Key Points
-
Chromosome segregation is a fundamental process that is necessary for the propagation of all organisms.
-
In meiosis there is a unique form of chromosome segregation in which sister chromatids co-segregate to the same pole, rather than moving apart to opposite poles as in mitosis.
-
This is facilitated by meiosis-specific changes to chromosomes, such as replacement of mitotic cohesin with meiotic cohesin and the presence of the centromeric protein shugoshin.
-
Comparative studies of meiosis and mitosis may draw a general principle that kinetochore geometry and tension exerted by microtubules synergistically generate chromosome orientation.
-
Errors in chromosome segregation in meiosis I may contribute to aneuploidy in aged oocytes.
Abstract
During mitosis, replicated chromosomes (sister chromatids) become attached at the kinetochore by spindle microtubules emanating from opposite poles and segregate equationally. In the first division of meiosis, however, sister chromatids become attached from the same pole and co-segregate, whereas homologous chromosomes connected by chiasmata segregate to opposite poles. Disorder in this specialized chromosome attachment in meiosis is the leading cause of miscarriage in humans. Recent studies have elucidated the molecular mechanisms determining chromosome orientation, and consequently segregation, in meiosis. Comparative studies of meiosis and mitosis have led to the general principle that kinetochore geometry and tension exerted by microtubules synergistically generate chromosome orientation.
Similar content being viewed by others
Main
Genetic information is packed into chromosomes, which have to be properly transmitted to replicated cells or offspring. In mitosis — the cell division observed in somatic cells — the replicated chromosomes (which consist of two sister chromatids) are split and equally divided between daughter cells through a process called equational division. For this process to be successful, the association of sister chromatids, which is known as 'cohesion' and is established during DNA replication, must be preserved until chromosome alignment, the process by which kinetochores of sister chromatids are each captured by microtubules emanating from opposite spindle poles (Box 1). Any mistake in chromosome segregation may increase the chance of tumorigenesis in humans1,2.
Eukaryotic cells have another cell division process called meiosis, which is required for sexual reproduction and reduces the chromosome number by half. Meiosis consists of two rounds of chromosome segregation following a single round of DNA replication. During the first division of meiosis (meiosis I), homologous chromosomes (homologues) rather than sister chromatids are pulled to opposite poles and segregated to the daughter cells (known as reductional division). Only at the second meiotic division (meiosis II) do sister chromatids segregate as they do in mitosis (Box 1). Thus, meiosis I has a unique form of chromosome segregation3,4.
Much has been learnt about how sister chromatids bi-orient (that is, how they are pulled towards opposite poles) in mitosis. In particular, tension across the paired kinetochores that are attached by microtubules has an essential role in stabilizing the attachment; an absence of tension leads to the release and recapture of microtubules (reorientation)5. Although meiosis must have evolved from mitosis, it is elusive how the principle that determines chromosome orientation in mitosis was modified to generate the new principle observed in meiosis, in which sister chromatids mono-orient (that is, when they move towards the same pole) and paired homologues (known as bivalents) bi-orient3,6.
Recent studies of meiosis have shed new light on the importance of kinetochore geometry in addition to the tension-dependent reorientation machinery. In this Review, I discuss the molecular mechanisms governing chromosome behaviour during meiosis, including the establishment of meiosis-specific chromosome orientation. I further discuss parallels between meiosis and mitosis and propose a common principle for determining chromosome orientation.
Kinetochore attachment to microtubules
Kinetochores are proteinaceous complexes that assemble on each chromosome and mediate their interaction with microtubules7,8. During chromosome alignment and segregation in mitosis, microtubules capture the resolved (paired but separated) sister kinetochores from opposite poles in a process known as amphitelic attachment (chromosomes are bi-oriented), and this generates full tension across centromeres (Fig. 1a). However, during the early stages of chromosome alignment, either or both sister kinetochores are transiently bound by microtubules from one pole, which is known as monotelic or syntelic attachment, respectively (chromosomes are mono-oriented) (Fig. 1a). These erroneous attachments are usually corrected into amphitelic attachment. A further complication arises from the fact that a single kinetochore binds to multiple microtubules (although this does not apply to budding yeast) so that one kinetochore can be bound simultaneously by microtubules emanating from both poles (known as merotelic attachment), which leads to undesirable chromosome bi-orientation (Fig. 1a). Because it is unlikely that merotelic attachment can be detected by surveillance mechanisms that monitor tension, it is thought that this form of attachment is the main source of aneuploidy in mammalian cells9.
Homologous chromosomes (homologues) have no interaction in mitosis, whereas in meiosis they are connected by chiasmata, forming bivalents. a | In mitosis (or meiosis II), kinetochores on sister chromatids are separated and oriented back-to-back. As a result, sister kinetochores are captured by microtubules from opposite poles, and this results in amphitelic attachment (during which chromosomes are bi-oriented). However, mono-orientation can occur if either one or both sister kinetochores are bound by microtubules from one pole, which gives rise to monotelic or syntelic attachment, respectively. If a kinetochore is bound by microtubules from both poles (merotelic attachment), chromosomes will bi-orient, but this may lead to aneuploidy. b | By contrast, in meiosis I sister kinetochores are juxtaposed or fused and orient side-by-side, thus binding to microtubules in a syntelic manner. Although bivalents are often captured from one pole (mono-orientation) during the initial stage of alignment, this tensionless attachment is so unstable that they are shifted to bi-orientation in the presence of chiasmata, which then produce tension. The univalent, which can be produced by the accidental loss of chiasmata or a recombination defect in meiosis I, is not identical to mitotic sister chromatids because sister kinetochores are fused. Such fused kinetochores often bi-orient by unstable merotelic attachment, otherwise it would be impossible to produce tension.
By contrast, during meiosis I, homologues are connected by reciprocal recombination (chiasmata), which produces paired chromosomes known as bivalents (Fig. 1b). Importantly, sister kinetochores are fused or juxtaposed side-by-side at this stage, so they tend to attach to microtubules in a syntelic manner. As a result, the bivalent is captured from opposite poles, thereby generating full tension. As in mitosis, merotelic attachment has to be avoided to ensure homologue disjunction in meiosis I (Fig. 1b). It is notable that if recombination does not occur or if chiasmata are lost (resulting in univalents), the kinetochores of sister chromatids are still fused (Fig. 1b). Consequently, and in contrast to the mitotic chromosome, univalents tend to maintain syntelic attachment, although they occasionally bi-orient by merotelic attachment. The principle kinetochore attachment to microtubules in meiosis II is thought to be virtually the same as what occurs in mitosis, as kinetochore geometry and orientation are similarly regulated in both divisions (Box 1).
Cohesion in meiotic chromosomes
Meiotic cell differentiation occurs in G1 phase of the cell cycle. In meiotic prophase, chromosomes are modified for meiosis-specific features, such as recombination and reductional division at meiosis I. Although sister chromatid cohesion mediated by cohesin is primarily required for preserving the association of paired chromosomes, accumulating evidence suggests that the cohesin complex, which constitutes the sister chromatid cohesion axis, provides a structural and functional basis for other factors that act in meiotic chromosome differentiation.
Meiosis-specific cohesin. During both mitosis and meiosis, sister chromatid cohesion is established usually during DNA replication by a multisubunit complex termed cohesin. The mitotic cohesin complex comprises two SMC (structural maintenance of chromosome) family proteins, which are termed Smc1 and Smc3, as well as two accessory subunits, which are named Rad21 (also known as kleisin subunit; Scc1 in Saccharomyces cerevisiae) and Scc3. Smc1–Smc3 heterodimers topologically embrace DNA strands through their long coiled-coil stretches. Rad21 interacts with the two ends of the cohesin ring, thereby closing it. It has been proposed that sister chromatid cohesion is established when the DNA strand is replicated while being entrapped in the cohesin ring, which thus 'glues' the sister chromatids together10,11.
Cohesin complexes are modified in meiosis. The most common modification is the replacement of Rad21 by its meiotic counterpart, Rec8 (Ref. 12), although mammals have another meiotic kleisin subunit, termed RAD21L, and Caenorhabditis elegans has COH-3 and COH-4 (Refs 13,14,15,16). Meiotic counterparts of other subunits are also reported in several organisms12,17. For example, the fission yeast protein Rec11, which is a meiosis-specific Scc3 subunit, is exclusively required for chromatid arm cohesion and recombination, whereas canonical (mitotic) Scc3 is required for centromeric cohesion in meiosis but not for recombination18. Thus, a combination of canonical and meiosis-specific cohesin subunits may diversify cohesin functions along meiotic chromosomes.
