarr-seq. Nat. Commun. 6, 6905 (2015).
Muerdter, F., Boryn, L. M. & Arnold, C. D. STARR-seq – principles and applications. Genomics 106, 145–150 (2015).
Arnold, C. D. et al. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 (2013).
Inoue, F. & Ahituv, N. Decoding enhancers using massively parallel reporter assays. Genomics 106, 159–164 (2015).
Zabidi, M. A. et al. Enhancer-core-promoter specificity separates developmental and housekeeping gene regulation. Nature 518, 556–559 (2015).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Wu, S. et al. Genome-wide association studies and CRISPR/Cas9-mediated gene editing identify regulatory variants influencing eyebrow thickness in humans. PLoS Genet. 14, e1007640 (2018).
Gasperini, M. et al. A genome-wide framework for mapping gene regulation via cellular genetic screens. Cell 176, 1516 (2019).
Schwank, G. et al. Functional repair of CFTR by CRISPR/Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell 13, 653–658 (2013).
Roper, J. & Yilmaz, O. H. Breakthrough moments: genome editing and organoids. Cell Stem Cell 24, 841–842 (2019).
Diabetes Genetics Replication and Meta-analysis (DIAGRAM) Consortium. et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat. Genet. 46, 234–244 (2014).
Soldner, F. et al. Parkinson-associated risk variant in distal enhancer of α-synuclein modulates target gene expression. Nature 533, 95–99 (2016).
Genovese, G. et al. Association of trypanolytic ApoL1 variants with kidney disease in African Americans. Science 329, 841–845 (2010).
van der Wijst, M. G. P. et al. Single-cell RNA sequencing identifies celltype-specific cis-eQTLs and co-expression QTLs. Nat. Genet. 50, 493–497 (2018).
Wang, X. et al. Bulk tissue cell type deconvolution with multi-subject single-cell expression reference. Nat. Commun. 10, 380 (2019).
Neumeyer, S., Hemani, G. & Zeggini, E. Strengthening causal inference for complex disease using molecular quantitative trait loci. Trends. Mol. Med. 26, 232–241 (2020).
Franke, L. & Jansen, R. C. eQTL analysis in humans. Methods Mol. Biol. 573, 311–328 (2009).
Guo, X. et al. A comprehensive cis-eQTL analysis revealed target genes in breast cancer susceptibility loci identified in genome-wide association studies. Am. J. Hum. Genet. 102, 890–903 (2018).
Yao, C. et al. Dynamic role of trans regulation of gene expression in relation to complex traits. Am. J. Hum. Genet. 100, 985–986 (2017).
Bryois, J. et al. Cis and trans effects of human genomic variants on gene expression. PLoS Genet. 10, e1004461 (2014).
Westra, H. J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Võsa, U. et al. Unraveling the polygenic architecture of complex traits using blood eQTL metaanalysis. Preprint at https://www.biorxiv.org/content/10.1101/447367v1 (2018)
Liu, X., Li, Y. I. & Pritchard, J. K. Trans effects on gene expression can drive omnigenic inheritance. Cell 177, 1022–1034.e6 (2019).
GTEx Consortium Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
GTEx Consortium et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Pastinen, T. & Hudson, T. J. Cis-acting regulatory variation in the human genome. Science 306, 647–650 (2004).
Barlow, D. P. & Bartolomei, M. S. Genomic imprinting in mammals. Cold Spring Harb Perspect Biol 6, a018382 (2014).
Khansefid, M. et al. Comparing allele specific expression and local expression quantitative trait loci and the influence of gene expression on complex trait variation in cattle. BMC Genomics 19, 793 (2018).
Degner, J. F. et al. Effect of read-mapping biases on detecting allele-specific expression from RNA-sequencing data. Bioinformatics 25, 3207–3212 (2009).
Baralle, F. E. & Giudice, J. Alternative splicing as a regulator of development and tissue identity. Nat. Rev. Mol. Cell Biol. 18, 437–451 (2017).
Li, Y. I. et al. RNA splicing is a primary link between genetic variation and disease. Science 352, 600–604 (2016).
Takata, A., Matsumoto, N. & Kato, T. Genome-wide identification of splicing QTLs in the human brain and their enrichment among schizophrenia-associated loci. Nat. Commun. 8, 14519 (2017).
Gate, R. E. et al. Genetic determinants of co-accessible chromatin regions in activated T cells across humans. Nat. Genet. 50, 1140–1150 (2018). This study used ATAC-seq and RNA-seq data and showed that human risk variants can be detected within accessible chromatin in activated T cells and localize with RNA-seq data.
