Abstract
During mouse embryonic development, pluripotent cells rapidly divide and diversify, yet the regulatory programs that define the cell repertoire for each organ remain ill-defined. To delineate comprehensive chromatin landscapes during early organogenesis, we mapped chromatin accessibility in 19,453 single nuclei from mouse embryos at 8.25 days post-fertilization. Identification of cell-type-specific regions of open chromatin pinpointed two TAL1-bound endothelial enhancers, which we validated using transgenic mouse assays. Integrated gene expression and transcription factor motif enrichment analyses highlighted cell-type-specific transcriptional regulators. Subsequent in vivo experiments in zebrafish revealed a role for the ETS factor FEV in endothelial identity downstream of ETV2 (Etsrp in zebrafish). Concerted in vivo validation experiments in mouse and zebrafish thus illustrate how single-cell open chromatin maps, representative of a mammalian embryo, provide access to the regulatory blueprint for mammalian organogenesis.








Similar content being viewed by others
Data availability
Raw sequencing data and processed data are available at GEO with accession number GSE133244. Previously published sequencing data that were re-analysed here are available under accession codes GSM1436367 and GSM1436368 (ETV2 ChIP-seq) and GSM1692843, GSM1692848 and GSM1692858 (TAL1 ChIP-seq). Processed TAL1 ChIP-seq data used in this publication is also available at http://codex.stemcells.cam.ac.uk/. Data are available in processed form for download and interactive browsing at https://gottgens-lab.stemcells.cam.ac.uk/snATACseq_E825. Cell type tracks can be explored at https://tinyurl.com/snATACseq-GSE133244-UCSC. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
All code is available upon request and at https://github.com/BPijuanSala/MouseOrganogenesis_snATACseq_2020.
References
Cao, J. et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 361, 1380–1385 (2018).
Cusanovich, D. A. et al. Multiplex single-cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Pijuan-Sala, B., Guibentif, C. & Göttgens, B. Single-cell transcriptional profiling: a window into embryonic cell-type specification. Nat. Rev. Mol. Cell Biol. 19, 399–412 (2018).
Preissl, S. et al. Single-nucleus analysis of accessible chromatin in developing mouse forebrain reveals cell-type-specific transcriptional regulation. Nat. Neurosci. 21, 432–439 (2018).
Cusanovich, D. A. et al. The cis-regulatory dynamics of embryonic development at single-cell resolution. Nature 555, 538–542 (2018).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496 (2019).
Ibarra-Soria, X. et al. Defining murine organogenesis at single-cell resolution reveals a role for the leukotriene pathway in regulating blood progenitor formation. Nat. Cell Biol. 20, 127–134 (2018).
Pijuan-Sala, B. et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566, 490 (2019).
González-Blas, C. B. et al. cisTopic: cis-regulatory topic modeling on single-cell ATAC-seq data. Nat. Methods 16, 397–400 (2019).
Bellomo, D., Lander, A., Harragan, I. & Brown, N. A. Cell proliferation in mammalian gastrulation: the ventral node and notochord are relatively quiescent. Dev. Dyn. 205, 471–485 (1996).
Ilgren, E. B. Polyploidization of extraembryonic tissues during mouse embryogenesis. Development 59, 103–111 (1980).
Anguita, E. et al. Deletion of the mouse α-globin regulatory element (HS−26) has an unexpectedly mild phenotype. Blood 100, 3450–3456 (2002).
Hay, D. et al. Genetic dissection of the α-globin super-enhancer in vivo. Nat. Genet. 48, 895–903 (2016).
Hughes, J. R. et al. Annotation of cis-regulatory elements by identification, subclassification, and functional assessment of multispecies conserved sequences. Proc. Natl Acad. Sci. USA 102, 9830–9835 (2005).
Craig, M. L. & Russell, E. S. A developmental change in hemoglobins correlated with an embryonic red cell population in the mouse. Dev. Biol. 10, 191–201 (1964).