Functions of meiotic cohesin. In addition to promoting sister chromatid cohesion, cohesin complexes that include Rec8 are thought to mediate numerous processes during meiosis, such as the formation of double-strand breaks (DSBs) and synaptonemal complexes, mono-orientation of sister kinetochores and persistent cohesion throughout anaphase I17,19,20,21,22,23.
For example, cohesin complexes have been proposed to mediate homologue pairing following recombination in meiotic chromosomes (the mechanism of homologue pairing and recombination is reviewed extensively elsewhere; see Refs 24,25,26). In most organisms, loose association (pairing) and subsequent tight association (synapsis) of homologues initiate through telomere clustering at the nuclear periphery, a chromosomal configuration known as a 'bouquet'. Accompanying nuclear movement promotes the side-by-side configuration of chromosomes, thus facilitating homologue search27,28. In C. elegans, this process is promoted by specific proteins that localize at a defined chromosomal site (called the pairing centre) rather than at the telomeres29. Although DSBs and their repair process promote pairing through the recognition of homologous DNA sequences, accumulating evidence in several organisms suggests that weak interstitial pairing occurs between homologues before or independently of DSB formation25,26. How this occurs is largely unclear, but recent cytological studies in mouse germ cells suggest a model in which cohesin complexes that distribute uniquely along the chromosome axis but identically between homologues may act to recognize one another13.
The two homologues become tightly connected through sister chromatid cohesion in the distal regions (Fig. 1b) through chiasmata. This occurs through the process of reciprocal recombination between non-sister strands of two homologues. Chiasmata are essential for the capture of homologues by microtubules from opposite poles and thus for their disjunction (separation) at meiosis I (see below). In the absence of chiasmata, separated homologues undergo random segregation at metaphase I. Thus, although meiotic recombination contributes to the generation of genetic diversity, its primary aim might be the production of physical linkages between homologues, which is essential for proper segregation of homologues at meiosis I3. Some insect species have different mechanisms for establishing the association of homologues even in the absence of recombination and thereby allowing correct segregation at meiosis I30.
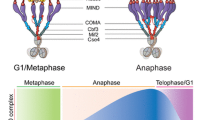
Persisting centromeric cohesion by shugoshin. In mitosis, sister chromatid cohesion is established in S phase and maintained until metaphase, when the sister chromatids are aligned by the balance of polewards pulling forces and sister chromatid cohesion. For the onset of anaphase, the APC/C (anaphase-promoting complex; also known as the cyclosome) triggers the degradation of securin, an inhibitory chaperone for the specific endopeptidase separase (also known as separin), which cleaves cohesin that includes Rad21 and removes cohesion along the entire chromosome10,11 (Box 1). This removal of cohesin triggers chromosome movement to opposite poles.
In meiosis, Rec8 largely replaces Rad21 and is cleaved only along the chromosome arms at anaphase I31,32. By contrast, centromeric Rec8 must be protected from cleavage because the residual cohesion at the centromeres is used to ensure equational division at meiosis II (see below) (Box 1). This protection of Rec8 at anaphase I is mediated by the conserved centromeric protein family shugoshin (SGO; which means guardian spirit in Japanese). Shugoshin was initially identified in the Drosophila melanogaster meiotic mutant MEI-S332, which shows precocious sister centromere separation in meiosis33,34 and was later rediscovered as a protector of Rec8 through functional screening in yeast35,36,37. Shugoshin proteins are conserved from yeast to plants and mammals38,39,40,41,42.
In yeast Sgo1 forms a complex with protein phosphatase 2A (PP2A), and the Rec8-mediated protection by Sgo1 depends on the recruitment of PP2A to the centromeres43,44. Recent studies in budding and fission yeast show that Rec8 phosphorylation by casein kinase 1 (CK1) and by cell division control protein 7 (Cdc7) is a prerequisite for the cleavage of Rec8 by separase45,46,47. Together, these findings have suggested that Sgo1–PP2A antagonizes Rec8 phosphorylation and thus its cleavage at centromeres in meiosis I (Fig. 2). Consistent with this, inactivation of Sgo1 or PP2A leads to the loss of sister chromatid cohesion during anaphase I; however, sister chromatids still co-segregate, which indicates that monopolar attachment is completed before the onset of anaphase I (Box 1). Consequently, chromosome segregation defects in Sgo1-depleted cells become visible mainly in meiosis II, as the premature separation of sister chromatids results in random segregation in the ensuing anaphase II35,36.
In meiosis I, centromeric cohesin carrying Rec8 is protected from separase cleavage. For Rec8 to be cleaved by separase, it must be phosphorylated by casein kinase 1 (CK1) in yeast. Shugoshin 1 (Sgo1) recruits protein phosphatase 2A (PP2A) to the pericentric heterochromatin regions and antagonizes this phosphorylation, thus preventing the cleavage of pericentric cohesin.
Mammals have two SGO-like proteins, SGOL1 and SGOL2. As centromeric cohesin is protected not only during meiosis I but also during prophase of mitosis, it is thought that SGOL1–PP2A may solely interact with and dephosphorylate mitotic cohesin and thereby prevent cohesin dissociation during prophase48,49,50, whereas SGOL2–PP2A protects meiotic cohesin from cleavage during anaphase I40,41. In human cells, SGOL2 has an auxiliary role in mitotic protection, presumably by supplying sufficient PP2A to centromeres43,51. Moreover, SGOL2 has a specific role in recruiting the microtubule depolymerizing mitotic centromere-associated kinesin (MCAK; also known as KIF2C) to centromeres and thereby promoting chromosome alignment at metaphase51,52,53. Recent studies in yeast and humans have shown that shugoshin proteins also act as a conserved centromeric adaptor of the Aurora B complex (see below).
Kinetochore geometry in meiosis
The importance of kinetochore orientation during mitosis has been long recognized and studied in several organisms54. Electron microscopy analyses of prometaphase chromosomes show that in mitosis, sister kinetochores face opposite directions before attachment by microtubules55. Ideally, this back-to-back assembly of sister kinetochores may facilitate the attachment of the two kinetochores to opposite poles by geometric restriction. In contrast to mitosis, sister kinetochores in meiosis I are fused or juxtaposed side-by-side, which may facilitate monopolar attachment54,56,57 (Figs 1, 3a). The molecular mechanism that regulates kinetochore geometry has recently been elucidated mainly in yeasts.
a | The distinct kinetochore geometries of mitotic and meiotic (bivalent) chromosomes in mice. Centromere protein C (CENPC) is labelled in red, and chromosomes are shown in green. b | Studies in fission yeast reveal that in mitosis there is no cohesion at the core centromere, and Rad21-containing cohesin accumulates in pericentric heterochromatin. By contrast, cohesion at the core centromere is established in meiosis I, and this depends on meiotic cohesin Rec8 and the meiotic kinetochore protein Moa1 (monopolar attachment protein 1). It is unknown how Moa1 regulates Rec8. During anaphase I, core centromeric Rec8 is likely to be cleaved by separase because the core centromeric cohesion is disrupted by prophase II, and only pericentric cohesin is protected by shugoshin75,134 (Fig. 2). The side panels indicate the orientation of microtubule attachment to kinetochores. Image in part a is courtesy of J. Kim, University of Tokyo, Japan.
Mono-orientation in budding yeast. Genetic studies in budding yeast have identified a protein complex called monopolin, which is required for mono-orientation of sister kinetochores in meiosis I58,59,60. The monopolin complex is composed of Csm1 (chromosome segregation in meiosis protein 1), Lrs4 (loss of rDNA silencing protein 4), Mam1 (monopolar microtubule attachment during meiosis I protein 1) and CK1 and localizes to centromeres specifically in meiosis I. Spo13 (sporulation-specific protein 13), another factor that is required for mono-orientation, associates with the Polo-like kinase Cdc5 and acts to recruit or stabilize the monopolin complex at centromeres61,62,63. Cdc7 kinase, which is known to be essential for the initiation of DNA replication, also acts to localize monopolin at centromeres63. Evidence suggests that the enrichment of CK1 activity at kinetochores, which depends on the presence of Csm1–Lrs4, might be an ultimate requirement for the establishment of mono-orientation in budding yeast60. However, which CK1 substrates are required for mono-orientation and the mechanism by which this occurs remain elusive.
