Abstract
Base editors (BEs) enable the generation of targeted single-nucleotide mutations, but currently used rat APOBEC1-based BEs are relatively inefficient in editing cytosines in highly methylated regions or in GpC contexts. By screening a variety of APOBEC and AID deaminases, we show that human APOBEC3A-conjugated BEs and versions we engineered to have narrower editing windows can mediate efficient C-to-T base editing in regions with high methylation levels and GpC dinucleotide content.


Similar content being viewed by others
Accession codes
References
Komor, A.C., Kim, Y.B., Packer, M.S., Zuris, J.A. & Liu, D.R. Nature 533, 420–424 (2016).
Nishida, K. et al. Science 353, 919–921 (2016).
Kim, K. et al. Nat. Biotechnol. 35, 435–437 (2017).
Zong, Y. et al. Nat. Biotechnol. 35, 438–440 (2017).
Hu, J.H. et al. Nature 556, 57–63 (2018).
Li, X. et al. Nat. Biotechnol. 36, 324–327 (2018).
Schübeler, D. Nature 517, 321–326 (2015).
Nabel, C.S. et al. Nat. Chem. Biol. 8, 751–758 (2012).
Chelico, L., Pham, P., Calabrese, P. & Goodman, M.F. Nat. Struct. Mol. Biol. 13, 392–399 (2006).
Chen, J., Miller, B.F. & Furano, A.V. Elife 3, e02001 (2014).
Carpenter, M.A. et al. J. Biol. Chem. 287, 34801–34808 (2012).
Ito, F., Fu, Y., Kao, S.A., Yang, H. & Chen, X.S. J. Mol. Biol. 429, 1787–1799 (2017).
Komor, A.C. et al. Sci. Adv. 3, eaao4774 (2017).
St. Martin, A. et al. Nucleic Acids Res. https://dx.doi.org/10.1093/nar/gky332 (2018).
Lei, L. et al. Nat. Struct. Mol. Biol. 25, 45–52 (2018).
Yang, B., Li, X., Lei, L. & Chen, J. J. Genet. Genomics 44, 423–437 (2017).
Shi, K. et al. Nat. Struct. Mol. Biol. 24, 131–139 (2017).
Mitra, M. et al. Nucleic Acids Res. 42, 1095–1110 (2014).
Wang, L. et al. Cell Res. 27, 1289–1292 (2017).
Kleinstiver, B.P. et al. Nature 523, 481–485 (2015).
Morita, S. et al. Nat. Biotechnol. 34, 1060–1065 (2016).
Stepper, P. et al. Nucleic Acids Res. 45, 1703–1713 (2017).
Acknowledgements
We are grateful to G.G. Carmichael and L.-L. Chen for critical reading of this paper, L. Lei and L. Wang for participating in the western blot and plasmid construction, and J. Liang for technical support. This work was supported by grants 31730111 (L.Y.), 91540115 (L.Y.), 31600654 (J.C.), 31600619 (B.Y.) and 31471241 (L.Y.) from the NSFC and grants 16PJ1407000 (J.C.) and 16PJ1407500 (B.Y.) from the Shanghai Pujiang Program.
Author information
Authors and Affiliations
Contributions
J.C., L.Y. and X.H. conceived, designed and supervised the project. J.C. managed the project. X.W. and J.L. performed most experiments with the help of J. Wu and R.W. on cell culture and plasmid construction. J. Wei prepared libraries for deep sequencing and Y.W. performed bioinformatics analyses, supervised by L.Y. J.C., L.Y. and B.Y. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Effect of DNA methylation on the base editing efficiency induced by BE3.
(a) DNMT3-induced DNA methylation leads to the decreased base editing efficiency induced by BE3. Top panels, DNA methylation status of indicated genomic regions from NT or dCas9-DNMT3-transfected HEK293T cells are shown. Bottom panels, BE3-induced C-to-T editing frequencies at indicated cytosines were individually determined under different conditions. (b) TET1-induced DNA demethylation increased the efficiency of BE3-mediated base editing. Top panels, DNA methylation status of indicated genomic regions from NT or dCas9-TET1-transfected HEK293T cells are shown. Bottom panels, BE3 induced C-to-T editing frequencies at indicated cytosines were individually determined under different conditions. (a,b) Target site sequences are shown with the editing window (position 4-8, setting the base distal to the protospacer adjacent motif (PAM) as position 1) in lavender, PAM in cyan and CpG site in capital. NT, native HEK293T cells with no treatment. M, methylable regions. (c) Normalized C-to-T editing frequencies at CpG sites shown in (a) and (b), setting the ones induced at low-methylation levels as 100%. (d) Statistical analysis of normalized C-to-T editing frequencies at CpG and non-CpG sites, setting the ones induced at low-methylation levels as 100%. The data analyzed are from panels (a) and (b). Left, n = 36 samples from three independent experiments; right, n = 18 samples from three independent experiments. P value, one-tailed Student’s t test. The median and interquartile range (IQR) are shown.
(a-c) Means ± s.d. were from three independent experiments.
Supplementary Figure 2 Screening for DNA methylation-resistant base editors.