Hanssen, L. L. P. et al. Tissue-specific CTCF–cohesin-mediated chromatin architecture delimits enhancer interactions and function in vivo. Nat. Cell Biol. 19, 952–961 (2017).
Tzouanacou, E., Wegener, A., Wymeersch, F. J., Wilson, V. & Nicolas, J.-F. Redefining the progression of lineage segregations during mammalian embryogenesis by clonal analysis. Dev. Cell 17, 365–376 (2009).
Tremblay, M., Sanchez-Ferras, O. & Bouchard, M. GATA transcription factors in development and disease. Development 145, dev164384 (2018).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Moon, K. R. et al. Visualizing structure and transitions in high-dimensional biological data. Nat. Biotechnol. 37, 1482–1492 (2019).
Ralston, A. et al. Gata3 regulates trophoblast development downstream of Tead4 and in parallel to Cdx2. Development 137, 395–403 (2010).
Nuez, B., Michalovich, D., Bygrave, A., Ploemacher, R. & Grosveld, F. Defective haematopoiesis in fetal liver resulting from inactivation of the EKLF gene. Nature 375, 316 (1995).
Parkins, A. C., Sharpe, A. H. & Orkin, S. H. Lethal β-thalassaemia in mice lacking the erythroid CACCC-transcription factor EKLF. Nature 375, 318 (1995).
Desjardins, C. A. & Naya, F. J. The function of the MEF2 family of transcription factors in cardiac development, cardiogenomics, and direct reprogramming. J. Cardiovasc. Dev. Dis. 3, 26 (2016).
Kallianpur, A. R., Jordan, J. E. & Brandt, S. J. The SCL/TAL-1 gene is expressed in progenitors of both the hematopoietic and vascular systems during embryogenesis. Blood 83, 1200–1208 (1994).
Shivdasani, R. A., Mayer, E. L. & Orkin, S. H. Absence of blood formation in mice lacking the T-cell leukaemia oncoprotein tal-1/SCL. Nature 373, 432–434 (1995).
Silver, L. & Palis, J. Initiation of murine embryonic erythropoiesis: a spatial analysis. Blood 89, 1154–1164 (1997).
Palis, J. Hematopoietic stem cell‐independent hematopoiesis: emergence of erythroid, megakaryocyte, and myeloid potential in the mammalian embryo. FEBS Lett. 590, 3965–3974 (2016).
Downs, K. M., Gifford, S., Blahnik, M. & Gardner, R. L. Vascularization in the murine allantois occurs by vasculogenesis without accompanying erythropoiesis. Dev. Camb. Engl. 125, 4507–4520 (1998).
Ng, C. E. L. et al. A Runx1 intronic enhancer marks hemogenic endothelial cells and hematopoietic stem cells. Stem Cells 28, 1869–1881 (2010).
Nottingham, W. T. et al. Runx1-mediated hematopoietic stem-cell emergence is controlled by a Gata/Ets/SCL-regulated enhancer. Blood 110, 4188–4197 (2007).
Goode, D. K. et al. Dynamic gene regulatory networks drive hematopoietic specification and differentiation. Dev. Cell 36, 572–587 (2016).
Beck, D. et al. Genome-wide analysis of transcriptional regulators in human HSPCs reveals a densely interconnected network of coding and noncoding genes. Blood 122, e12–e22 (2013).
Wilson, N. K. et al. Combinatorial transcriptional control in blood stem/progenitor cells: genome-wide analysis of ten major transcriptional regulators. Cell Stem Cell 7, 532–544 (2010).
Pinto do O, P., Kolterud, A. & Carlsson, L. Expression of the LIM-homeobox gene LH2 generates immortalized steel factor-dependent multipotent hematopoietic precursors. EMBO J. 17, 5744–5756 (1998).