In fission yeast, homologues of Csm1 and Lrs4 (termed Pcs1 and Mde4, respectively) are required to prevent merotelic attachment of single kinetochores in mitosis rather than to prevent bi-orientation of sister kinetochores in meiosis64. Therefore, it was hypothesized that the Csm1–Lrs4 complex has a conserved role in clamping together adjacent microtubule attachment sites to mediate mono-orientation. In budding yeast (in which paired sister centromeres assemble a single kinetochore and bind only one microtubule) this would involve conjoining two microtubule attachment sites on sister kinetochores, but in fission yeast there are multiple microtubule attachment sites on each kinetochore, so these multiple attachment sites would be joined rather than paired sister kinetochores. The structural analysis of Csm1–Lrs4 suggests that this complex forms a V-shape with two pairs of kinetochore-binding domains, indicating that Csm1–Lrs4 may bring kinetochores together, which favours the clamp model65. In fission yeast, however, the Csm1–Lrs4 complex can be functionally replaced by the condensin complex, which can compact chromatin and is enriched to kinetochores in a manner that depends on the presence of Csm1–Lrs4 (Ref. 66). Thus, the V-shaped clamp model is debatable.
Mono-orientation in fission yeast. The mechanisms that establish monopolar attachment of kinetochores during meiosis I seem to have diverged between budding yeast and fission yeast. For example, fission yeast Pcs1–Mde4 is dispensable for sister kinetochore mono-orientation (see above), and the fission yeast CK1 homologue is not required for the canonical mono-orientation pathway (T. Sakuno and Y.W., unpublished observations). Moreover, although cohesin has a crucial role in sister kinetochore mono-orientation in fission yeast (see below), this might be not the case in budding yeast60,67.
A pioneering study in this field postulated that fission yeast cohesin complexes regulate kinetochore orientation, as rec8 mutation causes equational, rather than reductional, division in meiosis I68. Like metazoan centromeres, centromeres in fission yeast comprise two domains, a kinetochore-assembling core centromere and heterochromatic pericentric regions (also known as the inner centromere in metazoans), and a single kinetochore can attach to several microtubules69. Intriguingly, in mitosis the cohesin complex that includes Rad21 accumulates preferentially at the pericentric heterochromatin regions, whereas in meiosis the Rec8-containing cohesin complex accumulates at the core centromere in addition to the pericentric regions68 (Fig. 3b). Removing Rec8 and replacing it with Rad21 during meiosis I results in accumulation of the Rad21-containing cohesin complex at the pericentric regions, but much less at the central core region, leading to bi-orientation of sister chromatids, as occurs during mitosis70. Importantly, inactivating Rec8 specifically at the core centromere while preserving its other functions leads to kinetochore bi-orientation at meiosis I, which confirms the crucial role of Rec8 for sister kinetochore mono-orientation71 (Fig. 3b).
Genetic screening to search for factors that regulate mono-orientation has further identified a meiosis-specific kinetochore protein, termed Moa1 (monopolar attachment protein 1). Moa1 interacts with Rec8 and localizes exclusively at the core centromere from prophase I to metaphase I but disappears in anaphase I71. Moa1 may assist Rec8 after pre-meiotic DNA replication to maintain the core centromere cohesion72, although its precise molecular function remains elusive.
Cohesion regulates mono-orientation in higher organisms. REC8 has been shown to have an essential role in sister kinetochore mono-orientation in meiosis I in plants and nematodes, as REC8 mutants in these organisms exhibit equational chromosome segregation at meiosis I, as observed in fission yeast16,73,74. In D. melanogaster, the orientation disruptor (ord) mutant, which has impaired sister chromatid cohesion, also exhibits a mono-orientation defect in meiosis I33. It was recently shown that inactivation of REC8 only at the kinetochore in mouse oocytes leads to bi-orientation of univalents in meiosis I (K. Tachibana-Konwalski and K. Nasmyth, personal communication). Thus, it seems that cohesion at the kinetochore mediated by the REC8-containing cohesin complex has an essential and conserved role in sister chromatid mono-orientation in meiosis I in higher organisms, including mammals.
Kinetochore geometry regulates chromosome orientation. To examine the role of cohesion at the centromere, and therefore establish the role of kinetochore geometry in chromosome orientation, the core centromere was visualized in fission yeast by looping out and excising this DNA region from the neighbouring chromosomal domains during prophase I, the stage before the attachment of kinetochores to the spindle microtubules. This analysis revealed that cohesion at the core centromere is indeed established and maintained, particularly during meiosis I, whereas it is lost in mutants that lack moa1 or rec8 and that are defective in sister kinetochore mono-orientation75 (Fig. 3b). Importantly, although pericentric cohesion is preserved throughout meiosis I until metaphase II in an Sgo1-dependent manner, cohesion at the central core region is resolved before prophase II, which is likely to be dependent on Rec8 cleavage by separase (note that Sgo1 localizes only at the pericentric heterochromatic regions but not at the core centromere in meiosis I (Fig. 2).
Remarkably, cohesin is intrinsically prevented at the core centromere in normal mitotic cells, and this is still true even if Rec8-containing cohesion is ectopically expressed in mitosis and localized to the central core region75. This finding is consistent with the notion that the establishment of cohesion at the core centromere requires not only Rec8 but also Moa1, even in meiosis I71. Finally, when an artificial tether is introduced at the core centromere, monopolar attachment and, therefore, co-segregation of sister kinetochores are restored in moa1-mutant or rec8-mutant meiotic cells and even established in mitotic cells, albeit abortively75. Thus, it seems that sister kinetochore mono-orientation is achieved by joining together DNA duplexes at the core centromeres and not by the action of a kinetochore protein itself.
Aurora B stabilizes chromosome orientation
Although the orientation of a kinetochore–microtubule attachment might be influenced by the geometry or the shape of kinetochores, erroneous attachment is commonly observed in the initial stage of chromosome alignment in mitosis and is corrected before anaphase. Tension has an important role in chromosome attachment correction, as only tension-exerted attachment is stabilized during chromosome alignment. Recent findings suggest that this may also be true in meiosis.
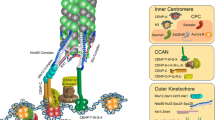
Lessons from the mitotic chromosome. In mitosis, centromeric cohesion produces tension by counteracting the pulling force of microtubules that capture sister kinetochores from opposite poles, and this tension has a crucial role in stabilizing attachment; indeed, monopolar attachment is highly unstable owing to the lack of tension (Fig. 1a). Studies in several organisms have established that the trial-and-error process of kinetochore–microtubule attachment relies largely on the mitotic kinase Aurora B (also known as Ipl1 in S. cerevisiae and Ark1 in S. pombe)5,76,77. Aurora B forms a complex with the regulatory subunits borealin, survivin and inner centromere protein (INCENP). This complex is called chromosomal passenger complex (CPC) and localizes at the centre between two sister kinetochores (known as the inner centromere) from prometaphase until metaphase78. In the absence of tension across centromeres (during monopolar attachment), Aurora B localizes in close proximity to the kinetochores, whereas bipolar (amphitelic) attachment produces tension that brings about the spatial separation of kinetochores from Aurora B79 (Fig. 4). Increasing evidence suggests that if Aurora B is close to kinetochores, Aurora B phosphorylates the microtubule-anchoring kinetochore proteins of the KMN network (comprising KNL1 (kinetochore null protein 1; also known as Spc105 in yeast), the MIS12 (also known as Mtw1 in yeast) complex and the NDC80 complex) and destabilizes the attachment80,81. By contrast, when kinetochores are separated from Aurora B under tension, overall kinetochore phosphorylation is suppressed, which stabilizes the attachment (Fig. 4).
a | Aurora B localizes at the inner centromeres in mitotic chromosomes and produces a phosphorylation gradient (dark and light blue areas), thus phosphorylating kinetochore proteins. During prometaphase, kinetochores are captured by microtubules, often in an erroneous manner (monotelic attachment, syntelic attachment or merotelic attachment; Fig. 1a). In either case, the attachment sites are in close proximity to Aurora B activity, which peaks at the inner centromeres, so attachment becomes unstable. However, once two kinetochores are captured from opposite poles, the tension that is generated across the centromeres moves the attachment sites away from Aurora B, thus selectively stabilizing the attachment. b | The KMN (kinetochore null protein 1 (KNL1)–MIS12 complex–NDC80 complex) network at kinetochores is conserved in eukaryotes and binds directly to microtubules7,135. A gradient of Aurora B phosphorylation that emanates from the inner centromere phosphorylates the KMN network, thereby destabilizing the attachment. When tension is present, the kinetochore is positioned away from Aurora B, and this facilitates the association of protein phophatase 1 (PP1) with the KMN component KNL1 (Ref. 82). Tension also acts to redistribute PP2A from the inner centromere to the kinetochore side84. Thus, the increased distance from Aurora B and the enrichment of phosphatase activity may act cumulatively to dephosphorylate the KMN network and thereby stabilize microtubule attachment.
