Abstract
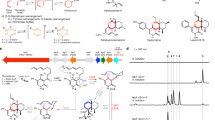
LepI is an S-adenosylmethionine (SAM)-dependent pericyclase that catalyses the formation of the 2-pyridone natural product leporin C. Biochemical characterization has shown that LepI can catalyse stereoselective dehydration to yield a reactive (E)-quinone methide that can undergo bifurcating intramolecular Diels–Alder (IMDA) and hetero-Diels–Alder (HDA) cyclizations from an ambimodal transition state, as well as a [3,3]-retro-Claisen rearrangement to recycle the IMDA product into leporin C. Here, we solve the X-ray crystal structures of SAM-bound LepI and in complex with a substrate analogue, the product leporin C, and a retro-Claisen reaction transition-state analogue to understand the structural basis for the multitude of reactions. Structural and mutational analysis reveals how nature evolves a classic methyltransferase active site into one that can serve as a dehydratase and a multifunctional pericyclase. Catalysis of both sets of reactions employs H133 and R295, two active-site residues that are not found in canonical methyltransferases. An alternative role of SAM, which is not found to be in direct contact with the substrate, is also proposed.





Similar content being viewed by others
Data availability
The data that support the findings of this study are available within this Article and its Supplementary Information, or are available from the corresponding authors upon reasonable request. The structural factors and coordinates of the LepI–SAM complex, LepI–SAM-1 complex, LepI (C52A)–SAH-1 complex, LepI (C52A)–SAM-8 complex and LepI (C52A)–SAM-10 complex have been deposited in the Protein Data Bank under ID nos. 6IX3, 6IX5, 6IX7, 6IX8 and 6IX9, respectively.
References
Nicolaou, K. C., Snyder, S. A., Montagnon, T. & Vassilikogiannakis, G. The Diels–Alder reaction in total synthesis. Angew. Chem. Int. Ed. 41, 1668–1698 (2002).
Cooper, S. J., Ohashi, M., Liu, F., Tang, Y. & Houk, K. N. The expanding world of biosynthetic pericyclases: cooperation of experiment and theory for discovery. Nat. Prod. Rep. 36, 698–713 (2018).
Chook, Y. M., Ke, H. & Lipscomb, W. N. Crystal structures of the monofunctional chorismate mutase from Bacillus subtilis and its complex with a transition state analog. Proc. Natl Acad. Sci. USA 90, 8600–8603 (1993).
Fage, C. D. et al. The structure of SpnF, a standalone enzyme that catalyzes [4+2] cycloaddition. Nat. Chem. Biol. 11, 256–258 (2015).
Zaitseva, J., Lu, J., Olechoski, K. L. & Lamb, A. L. Two crystal structures of the isochorismate pyruvate lyase from Pseudomonas aeruginosa. J. Biol. Chem. 281, 33441–33449 (2006).
Shipman, L. W., Li, D., Roessner, C. A., Scott, A. I. & Sacchettini, J. C. Crystal structure of precorrin-8x methyl mutase. Structure 9, 587–596 (2001).
Zheng, Q. et al. Enzyme-dependent [4+2] cycloaddition depends on lid-like interaction of the N-terminal sequence with the catalytic core in Pyrl4. Cell Chem. Biol. 23, 352–360 (2016).
Byrne, M. J. et al. The catalytic mechanism of a natural Diels–Alderase revealed in molecular detail. J. Am. Chem. Soc. 138, 6095–6098 (2016).
Cogan, D. P. et al. Structural insights into enzymatic [4+2] aza-cycloaddition in thiopeptide antibiotic biosynthesis. Proc. Natl Acad. Sci. USA 114, 12928–12933 (2017).
Zheng, Q. et al. Structural insights into a flavin-dependent [4+2] cyclase that catalyzes trans-decalin formation in pyrroindomycin biosynthesis. Cell Chem. Biol. 25, 718–727 (2018).
Newmister, S. A. et al. Structural basis of the Cope rearrangement and cyclization in hapalindole biogenesis. Nat. Chem. Biol. 14, 345–351 (2018).
Ohashi, M. et al. SAM-dependent enzyme-catalysed pericyclic reactions in natural product biosynthesis. Nature 549, 502–506 (2017).
Liscombe, D. K., Louie, G. V. & Noel, J. P. Architectures, mechanisms and molecular evolution of natural product methyltransferases. Nat. Prod. Rep. 29, 1238–1250 (2012).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Newmister, S. A. et al. Unveiling sequential late-stage methyltransferase reactions in the meleagrin/oxaline biosynthetic pathway. Org. Biomol. Chem. 16, 6450–6459 (2018).
Holm, L. & Rosenström, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, w545–w549 (2010).
Horowitz, S. et al. Conservation and functional importance of carbon–oxygen hydrogen bonding in AdoMet-dependent methyltransferases. J. Am. Chem. Soc. 135, 15536–15548 (2013).
Fick, R. J. et al. Water-mediated carbon–oxygen hydrogen bonding facilitates S-adenosylmethionine recognition in the reactivation domain of cobalamin-dependent methionine synthase. Biochemistry 57, 3733–3740 (2018).