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Wainberg, M. et al. Opportunities and challenges for transcriptome-wide association studies. Nat. Genet. 51, 592–599 (2019).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Gamazon, E. R. et al. A gene-based association method for mapping traits using reference transcriptome data. Nat. Genet. 47, 1091–1098 (2015).
Grubert, F. et al. Genetic control of chromatin states in humans involves local and distal chromosomal interactions. Cell 162, 1051–1065 (2015).
Porcu, E. et al. Mendelian randomization integrating GWAS and eQTL data reveals genetic determinants of complex and clinical traits. Nat. Commun. 10, 3300 (2019).
Lawlor, D. A., Tilling, K. & Davey Smith, G. Triangulation in aetiological epidemiology. Int. J. Epidemiol. 45, 1866–1886 (2016).
Davies, N. M., Holmes, M. V. & Davey Smith, G. Reading Mendelian randomisation studies: a guide, glossary, and checklist for clinicians. BMJ 362, k601 (2018).
Lawlor, D. A. et al. Mendelian randomization: using genes as instruments for making causal inferences in epidemiology. Stat. Med. 27, 1133–1163 (2008).
Smith, G. D. & Ebrahim, S. ‘Mendelian randomization’: can genetic epidemiology contribute to understanding environmental determinants of disease? Int. J. Epidemiol. 32, 1–22 (2003).
Del Greco, M. F. et al. Serum iron level and kidney function: a Mendelian randomization study. Nephrol. Dial. Transplant. 32, 273–278 (2017).
Gallagher, M. D. & Chen-Plotkin, A. S. The post-GWAS era: from association to function. Am. J. Hum. Genet. 102, 717–730 (2018).
Lan, F. et al. Droplet barcoding for massively parallel single-molecule deep sequencing. Nat. Commun. 7, 11784 (2016).
Zhang, X. et al. Comparative analysis of droplet-based ultra-high-throughput single-cell RNA-seq systems. Mol. Cell 73, 130–142.e5 (2019).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Cusanovich, D. A. et al. Multiplex single cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Jia, G. et al. Single cell RNA-seq and ATAC-seq analysis of cardiac progenitor cell transition states and lineage settlement. Nat. Commun. 9, 4877 (2018).
Giri, A. et al. Trans-ethnic association study of blood pressure determinants in over 750,000 individuals. Nat. Genet. 51, 51–62 (2019).
Teumer, A. et al. Genome-wide association meta-analyses and fine-mapping elucidate pathways influencing albuminuria. Nat. Commun. 10, 4130 (2019).
Nakatochi, M. et al. Genome-wide meta-analysis identifies multiple novel loci associated with serum uric acid levels in Japanese individuals. Commun. Biol 2, 115 (2019).
Sieber, K. B. et al. Integrated functional genomic analysis enables annotation of kidney genome-wide association study loci. J. Am. Soc. Nephrol. 30, 421–441 (2019). This study used DNase-seq and RNA-seq and found that disease-causing variants in GWAS are localized in regulatory gene elements and affect gene expression.
Ko, Y. A. et al. Genetic-variation-driven gene-expression changes highlight genes with important functions for kidney disease. Am. J. Hum. Genet. 100, 940–953 (2017).
Xu, X. et al. Molecular insights into genome-wide association studies of chronic kidney disease-defining traits. Nat. Commun. 9, 4800 (2018).
Gillies, C. E. et al. An eQTL landscape of kidney tissue in human nephrotic syndrome. Am. J. Hum. Genet. 103, 232–244 (2018). This human study identified eQTLs and potential target genes in patients with nephrotic syndrome.
Kottgen, A. et al. Uromodulin levels associate with a common UMOD variant and risk for incident CKD. J. Am. Soc. Nephrol. 21, 337–344 (2010).
Trudu, M. et al. Common noncoding UMOD gene variants induce salt-sensitive hypertension and kidney damage by increasing uromodulin expression. Nat. Med. 19, 1655–1660 (2013).
Bernascone, I. et al. A transgenic mouse model for uromodulin-associated kidney diseases shows specific tubulo-interstitial damage, urinary concentrating defect and renal failure. Hum. Mol. Genet. 19, 2998–3010 (2010).
Menon, M. C. et al. Intronic locus determines SHROOM3 expression and potentiates renal allograft fibrosis. J. Clin. Invest. 125, 208–221 (2015).