This suppression of phosphorylation is mediated not only by the separation of the substrates from Aurora B but also by the action of PP1, which is enriched at kinetochores when tension is exerted82. Indeed, either artificially tethering the Aurora B complex to the kinetochore or depleting PP1 from kinetochores dramatically destabilizes kinetochore–microtubule attachment in prometaphase or metaphase, and, consistently, inhibiting Aurora B or tethering PP1 at kinetochores stabilizes the attachment79,82,83. Moreover, a recent study revealed that centromeric PP2A, which redistributes in response to tension, also acts in silencing the phosphorylation of Aurora B substrates at kinetochores84.
Owing to the correction mechanism that occurs during kinetochore–microtubule attachment, the position of Aurora B within the centromere is crucial for establishing proper chromosome orientation. Recent studies in fission yeast and humans indicated that shugoshin proteins — which associate with nucleosomes that contain histone H2A phosphorylated at Ser121 (and H2A phosphorylated at Thr120 in humans)85, a modification mediated by BUB1— act as a centromeric adaptor of the CPC86,87,88. Because shugoshin proteins associate directly with a regulatory subunit of the CPC, both BUB1 and shugoshin proteins are required for the full localization of Aurora B at the centromeres. It is possible that the original role of shugoshin proteins was to recruit Aurora B or to establish chromosome bi-orientation, because fission and budding yeast shugoshin proteins retain only this role during mitosis86,89. The function of cohesin protection might have co-evolved for the generation of robust tension by counteracting the spindle force, thus improving the fidelity of chromosome bi-orientation.
Studies in fission yeast, Xenopus laevis and human cells have shown that, in addition to BUB1 and shugoshin proteins, haspin kinase is required for targeting Aurora B to centromeres90,91,92. Haspin phosphorylates histone H3 at Thr3 (H3T3), and survivin binds directly to phosphorylated H3T3 (Ref. 93). Thus, targeting of Aurora B to centromeres depends on two histone marks — phosphorylated H3T3 and H2AS121 (H2AT120 in humans) (Box 2). Because haspin localizes at the sister chromatid cohesion sites and BUB1 localizes at kinetochores, the Aurora B–shugoshin complex is most enriched at the intersection of the two histone marks, the site known as the inner centromere92 (Box 2). This is defined by the network comprising the kinetochore–BUB1–phosphorylated H2AS121 pathway and the cohesin–haspin–phosphorylated H3T3 pathway, which I propose to call the inner centromere–shugoshin (ICS) network.
In addition to the overall distance of the kinetochore from Aurora B (or inter-kinetochore distance), local tension exerted at the attachment sites, which is sensitively reflected by the intra-kinetochore stretch94,95, may also contribute to the stabilization of attachment96. Thus, lack of tension is translated to a loss of attachment, which can directly activate the spindle assembly checkpoint (SAC) — a signalling system that blocks cell cycle progression into anaphase if incorrect attachment is present97,98,99.
Tension stabilizes bi-orientation of bivalents in meiosis. During meiotic prophase I, homologues are held together by chiasmata, which are produced by reciprocal recombination between maternal and paternal chromatids, and this results in paired homologues, known as a bivalent (Fig. 1b). Because sister kinetochores are fused before chromosome alignment, tension is produced when a bivalent is captured by microtubules from opposite poles. Seminal experiments in grasshopper spermatocytes suggest that, as in mitosis (see above), tension is a fundamental requirement for stabilizing the bi-orientation of bivalents100. By using a micro-needle, the author exquisitely manipulated the bivalent to observe the process of chromosome alignment in meiosis I. This study demonstrated that monopolar attachment of a bivalent is not stable, so the bivalent reorients, which gives rise to bipolar attachment. Strikingly, the induced monopolar attachment of a bivalent could be stabilized by manually pulling the bivalent in the opposite direction to generate tension101, which indicates that tension is the trigger to stabilize the attachment of bivalents.
So how does tension stabilize bivalent orientation? Similarly to mitosis, genetic studies have shown that the Aurora B homologues in yeast, Ipl1 and Ark1, have an essential role in chromosome orientation in meiosis. Indeed, their inactivation stabilizes erroneous attachment and leads to SAC silencing67,102. Recent studies using Aurora kinase inhibitors indicate that this is also true in mouse oocytes103,104,105,106,107, although Aurora C (which is a meiosis-specific Aurora kinase) may largely replace the Aurora B function in mammalian meiosis107. Precise measurements of the position of Aurora B combined with inspection of kinetochore behaviour in fission yeast have provided mechanistic insights into how the monopolar attachment of fused sister kinetochores is stabilized in bivalents108. When bivalents bi-orient at metaphase I, the kinetochores localize at the very edge of the chromosome and oscillate along the spindle coordinately (but keeping kinetochore distance constant), whereas Aurora B always localizes inwards from the kinetochores. In bivalents, fused sister kinetochores can be separated from the Aurora B sites only when they are attached and pulled in the same direction and the other half-bivalent that is connected by chiasmata is pulled in the opposite direction (Fig. 5, bivalent). Thus, the monopolar attachment of sister kinetochores is stabilized by the bipolar attachment of the bivalent.
In meiosis I, sister kinetochores are juxtaposed or fused because of the specialized sister chromatid cohesion that is established at the core centromeres (Fig. 3). This geometry positions Aurora B (blue) next to paired kinetochores (red), rather than between them, as observed in mitosis. The initial attachment of fused kinetochores is often merotelic and unstable. This unstable status persists in univalents because the meiotic kinetochore geometry prevents the amphitelic attachment that is usually observed in mitosis and that can position attachment sites far from Aurora B. Only in the presence of chiasmata does the bi-orientation of bivalents change this unstable attachment of fused kinetochores into stable syntelic attachment. Immunofluorescence images show Aurora B (green), centromere protein C (CENPC; shown in magenta) and microtubules (blue) of fission yeast cells (three chromosomes in a haploid genome) in mitosis (left) and meiosis I (right), the latter in rec12 mutant and wild-type cells). Images are reproduced, with permission, from Ref. 108 © (2011) Elsevier.
In both meiosis I and II, the Aurora B yeast homologues (or Aurora C in mammals) are localized at the inner centromeres as in mitosis108,109, and their localization partly depends on shugoshin proteins and Bub1, at least in fission yeast86,102. Thus, the ICS network may have a crucial role in defining the localization of Aurora B in meiosis as well. The involvement of PP1 or PP2A in the stabilization of kinetochore–microtubule attachment has not been well defined in meiosis.
Behaviour of univalents. If homologues are not properly paired at prophase I through chiasmata, each chromatid (univalent) shows unstable attachment because the side-by-side configuration of sister kinetochores may force their mono-orientation (Fig. 1b). In this case, because of the lack of chiasmata, Aurora B localizes close to the fused kinetochores of the univalent, which randomly move along the spindle, indicative of unstable attachment (Fig. 5, univalent). Thus, the fused kinetochores of a univalent cannot stabilize any attachment in principle.
However, merotelic attachment is somehow stabilized during prolonged metaphase, thereby leading to evasion of the SAC108. For example, in trisomic maize, the univalent that is excluded from the exchange with the other two homologues often undergoes equational separation of sister kinetochores in meiosis I110. Similarly, in XO oocytes, which carry only one X chromosome, equational segregation of the X univalent at anaphase I reaches 32%111. Moreover, Mlh1-deficient mice, in which meiotic recombination is initiated but not repaired properly, produce substantial numbers of univalents that largely exhibit unstable attachment, although some establish bi-orientation at metaphase I112. Thus, univalents often, but not always, establish bi-orientation. Analysis of the fission yeast rec12 (a homologue of budding yeast SPO11) mutant, which is defective in recombination, reveals that a predominant population of univalents exhibits unstable bi-orientation of fused kinetochores in a merotelic manner108,113 (Fig. 5). This merotelic bipolar attachment is unstable, but is somehow stabilized after extended metaphase and finally evades the SAC, leading to anaphase. Because the protection of cohesion at the centromeres by shugoshin proteins is robust in fission yeast, unlike in oocytes, most merotelic attachment of univalents ends with the co-segregation of sister chromatids at anaphase I despite exhibiting transient splitting of sister kinetochores, whereas homologues segregate randomly108,113. Although the precise mechanism of this late stabilization of merotelic attachment remains elusive, one plausible explanation is that repeated challenge by microtubules might impair kinetochore rigidity or geometry, and this leads to the excessive extrusion of the attachment sites from the Aurora B sites108.