Jeon, B.-S., Wang, S.-A., Ruszczycky, M. W. & Liu, H.-W. Natural [4+2]-cyclases. Chem. Rev. 117, 5367–5388 (2017).
Preiswerk, N. et al. Impact of scaffold rigidity on the design and evolution of an artificial Diels–Alderase. Proc. Natl Acad. Sci. USA 111, 8013–8018 (2014).
Minami, A. & Oikawa, H. Recent advances of Diels–Alderases involved in natural product biosynthesis. J. Antibiot. 69, 500–506 (2016).
Breslow, R. Hydrophobic effects on simple organic reactions in water. Acc. Chem. Res. 24, 159–164 (1991).
Blokzijl, W., Blandamer, M. J. & Engberts, J. B. F. N. Diels–Alder reactions in aqueous solutions. Enforced hydrophobic interactions between diene and dienophile. J. Am. Chem. Soc. 113, 4241–4246 (1991).
Lee, A. Y., Stewart, J. D., Clardy, J. & Ganem, B. New insight into the catalytic mechanism of chorismite mutase from structural studies. Chem. Biol. 2, 195–203 (1995).
Burschowsky, D. et al. Electrostatic transition state stabilization rather than reactant destabilization provides the chemical basis for efficient chorismate mutase catalysis. Proc. Natl Acad. Sci. USA 111, 17516–17521 (2014).
Warshel, A. et al. Electrostatic basis for enzyme catalysis. Chem. Rev. 106, 3210–3235 (2006).
Smith, P. J. & Wilcox, C. S. The chemistry of functional group arrays. Electrostatic catalysis and the ‘intramolecular salt effect’. Tetrahedron 47, 2617–2628 (1991).
Fried, S. D. & Boxer, S. G. Electric fields and enzyme catalysis. Annu. Rev. Biochem. 86, 387–415 (2017).
Shaik, S., Mandal, D. & Ramanan, R. Oriented electric fields as future smart reagents in chemistry. Nat. Chem. 8, 1091–1098 (2016).
Aragonès, A. C. et al. Electrostatic catalysis of a Diels–Alder reaction. Nature 531, 88–91 (2016).
Welborn, V. V., Ruiz, P. L. & Head-Gordon, T. Computational optimization of electric fields for better catalysis design. Nat. Catal. 1, 649–655 (2018).
Yu, J., Zhou, Y., Tanaka, I. & Yao, M. Roll: a new algorithm for the detection of protein pockets and cavities with a rolling probe sphere. Bioinformatics 26, 46–52 (2010).
Acknowledgements
This work was supported by the NSFC (91856202), CAS (XDB20000000) and SMSTC (18430721500, 19XD1404800) (J.Z.), the NIH (1R01AI141481) and NSF (CHE-1806581) (Y.T. and K.N.H.). Chemical characterization studies were supported by shared instrumentation grants from the NSF (CHE-1048804) and NIH NCRR (S10RR025631). The authors thank the staff of beamlines BL17U1, BL18U1 and BL19U1 of Shanghai Synchrotron Radiation Facility for access and help with the X-ray data collection. The authors also thank J. Gan for help with structure refinement and J. Long for help with ultracentrifugation sedimentation measurements. The computational resources from the UCLA Institute of Digital Research and Education (IDRE) are acknowledged. M.O. is supported by an overseas postdoctoral fellowship from The Uehara Memorial Foundation, Japan. Y.H. is a Life Sciences Research Foundation fellow sponsored by the Mark Foundation for Cancer Research. The authors thank E.B. Go for careful proofreading of the manuscript.
Author information
Authors and Affiliations
Contributions
Y.C., Y.H., M.O., K.N.H., J.Z. and Y.T. developed the hypothesis and designed the study. Y.C. and Y.H. purified and crystallized protein samples and solved the X-ray structures. M.O. and Y.H. performed in vivo and in vitro experiments. M.O. performed compound isolation and characterization. C.S.J. and M.G.-B. performed the computational analysis. All authors analysed and discussed the results. Y.C., Y.H., K.N.H., J.Z. and Y.T. prepared the manuscript. Y.C., Y.H. and M.O. contributed equally to this work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
The Supplementary Information file includes methods, Supplementary tables and Supplementary figures
Rights and permissions
About this article
Cite this article
Cai, Y., Hai, Y., Ohashi, M. et al. Structural basis for stereoselective dehydration and hydrogen-bonding catalysis by the SAM-dependent pericyclase LepI. Nat. Chem. 11, 812–820 (2019). https://doi.org/10.1038/s41557-019-0294-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-019-0294-x
- Springer Nature Limited
This article is cited by
-
A cyclase that catalyses competing 2 + 2 and 4 + 2 cycloadditions
Nature Chemistry (2023)
-
Improving D-carbamoylase thermostability through salt bridge engineering for efficient D-p-hydroxyphenylglycine production
Systems Microbiology and Biomanufacturing (2023)
-
Discovery and investigation of natural Diels–Alderases
Journal of Natural Medicines (2021)
-
Broadening the scope of biocatalytic C–C bond formation
Nature Reviews Chemistry (2020)
-
An enzymatic Alder-ene reaction
Nature (2020)