Khalili, H. et al. Developmental origins for kidney disease due to Shroom3 deficiency. J. Am. Soc. Nephrol. 27, 2965–2973 (2016).
Yeo, N. C. et al. Shroom3 contributes to the maintenance of the glomerular filtration barrier integrity. Genome Res. 25, 57–65 (2015).
Lovric, S. et al. Genetic testing in steroid-resistant nephrotic syndrome: when and how? Nephrol. Dial. Transplant. 31, 1802–1813 (2016).
Bryer, J. S. & Susztak, K. Screening drugs for kidney disease: targeting the podocyte. Cell Chem. Biol. 25, 126–127 (2018).
Beck, L. H. Jr. et al. M-type phospholipase A2 receptor as target antigen in idiopathic membranous nephropathy. N. Engl. J. Med. 361, 11–21 (2009).
Foley, R. N. & Collins, A. J. The USRDS: what you need to know about what it can and can’t tell us about ESRD. Clin. J. Am. Soc. Nephrol. 8, 845–851 (2013).
Groopman, E. E. et al. Diagnostic utility of exome sequencing for kidney disease. N. Engl. J. Med. 380, 142–151 (2019).
Susztak, K. Understanding the epigenetic syntax for the genetic alphabet in the kidney. J. Am. Soc. Nephrol. 25, 10–17 (2014).
Braun, D. A. et al. Whole exome sequencing identifies causative mutations in the majority of consanguineous or familial cases with childhood-onset increased renal echogenicity. Kidney Int. 89, 468–475 (2016).
Flannick, J. et al. Exome sequencing of 20,791 cases of type 2 diabetes and 24,440 controls. Nature 570, 71–76 (2019).
Nelson, M. R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Ling, G. et al. Unbiased, genome-wide in vivo mapping of transcriptional regulatory elements reveals sex differences in chromatin structure associated with sex-specific liver gene expression. Mol. Cell Biol. 30, 5531–5544 (2010).
Lu, Q. & Richardson, B. DNaseI hypersensitivity analysis of chromatin structure. Methods Mol. Biol. 287, 77–86 (2004).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
O’Geen, H., Echipare, L. & Farnham, P. J. Using ChIP-seq technology to generate high-resolution profiles of histone modifications. Methods Mol. Biol. 791, 265–286 (2011).
Moran, S., Arribas, C. & Esteller, M. Validation of a DNA methylation microarray for 850,000 CpG sites of the human genome enriched in enhancer sequences. Epigenomics 8, 389–399 (2016).
Sandoval, J. et al. Validation of a DNA methylation microarray for 450,000 CpG sites in the human genome. Epigenetics 6, 692–702 (2011).
Li, Q., Hermanson, P. J. & Springer, N. M. Detection of DNA methylation by whole-genome bisulfite sequencing. Methods Mol. Biol. 1676, 185–196 (2018).
Gu, H. et al. Preparation of reduced representation bisulfite sequencing libraries for genome-scale DNA methylation profiling. Nat. Protoc. 6, 468–481 (2011).
Khulan, B. et al. Comparative isoschizomer profiling of cytosine methylation: the HELP assay. Genome Res. 16, 1046–1055 (2006).
Andersson, R. & Sandelin, A. Determinants of enhancer and promoter activities of regulatory elements. Nat Rev Genet 21, 71–87 (2019).
Angermueller, C. et al. Parallel single-cell sequencing links transcriptional and epigenetic heterogeneity. Nat. Methods 13, 229–232 (2016).
Clark, S. J. et al. scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells. Nat. Commun. 9, 781 (2018).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Peterson, V. M. et al. Multiplexed quantification of proteins and transcripts in single cells. Nat. Biotechnol. 35, 936–939 (2017).
Pelikan, R. C. et al. Enhancer histone-QTLs are enriched on autoimmune risk haplotypes and influence gene expression within chromatin networks. Nat. Commun. 9, 2905 (2018).
McRae, A. F. et al. Identification of 55,000 replicated DNA methylation QTL. Sci. Rep. 8, 17605 (2018).
Yao, C. et al. Genome-wide mapping of plasma protein QTLs identifies putatively causal genes and pathways for cardiovascular disease. Nat. Commun. 9, 3268 (2018).
Nicholson, G. et al. A genome-wide metabolic QTL analysis in Europeans implicates two loci shaped by recent positive selection. PLoS Genet. 7, e1002270 (2011).