Univalents that are produced as a consequence of recombination defects often lack the synaptonemal complex, which is usually formed between paired homologues. However, mouse oocytes in which Mlh1 has been deleted achieve full synaptonemal complex formation between homologues in prophase I, but the resultant univalents still bi-orient112. Thus, lack of the synaptonemal complex might not be the reason of the aberrant orientation of univalents. Moreover, in the fission yeast rec12 mutant, cohesion at the core centromere is preserved, which indicates that kinetochore geometry is kept intact108 (Fig. 1b). Thus, recombination components or the synaptonemal complex might not be directly involved in the regulation of kinetochore orientation of univalents. Crucially, bi-orientation of univalents in fission yeast rec12-mutant cells is completely suppressed by introducing an artificial reciprocal exchange of sister strands between two univalents108. Thus, monopolar attachment is indeed established, and this depends on the physical linkage between homologues.
Aneuploidy in aged oocytes
In mammals, oocytes start to undergo meiosis in the fetal ovary and then enter a prophase I arrest following the formation of bivalent chromosomes. Remarkably, this arrest lasts until ovulation, which is an interval of many months in mice but several decades in humans, and this potentially explains why maternal age-related miscarriage and birth defects originate predominantly from chromosome segregation errors during meiosis I114. The precise mechanism of this failure has been elusive, although age-related increases in univalents have been observed in both female mice and humans115,116.
The first mouse genetic mutant with enhanced age-related oocyte defects was obtained by deleting meiosis-specific cohesin Smc1b, which suggests that deficient cohesion is an underlying cause of age-related aneuploidy117,118. Mice that are heterozygous for Bub1 also show an age-dependent increase in aneuploidy119. Recent studies of old mice have revealed that the localization of REC8-containing cohesin is much lower in the chromosomes of aged oocytes120,121, leading to weakened sister chromatid cohesion and an increase in the incidence of univalents. Notably, the distance between sister kinetochores increases even in bivalents, suggesting that dysfunction of kinetochore geometry also contributes to the abnormal chromosome segregation in meiosis I120. Aged oocytes do not only show decreased levels of cohesin but also of its protector SGOL2 (Ref. 121), which, as in fission yeast, may cause a synergistic defect in chromosome segregation at meiosis I108. Recent studies suggest that cohesin turnover in oocytes is scarcely detectable during the growing phase and even several months after birth, and these results support the hypothesis that cohesin loaded onto chromosomes only before birth has to maintain the cohesion of bivalents and is lost in an age-related manner122,123.
In addition to weakened cohesion, the SAC might be a risk factor for aneuploidy in oocytes. Indeed, misalignment of one or a few chromosomes is occasionally neglected by the SAC in oocytes111,112, presumably because oocytes have a larger cell volume per chromosome than mitotic cells, in which SAC activation at a single kinetochore is sufficient to generate an activation signal that is transduced to the rest of the cell. Moreover, univalents, which usually show unstable attachment, occasionally establish merotelic bipolar attachment, which also contributes to the evasion of the SAC108,124. Thus, it is likely that age-related impairment of cohesion leads to defects in chiasmata, kinetochore geometry and centromeric protection. These defects and the intrinsic weakness of the SAC may contribute synergistically to aneuploidy in aged oocytes.
Conclusions and perspectives
Kinetochore are captured by the spindle microtubules and have a fundamental role in chromosome segregation. In mitosis, sister chromatid cohesion is tightly established at the pericentric region but not at the core centromere, thus promoting the separation of assembled sister kinetochores and bi-orientation at metaphase. By contrast, in meiosis I there is cohesion at the core centromere, which promotes monopolar attachment of sister kinetochores. Therefore, attachment is stabilized only when the bivalent is captured from opposite poles, which exerts full tension at the kinetochores. This principle might have been modified from that in mitosis, in which tension is produced in sister kinetochores that are captured from opposite poles. It has become increasingly clear that Aurora B kinase plays a central part in monitoring the tension that is exerted on kinetochores, promoting the re-orientation of incorrect (tensionless) attachment in both mitosis and meiosis.
Although kinetochore geometry might influence, at least in part, the initial access of microtubules to kinetochores, it has been shown that the mitotic kinetochores in rat kangaroo cells — when attached in a syntelic manner (this occurs frequently in normal prometaphase) — tend to remain side-by-side; they return to a back-to-back orientation only by the external force of microtubules125. Thus, kinetochore structure might be plastic rather than elastic, suggesting that the access of microtubules is not always properly biased by kinetochore geometry. Moreover, in the early stage of prometaphase I, fused sister kinetochores frequently attain merotelic attachment, which is challenged over time and corrected depending on the presence of chiasmata103,108 (Fig. 5). These observations indicate that kinetochore geometry plays a crucial part at the reorientation step rather than at the initial attachment step. The initial attachment would be relatively unstable, as the number of attached microtubules is not sufficient to produce full tension, but this premature tension may transiently define the relative position of Aurora B and microtubule attachment sites within the centromeres. Only favourable attachment that is located away from Aurora B would be stabilized, whereas unfavourable attachment that is close to Aurora B would be destabilized and re-oriented. This model seems applicable to both mitosis and meiosis (Figs 4,5) and supports the notion that chromosome orientation is determined by kinetochore geometry and tension exerted by microtubules. Recent developments in live-cell imaging enabled complete kinetochore tracking during chromosome congression in mitosis and meiosis106,126. Further progress of this technique will help to visualize the complete process of 'search and capture' on the single microtubule level and will prove or improve the above-mentioned geometry and tension model for kinetochore orientation.
Cohesion at the inner centromere (pericentric region) is integral for joining sister centromeres to ensure the back-to-back configuration of kinetochores and to promote bi-orientation. In fact, weakened centromeric cohesion triggers merotelic attachment and lagging of chromosomes at anaphase, which might be one of the major reasons for the chromosomal instability that is frequently observed in cancer cells9,127,128,129. It is notable that haspin kinase, which directly regulates the centromeric localization of Aurora B, binds to the cohesin-associating protein PDS5 and mislocalizes in cohesin mutants92 (Box 2). Thus, cohesion defects may impair not only tension across centromeres or the basis of kinetochore geometry but also the ICS network, thereby predisposing cells to chromosomal instability in multiple ways. It is reasonable to consider that the ICS network, which has a fundamental role in linking sister chromatid cohesion to chromosome orientation (Box 2), substantially contributes to the prevention of tumorigenesis or birth defects in humans. This possibility should be examined in future studies. Further work will define the basic molecular mechanisms for the regulation of chromosome segregation and will contribute to our understanding of how this miraculous system sustains the succession of life.
References
Ricke, R. M., van Ree, J. H. & van Deursen, J. M. Whole chromosome instability and cancer: a complex relationship. Trends Genet. 24, 457–466 (2008).
Gordon, D. J., Resio, B. & Pellman, D. Causes and consequences of aneuploidy in cancer. Nature Rev. Genet. 13, 189–203 (2012).
Moore, D. P. & Orr-Weaver, T. L. Chromosome segregation during meiosis: building an unambivalent bivalent. Curr. Top. Dev. Biol. 37, 263–299 (1998).
Petronczki, M., Siomos, M. F. & Nasmyth, K. Un menage a quatre: the molecular biology of chromosome segregation in meiosis. Cell 112, 423–440 (2003).
Tanaka, T. U. Kinetochore-microtubule interactions: steps towards bi-orientation. EMBO J. 29, 4070–4082 (2010).
Hauf, S. & Watanabe, Y. Kinetochore orientation in mitosis and meiosis. Cell 119, 317–327 (2004).
Cheeseman, I. M. & Desai, A. Molecular architecture of the kinetochore-microtubule interface. Nature Rev. Mol. Cell Biol. 9, 33–46 (2008).
Santaguida, S. & Musacchio, A. The life and miracles of kinetochores. EMBO J. 28, 2511–2531 (2009).
Cimini, D. et al. Merotelic kinetochore orientation is a major mechanism of aneuploidy in mitotic mammalian tissue cells. J. Cell Biol. 153, 517–527 (2001).
Uhlmann, F. A matter of choice: the establishment of sister chromatid cohesion. EMBO Rep. 10, 1095–1102 (2009).
Nasmyth, K. Cohesin: a catenase with separate entry and exit gates? Nature Cell Biol. 13, 1170–1177 (2011).
Watanabe, Y. Sister chromatid cohesion along arms and at centromeres. Trends Genet. 21, 405–412 (2005).
Ishiguro, K., Kim, J., Fujiyama-Nakamura, S., Kato, S. & Watanabe, Y. A new meiosis-specific cohesin complex implicated in the cohesin code for homologous pairing. EMBO Rep. 12, 267–275 (2011).