Butko, E., Pouget, C. & Traver, D. Complex regulation of HSC emergence by the Notch signaling pathway. Dev. Biol. 409, 129–138 (2016).
Kothary, R. et al. A transgene containing lacZ inserted into the dystonia locus is expressed in neural tube. Nature 335, 435 (1988).
Pennacchio, L. A. et al. In vivo enhancer analysis of human conserved non-coding sequences. Nature 444, 499 (2006).
Craig, M. P. & Sumanas, S. ETS transcription factors in embryonic vascular development. Angiogenesis 19, 275–285 (2016).
Koyano-Nakagawa, N. et al. Etv2 is expressed in the yolk sac hematopoietic and endothelial progenitors and regulates Lmo2 gene expression. Stem Cells 30, 1611–1623 (2012).
Liu, F. et al. Induction of hematopoietic and endothelial cell program orchestrated by ETS transcription factor ER71/ETV2. EMBO Rep. 16, 654–669 (2015).
Wang, L. et al. Fev regulates hematopoietic stem cell development via ERK signaling. Blood 122, 367–375 (2013).
Pham, V. N. et al. Combinatorial function of ETS transcription factors in the developing vasculature. Dev. Biol. 303, 772–783 (2007).
Landry, J.-R. et al. Expression of the leukemia oncogene Lmo2 is controlled by an array of tissue-specific elements dispersed over 100 kb and bound by Tal1/Lmo2, Ets, and Gata factors. Blood 113, 5783–5792 (2009).
Göttgens, B. Establishing the transcriptional programme for blood: the SCL stem cell enhancer is regulated by a multiprotein complex containing Ets and GATA factors. EMBO J. 21, 3039–3050 (2002).
Göttgens, B. et al. The scl +18/19 stem cell enhancer is not required for hematopoiesis: identification of a 5′ bifunctional hematopoietic-endothelial enhancer bound by Fli-1 and Elf-1. Mol. Cell. Biol. 24, 1870–1883 (2004).
Fang, R.et al. Fast and accurate clustering of single cell epigenomes reveals cis-regulatory elements in rare cell types. Preprint available at https://www.biorxiv.org/content/10.1101/615179v2 (2019).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Preprint at https://www.biorxiv.org/content/10.1101/357368v1 (2018).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Sumanas, S. & Lin, S. Ets1-related protein is a key regulator of vasculogenesis in zebrafish. PLoS Biol. 4, e10 (2006).
Xue, Y. et al. A 3D atlas of hematopoietic stem and progenitor cell expansion by multi-dimensional RNA-seq analysis. Cell Rep. 27, 1567–1578.e5 (2019).
Acknowledgements
We thank B. Ren and K. Zhang for making this collaboration between the University of California San Diego and the University of Cambridge possible; I. Imaz-Rosshandler for statistical advice; T. L. Hamilton and Central Biomedical Services for technical support in embryo collection; and R. Fang for kindly providing us with the list of constitutive promoters. We also thank S. Kuan for sequencing and B. Li for bioinformatics support. We would like to extend our gratitude to the QB3 Macrolab at UC Berkeley for purification of the Tn5 transposase. B.P.-S. is funded by the Wellcome Trust 4-Year PhD Programme in Stem Cell Biology and Medicine and the University of Cambridge, UK. B.P.-S was awarded a Travelling Fellowship from The Company of Biologists (DEV–180505) to perform this study. Research in the authors’ laboratories is supported by the Wellcome, MRC, Bloodwise, CRUK and NIH-NIDDK; as well as core support grants from the Wellcome to the Wellcome-MRC Cambridge Stem Cell Institute. This work was funded as part of a Wellcome Strategic Award (105031/Z/14/Z) awarded to W. Reik, B.G., J. Marioni, J. Nichols, L. Vallier, S. Srinivas, B. Simons, S. Teichmann and T. Voet. Work at the Center for Epigenomics was supported in part by the UC San Diego School of Medicine.