Ko, Y. A. et al. Cytosine methylation changes in enhancer regions of core pro-fibrotic genes characterize kidney fibrosis development. Genome Biol. 14, R108 (2013). First study to examine methylation changes in patients with chronic kidney disease showing that these changes occur in enhancer regions and affect fibrosis-related genes.
Qiu, C. et al. Cytosine methylation predicts renal function decline in American Indians. Kidney Int. 93, 1417–1431 (2018).
Simon, J. M., Giresi, P. G., Davis, I. J. & Lieb, J. D. A detailed protocol for formaldehyde-assisted isolation of regulatory elements (FAIRE). Curr. Protoc. Mol. Biol. 102, 21.26.1–21.26.15 (2013).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Futema, M. et al. Whole exome sequencing of familial hypercholesterolaemia patients negative for LDLR/APOB/PCSK9 mutations. J. Med. Genet. 51, 537–544 (2014).
Lata, S. et al. Whole-exome sequencing in adults with chronic kidney disease: a pilot study. Ann. Intern. Med. 168, 100–109 (2018).
Salem, R. M. et al. Genome-wide association study of diabetic kidney disease highlights biology involved in glomerular basement membrane collagen. J. Am. Soc. Nephrol. 30, 2000–2016 (2019).
Acknowledgements
K.S. is supported by NIH National Institute of Diabetes and Digestive and Kidney Diseases grants R01DK076077, R01 DK087635 and DP3 DK108220.
Author information
Authors and Affiliations
Contributions
Both authors wrote, reviewed or edited the manuscript before submission. K.M.S. researched data for the article and K.S. made substantial contributions to the discussion of content.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Nephrology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Encyclopedia of DNA Elements (ENCODE): https://www.encodeproject.org
Genotype–Tissue Expression (GTEx) Portal: https://gtexportal.org/home/
International Human Epigenome Consortium (IHEC): http://ihec-epigenomes.org
NEPH eQTL browser: http://nephqtl.org
NIH Roadmap Epigenomics Project: http://www.roadmapepigenomics.org
Susztaklab Human Kidney eQTL atlas: http://susztaklab.com/eqtl/
Glossary terms
- Secondary chromatin structure
-
The structure formed by the folding of primary chromatin.
- Fine mapping
-
The process by which a variant is assigned to a complex trait.
- Quantitative trait loci
-
(QTL) Genetic loci that are associated with variation in a phenotype.
- Imputation
-
The inclusion of genotypes that are not directly measured, by estimating the missing data based on reference genome datasets and SNP linkage.
- Haplotype
-
A set of genetic polymorphisms that are inherited together.
- Promoter
-
The DNA sequences on which transcription is initiated.
- Enhancer
-
A DNA sequence that is bound by transcription factors to increase the transcription of a gene.
- Insulator
-
A DNA sequence that limits chromatin activation.
- Transposable elements
-
DNA sequences whose position can change within the genome.
- Cis-regulatory elements
-
Regions of non-coding DNA that affect the expression of nearby genes.
- Massively parallel reporter assays
-
High-throughput experiments, in which the transcriptional activities of many regulatory sequences can be determined.
- Alternative splicing
-
A process by which multiple transcripts are produced from a single gene.
- Posterior probability
-
The probability of an event occurring based on information from a prior event.
- Mendelian randomization
-
A technique that uses genetic data to make causative inference.
- Orthogonal datasets
-
Independent and uncorrelated datasets.
- Cellular encapsulation
-
A technique whereby cells are entrapped in a spherical semipermeable polymeric membrane.
- Cellular deconvolution
-
The estimation of gene expression data for individual cell types within a bulk expression dataset.
Rights and permissions
About this article
Cite this article
Sullivan, K.M., Susztak, K. Unravelling the complex genetics of common kidney diseases: from variants to mechanisms. Nat Rev Nephrol 16, 628–640 (2020). https://doi.org/10.1038/s41581-020-0298-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41581-020-0298-1
- Springer Nature Limited
This article is cited by
-
Unraveling the epigenetic code: human kidney DNA methylation and chromatin dynamics in renal disease development
Nature Communications (2024)
-
Identification of genetic variants associated with diabetic kidney disease in multiple Korean cohorts via a genome-wide association study mega-analysis
BMC Medicine (2023)
-
Comprehensive transcriptional variability analysis reveals gene networks regulating seed oil content of Brassica napus
Genome Biology (2022)
-
Linking genetic variants to kidney disease via the epigenome
Nature Genetics (2022)
-
From mapping kidney function to mechanism and prediction
Nature Reviews Nephrology (2022)