Lee, J. & Hirano, T. RAD21L, a novel cohesin subunit implicated in linking homologous chromosomes in mammalian meiosis. J. Cell Biol. 192, 263–276 (2011).
Herran, Y. et al. The cohesin subunit RAD21L functions in meiotic synapsis and exhibits sexual dimorphism in fertility. EMBO J. 30, 3091–3105 (2011).
Severson, A. F., Ling, L., van Zuylen, V. & Meyer, B. J. The axial element protein HTP-3 promotes cohesin loading and meiotic axis assembly in C. elegans to implement the meiotic program of chromosome segregation. Genes Dev. 23, 1763–1778 (2009).
Revenkova, E. & Jessberger, R. Keeping sister chromatids together: cohesins in meiosis. Reproduction 130, 783–790 (2005).
Kitajima, T. S., Yokobayashi, S., Yamamoto, M. & Watanabe, Y. Distinct cohesin complexes organize meiotic chromosome domains. Science 300, 1152–1155 (2003).
Klein, F. et al. A central role for cohesins in sister chromatid cohesion, formation of axial elements, and recombination during yeast meiosis. Cell 98, 91–103 (1999). The first study describing the central role of meiotic cohesin in chromosome differentiation.
Watanabe, Y. & Nurse, P. Cohesin Rec8 is required for reductional chromosome segregation at meiosis. Nature 400, 461–464 (1999). The first study to show that cohesin Rec8 is required for establishing mono-orientation of sister kinetochores.
Xu, H., Beasley, M. D., Warren, W. D., van der Horst, G. T. & McKay, M. J. Absence of mouse REC8 cohesin promotes synapsis of sister chromatids in meiosis. Dev. Cell 8, 949–961 (2005).
Kim, K. P. et al. Sister cohesion and structural axis components mediate homolog bias of meiotic recombination. Cell 143, 924–937 (2010).
Ellermeier, C. & Smith, G. R. Cohesins are required for meiotic DNA breakage and recombination in Schizosaccharomyces pombe. Proc. Natl Acad. Sci. USA 102, 10952–10957 (2005).
Neale, M. J. & Keeney, S. Clarifying the mechanics of DNA strand exchange in meiotic recombination. Nature 442, 153–158 (2006).
Gerton, J. L. & Hawley, R. S. Homologous chromosome interactions in meiosis: diversity amidst conservation. Nature Rev. Genet. 6, 477–487 (2005).
Bhalla, N. & Dernburg, A. F. Prelude to a division. Annu. Rev. Cell Dev. Biol. 24, 397–424 (2008).
Scherthan, H. A bouquet makes ends meet. Nature Rev. Mol. Cell Biol. 2, 621–627 (2001).
Yamamoto, A. & Hiraoka, Y. How do meiotic chromosomes meet their homologous partners?: lessons from fission yeast. Bioessays 23, 526–533 (2001).
Sato, A. et al. Cytoskeletal forces span the nuclear envelope to coordinate meiotic chromosome pairing and synapsis. Cell 139, 907–919 (2009).
Wolf, K. W. How meiotic cells deal with non-exchange chromosomes. Bioessays 16, 107–114 (1994).
Buonomo, S. B. et al. Disjunction of homologous chromosomes in meiosis I depends on proteolytic cleavage of the meiotic cohesin Rec8 by separin. Cell 103, 387–398 (2000).
Kitajima, T. S., Miyazaki, Y., Yamamoto, M. & Watanabe, Y. Rec8 cleavage by separase is required for meiotic nuclear divisions in fission yeast. EMBO J. 22, 5643–5653 (2003).
Goldstein, L. S. Mechanisms of chromosome orientation revealed by two meiotic mutants in Drosophila melanogaster. Chromosoma 78, 79–111 (1980).
Miyazaki, W. Y. & Orr-Weaver, T. L. Sister-chromatid cohesion in mitosis and meiosis. Annu. Rev. Genet. 28, 167–168 (1994).
Kitajima, T. S., Kawashima, S. A. & Watanabe, Y. The conserved kinetochore protein shugoshin protects centromeric cohesion during meiosis. Nature 427, 510–517 (2004). The first study to identify shugoshins as protectors of cohesin at the centromeres.
Rabitsch, K. P. et al. Two fission yeast homologs of Drosophila Mei-S332 are required for chromosome segregation during meiosis I and II. Curr. Biol. 14, 287–301 (2004).
Marston, A. L., Tham, W. H., Shah, H. & Amon, A. A genome-wide screen identifies genes required for centromeric cohesion. Science 303, 1367–1370 (2004).
Katis, V. L., Galova, M., Rabitsch, K. P., Gregan, J. & Nasmyth, K. Maintenance of cohesin at centromeres after meiosis I in budding yeast requires a kinetochore-associated protein related to MEI-S332. Curr. Biol. 14, 560–572 (2004).
Hamant, O. et al. A REC8-dependent plant shugoshin is required for maintenance of centromeric cohesion during meiosis and has no mitotic functions. Curr. Biol. 15, 948–954 (2005).
Lee, J. et al. Unified mode of centromeric protection by shugoshin in mammalian oocytes and somatic cells. Nature Cell Biol. 10, 42–52 (2008).
Llano, E. et al. Shugoshin-2 is essential for the completion of meiosis but not for mitotic cell division in mice. Genes Dev. 22, 2400–2413 (2008).
Wang, M. et al. OsSGO1 maintains synaptonemal complex stabilization in addition to protecting centromeric cohesion during rice meiosis. Plant J. 67, 583–594 (2011).
Kitajima, T. S. et al. Shugoshin collaborates with protein phosphatase 2A to protect cohesin. Nature 441, 46–52 (2006).
Riedel, C. G. et al. Protein phosphatase 2A protects centromeric sister chromatid cohesion during meiosis I. Nature 441, 53–61 (2006).
Brar, G. A. et al. Rec8 phosphorylation and recombination promote the step-wise loss of cohesins in meiosis. Nature 441, 532–536 (2006).
Ishiguro, T., Tanaka, K., Sakuno, T. & Watanabe, Y. Shugoshin-PP2A counteracts casein-kinase-1-dependent cleavage of Rec8 by separase. Nature Cell Biol. 12, 500–506 (2010).
Katis, V. L. et al. Rec8 phosphorylation by casein kinase 1 and Cdc7–Dbf4 kinase regulates cohesin cleavage by separase during meiosis. Dev. Cell 18, 397–409 (2010).
Salic, A., Waters, J. C. & Mitchison, T. J. Vertebrate shugoshin links sister centromere cohesion and kinetochore microtubule stability in mitosis. Cell 118, 567–578 (2004). The first demonstration that shugoshins protect centromeric cohesion in vertebrate mitotic cells.
McGuinness, B. E., Hirota, T., Kudo, N. R., Peters, J.-M. & Nasmyth, K. Shugoshin prevents dissociation of cohesin from centromeres during mitosis in vertebrate cells. PLoS Biol. 3, e86 (2005).
Kitajima, T. S., Hauf, S., Ohsugi, M., Yamamoto, T. & Watanabe, Y. Human Bub1 defines the persistent cohesion site along the mitotic chromosome by affecting shugoshin localization. Curr. Biol. 15, 353–359 (2005).
Tanno, Y. et al. Phosphorylation of mammalian Sgo2 by Aurora B recruits PP2A and MCAK to centromeres. Genes Dev. 24, 2169–2179 (2010).
Huang, H. et al. Tripin/hSgo2 recruits MCAK to the inner centromere to correct defective kinetochore attachments. J. Cell Biol. 177, 413–424 (2007).
Rivera, T. et al. Xenopus Shugoshin 2 regulates the spindle assembly pathway mediated by the chromosomal passenger complex. EMBO J. 31, 1467–1479 (2012).
Östergren, G. The mechanism of co-orientation in bivalents and multivalents. Hereditas 37, 85–156 (1951).
Journey, L. J. & Whaley, A. Kinetochore ultrastructure in vincristine-treated mammalian cells. J. Cell Sci. 7, 49–54 (1970).
Goldstein, L. S. Kinetochore structure and its role in chromosome orientation during the first meiotic division in male D. melanogaster. Cell 25, 591–602 (1981). This paper outlines the morphological differences between mitotic and meiotic sister kinetochores.
Hodges, C. A. & Hunt, P. A. Simultaneous analysis of chromosomes and chromosome-associated proteins in mammalian oocytes and embryos. Chromosoma 111, 165–169 (2002).
Toth, A. et al. Functional genomics identifies monopolin: a kinetochore protein required for segregation of homologs during meiosis I. Cell 103, 1155–1168 (2000). By using a genetic approach, this study identifies monopolin as a necessary complex for sister kinetochore mono-orientation in budding yeast.