Author information
Authors and Affiliations
Contributions
B.P.-S performed embryo dissections, bioinformatic analysis (both data pre-processing and biological analysis), created the website and coordinated the study. N.K.W., S.K. and F.J.C.-N. performed enhancer validation experiments. J.X. performed experiments in zebrafish. X.H. performed snATAC-seq and was assisted by B.P.-S. R.L.H. processed the ETV2 ChIP-seq dataset. O.P. performed data demultiplexing and barcode extraction. S.P. supervised the snATAC-seq experiment, sequencing and initial data pre-processing. F.L. supervised experiments in zebrafish. B.G. supervised the study. B.P.-S. and B.G. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 snATAC-seq experiment.
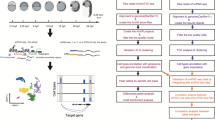
a, E8.25 embryos used for snATAC-seq. This panel includes the embryo in Fig. 1a (top right in this panel). Scale bars: 0.5 mm. Experiment was performed with 10 embryos. b, Representative FACS gating strategy. The gate used to sort the nuclei regardless of DNA ploidy can be found in the bottom left panel. Gates for nuclei with 2 (2n) and 4 copies (4n) of DNA can be found in the bottom right panel.
Extended Data Fig. 2 Data quality control and cell type annotation.
a, Quality control (QC) thresholds. Top: X-Y plot showing the number of reads in peaks and promoter coverage for each barcode. Promoter coverage is defined as the number of reads in constitutive promoters divided by the total number of constitutive promoters. Values have been log-transformed. Red square box delimits the nuclei that passed QC for these parameters. Middle: Histogram showing the doublet scores for the nuclei that passed the first QC. Red line delimits the threshold; those below the line passed QC. y axis has been log-transformed. Bottom: Histogram showing the ratio of reads in peaks for those nuclei that passed QC in the panels above. Red line delimits the threshold; nuclei above this line passed QC. b, Heatmap illustrating the row-normalised frequency (from dark blue/low to yellow/high) of nuclei for each cell type with open chromatin in the transcription start site (TSS) of genes that are expressed specifically in them. Marker gene list has been curated by using a previously reported transcriptomic atlas, containing this stage8. c, Frequency of nuclei based on their DNA content per cell type. For this plot, we only considered the nuclei sorted with the “4n” and “2n” gates from Extended Data Fig. 1b.
Extended Data Fig. 3 Transcription factor motif enrichment analyses.
a, Heatmap showing the motif enrichment scores (NES) for transcription factor (TF) motifs enriched in OCRs uniquely contributing to topics 38, 51 and/or 100. Values are represented by a colour gradient from dark blue (0) to dark red (8.9). Sequence logos are shown on the left. b, UMAP visualisation showing the motif enrichment scores for GATA1–6 using chromVAR on the 19,453 cells. Values are represented by a colour gradient from dark blue (low, below 0) to red (high, above 0). Cells with values of 0 are depicted in grey. Sequence logos for each member can be found at the bottom right corner of each plot. c, Histogram showing the number of regions containing GATA binding sites per topic.
Extended Data Fig. 4 Motif enrichment scores and sharing between gut and surface ectoderm.
a, Complete heatmap of transcription factor motif enrichment Z-scores (from blue/low/−1 to red/high/+1) showing all transcription factor (TF) names (extended from Fig. 3b). b, Barplot showing the number of cell-type-specific OCRs that are shared in a defined number of cell types, highlighted on the x axis. c, GO terms for genes associated with regions specific for surface ectoderm that are not shared with gut (n=1,018). d, GO terms for genes associated with regions specific for gut that are not shared with surface ectoderm (n=1,058). e, GO terms for genes associated with regions specific for gut and surface ectoderm that are shared between these lineages (n=227). Values obtained from one-sided hyperGTest and BH-corrected.