Rabitsch, K. P. et al. Kinetochore recruitment of two nucleolar proteins is required for homolog segregation in meiosis I. Dev. Cell 4, 535–548 (2003).
Petronczki, M. et al. Monopolar attachment of sister kinetochores at meiosis I requires casein kinase 1. Cell 126, 1049–1064 (2006).
Katis, V. L. et al. Spo13 facilitates monopolin recruitment to kinetochores and regulates maintenance of centromeric cohesion during yeast meiosis. Curr. Biol. 14, 2183–2196 (2004).
Lee, B. H., Kiburz, B. M. & Amon, A. Spo13 maintains centromeric cohesion and kinetochore coorientation during meiosis I. Curr. Biol. 14, 2168–2182 (2004).
Matos, J. et al. Dbf4-dependent CDC7 kinase links DNA replication to the segregation of homologous chromosomes in meiosis I. Cell 135, 662–678 (2008).
Gregan, J. et al. The kinetochore proteins Pcs1 and Mde4 and heterochromatin are required to prevent merotelic orientation. Curr. Biol. 17, 1190–1200 (2007).
Corbett, K. D. et al. The monopolin complex crosslinks kinetochore components to regulate chromosome-microtubule attachments. Cell 142, 556–567 (2010).
Tada, K., Susumu, H., Sakuno, T. & Watanabe, Y. Condensin association with histone H2A shapes mitotic chromosomes. Nature 474, 477–483 (2011).
Monje-Casas, F., Prabhu, V. R., Lee, B. H., Boselli, M. & Amon, A. Kinetochore orientation during meiosis is controlled by Aurora B and the monopolin complex. Cell 128, 477–490 (2007).
Watanabe, Y., Yokobayashi, S., Yamamoto, M. & Nurse, P. Pre-meiotic S phase is linked to reductional chromosome segregation and recombination. Nature 409, 359–363 (2001).
Pidoux, A. & Allshire, R. Kinetochore and heterochromatin domains of the fission yeast centromere. Chromosome Res. 12, 521–534 (2004).
Yokobayashi, S., Yamamoto, M. & Watanabe, Y. Cohesins determine the attachment manner of kinetochores to spindle microtubules at meiosis I in fission yeast. Mol. Cell. Biol. 23, 3965–3973 (2003).
Yokobayashi, S. & Watanabe, Y. The kinetochore protein Moa1 enables cohesion-mediated monopolar attachment at meiosis I. Cell 123, 803–817 (2005).
Kagami, A. et al. Acetylation regulates monopolar attachment at multiple levels during meiosis I in fission yeast. EMBO Rep. 12, 1189–1195 (2011).
Yu, H.-G. & Dawe, R. K. Functional redundancy in the maize meiotic kinetochore. J. Cell Biol. 151, 131–141 (2000).
Chelysheva, L. et al. AtREC8 and AtSCC3 are essential to the monopolar orientation of the kinetochores during meiosis. J. Cell Sci. 118, 4621–4632 (2005).
Sakuno, T., Tada, K. & Watanabe, Y. Kinetochore geometry defined by cohesion within the centromere. Nature 458, 852–858 (2009). This study establishes the causal link between kinetochore geometry and cohesion within the centromere.
Tanaka, T. U. et al. Evidence that the Ipl1–Sli15 (Aurora kinase-INCENP) complex promotes chromosome bi-orientation by altering kinetochore-spindle pole connections. Cell 108, 317–329 (2002). This first study to show that Aurora B destabilizes microtubule–kinetochore attachment and thereby promotes chromosome bi-orientation.
Hauf, S. et al. The small molecule Hesperadin reveals a role for Aurora B in correcting kinetochore-microtubule attachment and in maintaining the spindle assembly checkpoint. J. Cell Biol. 161, 281–294 (2003).
Ruchaud, S., Carmena, M. & Earnshaw, W. C. Chromosomal passengers: conducting cell division. Nature Rev. Mol. Cell Biol. 8, 798–812 (2007).
Liu, D., Vader, G., Vromans, M. J., Lampson, M. A. & Lens, S. M. Sensing chromosome bi-orientation by spatial separation of aurora B kinase from kinetochore substrates. Science 323, 1350–1353 (2009). This study demonstrates the influence of the relative location of Aurora B and kinetochores on the stability of the microtubule–kinetochore attachment.
DeLuca, J. G. et al. Kinetochore microtubule dynamics and attachment stability are regulated by Hec1. Cell 127, 969–982 (2006).
Welburn, J. P. et al. Aurora B phosphorylates spatially distinct targets to differentially regulate the kinetochore-microtubule interface. Mol. Cell 38, 383–392 (2010).
Liu, D. et al. Regulated targeting of protein phosphatase 1 to the outer kinetochore by KNL1 opposes Aurora B kinase. J. Cell Biol. 188, 809–820 (2010).
Lampson, M. A. & Cheeseman, I. M. Sensing centromere tension: Aurora B and the regulation of kinetochore function. Trends Cell Biol. 21, 133–140 (2011).
Foley, E. A., Maldonado, M. & Kapoor, T. M. Formation of stable attachments between kinetochores and microtubules depends on the B56-PP2A phosphatase. Nature Cell Biol. 13, 1265–1271 (2011).
Kawashima, S. A., Yamagishi, Y., Honda, T., Ishiguro, K. & Watanabe, Y. Phosphorylation of H2A by Bub1 prevents chromosomal instability through localizing shugoshin. Science 327, 172–177 (2010).
Kawashima, S. A. et al. Shugoshin enables tension-generating attachment of kinetochores by loading Aurora to centromeres. Genes Dev. 21, 420–435 (2007).
Vanoosthuyse, V., Prykhozhij, S. & Hardwick, K. G. Shugoshin 2 regulates localization of the chromosomal passenger proteins in fission yeast mitosis. Mol. Biol. Cell 18, 1657–1669 (2007).
Tsukahara, T., Tanno, Y. & Watanabe, Y. Phosphorylation of the CPC by Cdk1 promotes chromosome bi-orientation. Nature 467, 719–723 (2010).
Indjeian, V. B., Stern, B. M. & Murray, A. W. The centromeric protein Sgo1 is required to sense lack of tension on mitotic chromosomes. Science 307, 130–133 (2005).
Kelly, A. E. et al. Survivin reads phosphorylated histone H3 threonine 3 to activate the mitotic kinase Aurora B. Science 330, 235–239 (2010).
Wang, F. et al. Histone H3 Thr-3 phosphorylation by Haspin positions Aurora B at centromeres in mitosis. Science 330, 231–235 (2010).
Yamagishi, Y., Honda, T., Tanno, Y. & Watanabe, Y. Two histone marks establish the inner centromere and chromosome bi-orientation. Science 330, 239–243 (2010). This study, together with references 85, 90 and 91, establishes that the inner centromere (or Aurora B position) is defined by the ICS network, which is composed of two histone phosphorylation pathways.
Jeyaprakash, A. A., Basquin, C., Jayachandran, U. & Conti, E. Structural basis for the recognition of phosphorylated histone h3 by the survivin subunit of the chromosomal passenger complex. Structure 19, 1625–1634 (2011).
Maresca, T. J. & Salmon, E. D. Intrakinetochore stretch is associated with changes in kinetochore phosphorylation and spindle assembly checkpoint activity. J. Cell Biol. 184, 373–381 (2009).
Uchida, K. S. et al. Kinetochore stretching inactivates the spindle assembly checkpoint. J. Cell Biol. 184, 383–390 (2009).
Akiyoshi, B. et al. Tension directly stabilizes reconstituted kinetochore–microtubule attachments. Nature 468, 576–579 (2010).
Pinsky, B. A. & Biggins, S. The spindle checkpoint: tension versus attachment. Trends Cell Biol. 15, 486–493 (2005).
Nezi, L. & Musacchio, A. Sister chromatid tension and the spindle assembly checkpoint. Curr. Opin. Cell Biol. 21, 785–795 (2009).
Murray, A. W. A brief history of error. Nature Cell Biol. 13, 1178–1182 (2011).
Nicklas, R. B. How cells get the right chromosomes. Science 275, 632–637 (1997).
Nicklas, R. B. & Koch, C. A. Chromosome micromanipulation. 3. Spindle fiber tension and the reorientation of mal-oriented chromosomes. J. Cell Biol. 43, 40–50 (1969). Micromanipulation experiments in meiotic cells demonstrate that the physical appliance of tension can lead to a stabilization of attachment.