Extended Data Fig. 5 Allantoic-haemato-endothelial development.
a, Simplified diagram of allantoic-haemato-endothelial development. Early mesoderm generates different lineages including allantoic cells, erythrocytes and endothelial cells (ECs). The precursors of the first wave of primitive erythrocytes express genes commonly associated with ECs, thus making the distinction between erythroid and endothelial precursors (Haemato-endothelial precursors) difficult by their transcriptomes. Yolk sac (YS) ECs give rise to the definitive blood wave by generating erythro-myeloid progenitors (EMPs). The allantois also contributes to the EC pool by generating allantoic ECs. b, UMAP visualisation (left) and PAGA representation as in Fig. 7a (right) of the allantoic-haemato-endothelial landscape (n=3,284 cells) showing the chromatin accessibility at the Runx1 +23 kb enhancer. Black dots in UMAP on the right correspond to nuclei where the region is accessible. Accessibility in PAGA is represented by the ratio of nuclei per cluster (from grey to dark blue) that have Runx1 +23 kb accessible. c, PAGA representation as in Fig. 7a showing the mean enrichment scores per cluster (from grey=0 to dark blue=1) for TAL1 ChIP-seq peaks from haemangioblasts, haemogenic endothelium and haematopoietic progenitors. Subclusters for PAGA have been defined in Fig. 7a. d, Venn diagram showing the number of endothelial-specific regions from the snATAC-seq dataset, the number of TAL1-bound regions obtained by ChIP-seq in haemogenic endothelial cells from32, and their overlap. ChIP-seq peaks were taken from http://codex.stemcells.cam.ac.uk/.
Extended Data Fig. 6 Erg +85 kb and Fli1 −15 kb enhancers.
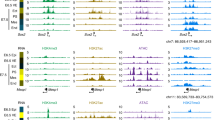
a,b, Genome browser tracks showing the Erg (a) and the Fli1 (b) loci. Black arrowheads indicate the Erg +85 kb (top) and the Fli1 −15 kb (bottom) enhancers. Tracks correspond to the snATAC-seq profiles of the erythroid, endothelium and allantois cell types after cell pooling, TAL1 ChIP-seq for haemogenic endothelial cells (“HE TAL1 ChIP-seq”, grey), H3K27ac ChIP-seq for haemogenic endothelial cells (“HE H3K27ac”, gold), TAL1 ChIP-seq for haemangioblasts (“Haem. TAL1 ChIP-seq”, grey), H3K27ac ChIP-seq for haemangioblasts (“Haem. H3K27ac”, gold), TAL1 ChIP-seq for haematopoietic progenitors (“HP TAL1 ChIP-seq”, grey), H3K27ac ChIP-seq for haematopoietic progenitors (“HP H3K27ac”, gold) from32, TAL1 ChIP-seq for HPC-7 cells (“HPC-7 TAL1 ChIP-seq”, grey) and DNase-seq for HPC-7 cells (blue). Publicly available tracks were obtained from http://codex.stemcells.cam.ac.uk/. c, UMAP visualisation of the allantoic-haemato-endothelial landscape (n=3,284 cells) showing the enrichment score (from grey/low to dark blue/high) for HPC-7 TAL1 ChIP-seq peaks.
Extended Data Fig. 7 Flt1 +67 kb and Maml3 +360 kb enhancers.
a,b, Genome browser tracks showing the Flt1 (a) and the Maml3 (b) loci. Black arrowheads indicate the Flt1 +67 kb (top) and the Maml3 +360 kb (bottom) enhancers. Tracks correspond to the snATAC-seq profiles of the erythroid, endothelium and allantois cell types after cell pooling, TAL1 ChIP-seq for haemogenic endothelial cells (“HE TAL1 ChIP-seq”, grey), H3K27ac ChIP-seq for haemogenic endothelial cells (“HE H3K27ac”, gold), TAL1 ChIP-seq for haemangioblasts (“Haem. TAL1 ChIP-seq”, grey), H3K27ac ChIP-seq for haemangioblasts (“Haem. H3K27ac”, gold), TAL1 ChIP-seq for haematopoietic progenitors (“HP TAL1 ChIP-seq”, grey), H3K27ac ChIP-seq for haematopoietic progenitors (“HP H3K27ac”, gold) from32, TAL1 ChIP-seq for HPC-7 cells (“HPC-7 TAL1 ChIP-seq”, grey) and DNase-seq for HPC-7 cells (blue). Publicly available tracks were obtained from http://codex.stemcells.cam.ac.uk/.