Hauf, S. et al. Aurora controls sister kinetochore mono-orientation and homolog bi-orientation in meiosis-I. EMBO J. 26, 4475–4486 (2007).
Kitajima, T. S., Ohsugi, M. & Ellenberg, J. Complete kinetochore tracking reveals error-prone homologous chromosome biorientation in mammalian oocytes. Cell 146, 568–581 (2011).
Lane, S. I., Chang, H. Y., Jennings, P. C. & Jones, K. T. The Aurora kinase inhibitor ZM447439 accelerates first meiosis in mouse oocytes by overriding the spindle assembly checkpoint. Reproduction 140, 521–530 (2010).
Shuda, K., Schindler, K., Ma, J., Schultz, R. M. & Donovan, P. J. Aurora kinase B modulates chromosome alignment in mouse oocytes. Mol. Reprod. Dev. 76, 1094–1105 (2009).
Sharif, B. et al. The chromosome passenger complex is required for fidelity of chromosome transmission and cytokinesis in meiosis of mouse oocytes. J. Cell Sci. 123, 4292–4300 (2010).
Yang, K. T. et al. Aurora-C kinase deficiency causes cytokinesis failure in meiosis I and production of large polyploid oocytes in mice. Mol. Biol. Cell 21, 2371–2383 (2010).
Sakuno, T., Tanaka, K., Hauf, S. & Watanabe, Y. Repositioning of Aurora B promoted by chiasmata ensures sister chromatid mono-orientation in meiosis I. Dev. Cell 21, 534–545 (2011). This study identifies a molecular mechanism that explains how chiasmata are more likely to cause the bi-orientation of bivalents than the bi-orientation of univalents at meiosis I.
Parra, M. T. et al. A perikinetochoric ring defined by MCAK and Aurora-B as a novel centromere domain. PLoS Genet. 2, e84 (2006).
Maguire, M. P. A possible role for the synaptonemal complex in chiasma maintenance. Exp. Cell Res. 112, 297–308 (1978).
LeMaire-Adkins, R., Radke, K. & Hunt, P. A. Lack of checkpoint control at the metaphase/anaphase transition: a mechanism of meiotic nondisjunction in mammalian females. J. Cell Biol. 139, 1611–1619 (1997).
Nagaoka, S. I., Hodges, C. A., Albertini, D. F. & Hunt, P. A. Oocyte-specific differences in cell-cycle control create an innate susceptibility to meiotic errors. Curr. Biol. 21, 651–657 (2011).
Hirose, Y. et al. Chiasmata promote monopolar attachment of sister chromatids and their co-segregation toward the proper pole during meiosis I. PLoS Genet. 7, e1001329 (2011).
Hassold, T. & Hunt, P. To err (meiotically) is human: the genesis of human aneuploidy. Nature Rev. Genet. 2, 280–291 (2001).
Henderson, S. A. & Edwards, R. G. Chiasma frequency and maternal age in mammals. Nature 218, 22–28 (1968).
Angell, R. R., Xian, J., Keith, J., Ledger, W. & Baird, D. T. First meiotic division abnormalities in human oocytes: mechanism of trisomy formation. Cytogenet. Cell Genet. 65, 194–202 (1994).
Revenkova, E. et al. Cohesin SMC1β is required for meiotic chromosome dynamics, sister chromatid cohesion and DNA recombination. Nature Cell Biol. 6, 555–562 (2004).
Hodges, C. A., Revenkova, E., Jessberger, R., Hassold, T. J. & Hunt, P. A. SMC1β-deficient female mice provide evidence that cohesins are a missing link in age-related nondisjunction. Nature Genet. 37, 1351–1355 (2005).
Leland, S. et al. Heterozygosity for a Bub1 mutation causes female-specific germ cell aneuploidy in mice. Proc. Natl Acad. Sci. USA 106, 12776–12781 (2009).
Chiang, T., Duncan, F. E., Schindler, K., Schultz, R. M. & Lampson, M. A. Evidence that weakened centromere cohesion is a leading cause of age-related aneuploidy in oocytes. Curr. Biol. 20, 1522–1528 (2010).
Lister, L. M. et al. Age-related meiotic segregation errors in mammalian oocytes are preceded by depletion of cohesin and Sgo2. Curr. Biol. 20, 1511–1521 (2010).
Revenkova, E., Herrmann, K., Adelfalk, C. & Jessberger, R. Oocyte cohesin expression restricted to predictyate stages provides full fertility and prevents aneuploidy. Curr. Biol. 20, 1529–1533 (2010).
Tachibana-Konwalski, K. et al. Rec8-containing cohesin maintains bivalents without turnover during the growing phase of mouse oocytes. Genes Dev. 24, 2505–2516 (2010).
Kouznetsova, A., Lister, L., Nordenskjold, M., Herbert, M. & Hoog, C. Bi-orientation of achiasmatic chromosomes in meiosis I oocytes contributes to aneuploidy in mice. Nature Genet. 39, 966–968 (2007). The first study to suggest that bi-orientated univalents evade the SAC, thus raising the risk of aneuploidy.
Loncarek, J. et al. The centromere geometry essential for keeping mitosis error free is controlled by spindle forces. Nature 450, 745–749 (2007).
Magidson, V. et al. The spatial arrangement of chromosomes during prometaphase facilitates spindle assembly. Cell 146, 555–567 (2011).
Barber, T. D. et al. Chromatid cohesion defects may underlie chromosome instability in human colorectal cancers. Proc. Natl Acad. Sci. USA 105, 3443–3448 (2008).
Solomon, D. A. et al. Mutational inactivation of STAG2 causes aneuploidy in human cancer. Science 333, 1039–1043 (2011).
Holland, A. J. & Cleveland, D. W. Boveri revisited: chromosomal instability, aneuploidy and tumorigenesis. Nature Rev. Mol. Cell Biol. 10, 478–487 (2009).
Kiburz, B. M., Amon, A. & Marston, A. L. Shugoshin promotes sister kinetochore biorientation in Saccharomyces cerevisiae. Mol. Biol. Cell 19, 1199–1209 (2008).
Tang, Z., Sun, Y., Harley, S. E., Zou, H. & Yu, H. Human Bub1 protects centromeric sister-chromatid cohesion through Shugoshin during mitosis. Proc. Natl Acad. Sci. USA 101, 18012–18017 (2004).
Boyarchuk, Y., Salic, A., Dasso, M. & Arnaoutov, A. Bub1 is essential for assembly of the functional inner centromere. J. Cell Biol. 176, 919–928 (2007).
Yamagishi, Y., Sakuno, T., Shimura, M. & Watanabe, Y. Heterochromatin links to centromeric protection by recruiting shugoshin. Nature 455, 251–255 (2008).
Kiburz, B. M. et al. The core centromere and Sgo1 establish a 50-kb cohesin-protected domain around centromeres during meiosis I. Genes Dev. 19, 3017–3030 (2005).
Lampert, F. & Westermann, S. A blueprint for kinetochores — new insights into the molecular mechanics of cell division. Nature Rev. Mol. Cell Biol. 12, 407–412 (2011).
Acknowledgements
The author thanks S. Hauf for critically reading the manuscript and his current and previous laboratory members for discussions. The author apologizes to authors whose work was not discussed in this Review owing to space limitations. Work in the Y.W. laboratory was supported by a Grant-in-Aid for Specially Promoted Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Related links
Glossary
- Kinetochores
-
Large protein complexes that assemble on chromosomes and mediate the attachment to spindle microtubules.
- Centromeres
-
Specialized genomic regions where the kinetochore assembles.
- Synaptonemal complexes
-
Ribbon-like protein structures between pachytene chromosomes that mediate synapsis.
- APC/C
-
(Anaphase-promoting complex; also known as the cyclosome). A multicomponent ubiquitin ligase that targets proteins for degradation by the proteasome.
- Condensin
-
A major protein component of mitotic chromosomes that is required for chromosome condensation. The molecular architecture of condensin is similar to that of cohesin, in which two SMC (structural maintenance of chromosome) proteins are linked to each other at one end, and the other end is closed by a kleisin subunit.
- Pericentric heterochromatin
-
Heterochromatin that is assembled at the pericentric region and is composed of repeated specific DNA sequences.
- KMN network
-
The conserved kinetochore linker complex that directly binds to the microtubule plus end and is composed of KNL1 (kinetochore null protein 1), MIS12 and NDC80 subcomplexes.
Rights and permissions
About this article
Cite this article
Watanabe, Y. Geometry and force behind kinetochore orientation: lessons from meiosis. Nat Rev Mol Cell Biol 13, 370–382 (2012). https://doi.org/10.1038/nrm3349
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3349
- Springer Nature Limited