Extended Data Fig. 8 Evolutionary conservation of Flt1 +67 kb and Maml3 +360 kb.
Alignment of Flt1 +67 kb (a) and Maml3 +360 kb (b) across species. Transcription factor (TF) binding motifs are boxed: red: Ets sites; yellow: Gata sites; blue: E-box sites; purple box: Runx site.
Extended Data Fig. 9 Endothelial development from allantoic cells.
a, UMAP visualisation (n=3,284 cells) showing the pseudotime trajectory from allantois to endothelium as a gradient from grey to blue. Cells scored as 0 in the plot (grey) were not part of the trajectory. b, Heatmap showing the -log(P value) obtained from a TF motif enrichment analysis on the accessibility patterns found in Fig. 7b using HOMER. -log(P value) ranges from 0 (dark blue) to 311 (dark red). c, UMAP visualisation of the allantoic-haemato-endothelial landscape (n=3,284 cells) showing the enrichment score (from grey=0 to dark blue=1) for ETV2 ChIP-seq peaks from41. d, Force-directed graph showing cells from the “Mixed mesoderm”, “Allantois”, “Haemato-endothelial progenitors” and “Endothelium” that have been profiled with single-cell RNA-seq in8 (n=7,631 cells). Cell colours show the different subclusters found when re-analysing this dataset. e, Expression dynamics of highly variable ETS factors (variance > 0.15) along the trajectory from mixed mesoderm to endothelium (top) and from allantois to endothelium (bottom). Cdh5 and Pecam1 have been added as positive controls for mature endothelium. Dots below plots represent the ordered cells coloured by the subclusters in panel (d).
Extended Data Fig. 10 fev plays a role in haematopoietic and endothelial development.
a, WISH showing the expression of lmo2, tal1, flk1 and gata1a in fev mutants at 10 s. Dorsal view, anterior to the top. Red arrowheads indicate increased expression in fev+/+. b, WISH showing that the expression of fev at 12 s was increased hsp70-fev-GFP transgenic embryos after heat-shock at 3 s. c, Western blot showing the protein level of Fev increased in hsp70-fev-GFP transgenic embryos compared to control at 12 s after heat-shock at 3 s. d, WISH of lmo2, tal1, gata1a, flk1, myod and runx1 in control embryos (top) and embryos injected with hsp70-fev-GFP and tol2 mRNA and heat-shocked at 3 s (bottom). Black arrowheads indicate the expression (top) and expanded expression (bottom) in the PLPM. White arrowheads indicate expression (top) and expanded expression (bottom) in the trunk vessels. Embryos are shown on the dorsal view at 12 s stage, and the lateral view at 28 hpf. e, Genome browser tracks showing the Fev locus. Black arrowhead indicates the Fev +0.7 kb region accessible in endothelium and bound by ETV2 in vitro. Tracks correspond to the snATAC-seq profiles of the erythroid, endothelium and allantois cell types after cell pooling, the ETV2 ChIP-seq from41 and evolutionary conservation tracks from UCSC. f, UMAP visualisation (left) and PAGA representation (right) of the allantoic-haemato-endothelial landscape (n=3,284 cells) showing the chromatin accessibility at the Fev +0.7 kb region. Sub-clusters are as in Fig. 7a. Black dots in UMAP on the right correspond to nuclei where the region is accessible. Accessibility in PAGA is represented by the ratio of nuclei per cluster (from grey to dark blue) that have Fev +0.7 kb accessible. g, WISH analyses showing the expression of fev in PLPM from etsrpy11-/- mutants, and etsrpy11-/- mutants with hsp70-fev-GFP and tol2 mRNA under heat-shock treatment at 3 s. Red arrowheads highlight the area where fev is reduced in etsrpy11-/- mutants and where it is ectopically expressed after heat-shock treatment. h, Western blot analysis showing the protein level of Fev in sibling and etsrpy11-/- mutants. i, WISH of lmo2, tal1, gata1a, flk1, myod and runx1 in sibling embryos (top), etsrpy11-/- embryos (middle), and etsrpy11-/- embryos co-injected with hsp70-fev-GFP and tol2 mRNA and heat-shocked at 3 s (bottom). Black arrowheads indicate the expression (top), reduction of expression (middle) and expanded expression (bottom) in the PLPM. White arrowheads indicate these patterns in the trunk vessels. Embryos are shown on the dorsal view at 12 s stage, and the lateral view at 28 hpf. Fractions in the panels with zebrafish images depict the number of embryos that showed similar results out of the total number of embryos analysed. Full, unmodified Western blots corresponding to panels (c) and (h) can be found in the Source Data file corresponding to this figure. Scale bars: 200 µm.
Supplementary information
Supplementary Tables 1–10.
Supplementary Tables 1. Nuclear barcodes. Barcode sequences used to label nuclei in single-nucleus ATAC-seq. File tabs: I1_index_Tn5_i7: tagmentation barcodes used in the first two plates; I2_E85_embryo_all, I2_E85_embryo_smallnuclei_2n, E85_embryo_largenuclei_4n: barcodes added to nuclei in the PCR step for the sample sorted indiscriminately, the sample sorted in the “2n” gate and the sample sorted in the “4n” gate of Extended Data Fig. 1b, respectively. Supplementary Table 2. Number of reads per sequencing run. Table indicating the number of reads sequenced in each run. Each read pair from the paired-end sequencing is counted as 2 reads. Supplementary Table 3. Retained reads throughout the pre-processing pipeline. Table specifying the number of reads in the different categories highlighted in the row names. Supplementary Table 4. Constitutive promoters. List of mm10 constitutive promoters, containing the coordinates of 5,006 promoters (TSS / TSS – 2 kb) that are accessible in the majority of datasets based on ENCODE DNase Hypersensitive Sites and ATAC-seq data. This list was originally generated for ref. 4. Supplementary Table 5. Metadata for each nucleus. File containing information for each nucleus analysed in this study that passed quality control (19,453 nuclei). For each nucleus, we have provided the barcode (“barcode” column), gating based on DNA content from Extended Data Fig. 1b (“nuclei_type” column), number of reads (“num_of_reads” column), promoter coverage (“promoter_coverage” column), number of reads in promoters (“read_in_promoter” column), doublet scores (“doublet_scores” column), number of reads in peaks (“read_in_peak” column), ratio of reads in peaks (“ratio_peaks” column), UMAP coordinates (“umap_X” and “umap_Y” columns), final clusters (“final_clusters” column), cell type annotation (“ann” column) and sub-clusters for the allantoic-haemato-endothelial landscape (“al_haem_endo_clusters” column). Values for each topic are also included. Supplementary Table 6. Metadata for each genomic region. File containing information for each genomic region analysed in this study. For each genomic region, we have provided the peak ID; peak coordinates (chromosome, start and end); their general annotation (TSS (−1kb to +100 bp), TTS (−100 bp to +1 kb), intron, exon, intergenic); their distance from the TSS that have been annotated to if the region is intergenic; and the gene name, ensemble ID and strand of the genes they has been annotated to (if multiple genes have been annotated to the peak, the peak entry will be repeated). If the region is cell-type-specific, the cell type(s) where it is specific can be found in the “celltype_specificity” column. If the region contributes to a particular topic, you can find what topic(s) it contributes to in the “topic” column. “topic_stringent” gives the topic information if the regions only contribute to one topic. This table also gives information on the UMAP coordinates for visualisation in Fig. 4a, and the number of nuclei with each genomic region accessible in linear (“accessibility”) and log10 form (“accessibility_log”). If the region is part of a dynamic pattern during endothelial establishment, you will find the pattern number in the “Pattern_endothelium” column. Please note that some peak entries may be repeated due to them being annotated to multiple genes. Therefore, if one wants to plot unique regions independently of metadata regarding gene annotation, we advise to make metadata unique by using the peakID column. Supplementary Table 7. Endothelial-specific TAL1-bound open chromatin regions. File containing the coordinates of endothelial-specific open chromatin regions that intersect with TAL1 ChIP-seq peaks from haemogenic endothelium. Supplementary Table 8. Endothelial-specific heptad-bound open chromatin regions. File containing the coordinates of endothelial-specific open chromatin regions, already intersected with TAL1 ChIP-seq peaks from haemogenic endothelium, that intersect with HPC-7 ChIP-seq peaks reported as heptad peaks in ref. 34. Supplementary Table 9. Mouse transgenic assays in numbers. Number of E11.5 mouse transgenic embryos with LacZ staining in different regions (column names). FL: Fetal Liver. YS: Yolk Sac. Supplementary Table 10. Metadata for single-cell RNA-seq samples. File containing information for each cell from ref. 8 analysed in this study. For each cell (row), we have provided the cell name, cell barcode, sample stage, sequencing batch and cell type as in ref. 8. Additionally, we provide the force-directed graph coordinates computed for Fig. 7d-f and Extended Data Fig. 9e (“FA_X”, “FA_Y”), the subcluster identity (“Louvain subclust”), the pseudotime values for the allantoic-to-endothelium trajectory (“DPT_al”) and for the mesoderm-to-endothelium trajectory (“DPT_meso”).
Source data
Source Data Fig. 1
Statistical source data to reproduce figure
Source Data Fig. 2
Statistical source data to reproduce figure
Source Data Fig. 3
Statistical source data to reproduce figure
Source Data Fig. 4
Statistical source data to reproduce figure
Source Data Fig. 5
Statistical source data to reproduce figure
Source Data Fig. 7
Statistical source data to reproduce figure
Source Data Extended Data Fig. 2
Statistical source data to reproduce figure
Source Data Extended Data Fig. 3
Statistical source data to reproduce figure
Source Data Extended Data Fig. 4
Statistical source data to reproduce figure
Source Data Extended Data Fig. 5
Statistical source data to reproduce figure
Source Data Extended Data Fig. 6
Statistical source data to reproduce figure
Source Data Extended Data Fig. 9
Statistical source data to reproduce figure
Source Data Extended Data Fig. 10
Statistical source data to reproduce figure
Source Data Extended Data Fig. 10
Unprocessed western blots
Rights and permissions
About this article
Cite this article
Pijuan-Sala, B., Wilson, N.K., Xia, J. et al. Single-cell chromatin accessibility maps reveal regulatory programs driving early mouse organogenesis. Nat Cell Biol 22, 487–497 (2020). https://doi.org/10.1038/s41556-020-0489-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0489-9
- Springer Nature Limited
This article is cited by
-
An atlas of rabbit development as a model for single-cell comparative genomics
Nature Cell Biology (2023)
-
Single-cell multi-omics profiling links dynamic DNA methylation to cell fate decisions during mouse early organogenesis
Genome Biology (2022)
-
Ranking reprogramming factors for cell differentiation
Nature Methods (2022)
-
Inferring transcription factor regulatory networks from single-cell ATAC-seq data based on graph neural networks
Nature Machine Intelligence (2022)
-
Systematic reconstruction of cellular trajectories across mouse embryogenesis
Nature Genetics (2022)