(a) Characteristics of the APOBEC/AID deaminases used to construct BEs. (b) Schematic diagram illustrating the in vitro CpG methylation of plasmid vectors. (c) BstUI restriction of unmethylated, partially methylated and fully methylated plasmid vectors. Plasmids were treated with different amounts of CpG methyltransferase M.SssI for in vitro methylation and then digested with methylation-sensitive enzyme BstUI. The methylation of cytidines in plasmid DNA used in (d) and Fig. 1c,d reached almost 100% (arrow head), as the methylated plasmid was almost completely resistant to BstUI restriction. In contrast, decreased amount of CpG methyltransferase resulted in partially methylated plasmid DNAs, which were partially digested accordingly. Gel images are representative of three independent experiments and uncropped gel images are shown in Supplementary Fig. 11. (d) Immunoblots of the BEs co-transfected with unmethylated or methylated vectors. Tubulin was used as a loading control and immunoblot images are representative of three independent experiments. Uncropped blot images are shown in Supplementary Fig. 11.
Supplementary Figure 3 Comparison of base editing efficiencies induced by BE3 and hA3A-BE3 in genomic regions with induced high levels of DNA methylation.
(a) Base editing efficiencies induced by BE3 and hA3A-BE3 in genomic regions with induced high levels of DNA methylation. The C-to-T editing frequencies of indicated cytosines were individually determined and target site sequences are shown with the BE3 editing window (position 4-8, setting the base distal to the PAM as position 1) in lavender, PAM in cyan and CpG site in capital. Means ± s.d. were from three independent experiments. (b) Statistical analysis of normalized C-to-T editing frequencies in regions with induced high levels of DNA methylation shown in (a), setting the ones induced by BE3 as 100%. n = 21 samples from three independent experiments. (c) Statistical analysis of normalized C-to-T editing frequencies at GpC sites in regions with natively or induced high levels of DNA methylation shown in (a) and Figure 1e, setting the ones induced by BE3 as 100%. n = 36 samples from three independent experiments.
(b,c) P value, one-tailed Student’s t test. The median and IQR are shown.
Supplementary Figure 4 Comparison of base editing frequencies induced by BE3 and hA3A-BE3 in genomic regions with low levels of DNA methylation.
(a) Base editing efficiencies induced by BE3 and hA3A-BE3 in genomic regions with natively low levels of DNA methylation. (b) Base editing efficiencies induced by BE3 and hA3A-BE3 in genomic regions with induced low levels of DNA methylation. (c) Statistical analysis of normalized C-to-T editing frequencies at C of GpC sites in regions with natively or induced low levels of DNA methylation shown in (a,b), setting the ones induced by BE3 as 100%. n = 39 samples from three independent experiments. (d) Statistical analysis of normalized C-to-T editing frequencies at non-CpG sites in regions with natively or induced low levels of DNA methylation shown in (a,b), setting the ones induced by BE3 as 100%. n = 24 samples from three independent experiments.
(a,b) Target site sequences are shown with the BE3 editing window (position 4-8, setting the base distal to the PAM as position 1) in lavender, PAM in cyan and CpG site in capital. Shaded gray, the guanines at 5’ end of editable cytosines. Means ± s.d. were from three independent experiments.
(c,d) P value, one-tailed Student’s t test. The median and IQR are shown.
Supplementary Figure 5 Product purity and indel formation yielded by BE3 and hA3A-BE3.
(a) Comparison of product purity yielded by BE3 and hA3A-BE3 in genomic DNA regions with natively or induced high levels of DNA methylation. Fractions of cytosine substitutions induced by BE3 and hA3A-BE3 were individually determined at the indicated cytosines under different conditions. (b) Statistical analysis of the data in (a) showed that C-to-T fractions of base editing products induced by hA3A-BE3 was significantly higher than that by BE3 in genomic regions with high-methylation levels. n = 69 samples from three independent experiments. (c) Comparison of product purity yielded by BE3 and hA3A-BE3 in genomic DNA regions with natively or induced low levels of DNA methylation. Fractions of cytosine substitutions induced by BE3 and hA3A-BE3 were individually determined at indicated cytosines under different conditions. (d) Statistical analysis of the data in (c) showed that C-to-T fractions of base editing products induced by hA3A-BE3 was significantly higher than that by BE3 in genomic regions with low-methylation levels. n = 63 samples from three independent experiments. (e) Comparison of indel formation induced by BE3 and hA3A-BE3 in genomic DNA regions with natively or induced high levels of DNA methylation. (f) Comparison of indel formation induced by BE3 and hA3A-BE3 in genomic DNA regions with natively or induced low levels of DNA methylation. (g) Statistical analysis of the data in (a,c) showed that C-to-T fractions of base editing products induced by BE3 and hA3A-BE3 at CpG sites with high-methylation levels were significantly higher than those at CpG sites with low-methylation levels. n = 36 samples from three independent experiments. (h) Statistical analysis of normalized indel frequencies induced by BE3 and hA3A-BE3 shown in (e) and (f), setting the ones induced by BE3 as 100%. n = 39 (left panel) and 36 (right panel) samples from three independent experiments.
(b,d,g,h) P value, one-tailed Student’s t test. The median and IQR are shown.
(e,f) Means ± s.d. were from three independent experiments. Asterisks denote unusually high basal indel frequencies (or amplification, sequencing or alignment artifact) at examined VEGFA-M-a, VEGFA-M-b and VEGFA-M-c sites in NT.
Supplementary Figure 6 Y130F, D131Y and Y132D mutations of hA3A narrowed the editing window of hA3A-BE3.
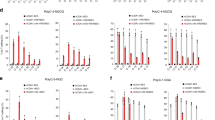
Summary of base editing frequencies induced by hA3A-BE3 (a), hA3A-BE3-Y130F (b), hA3A-BE3-D131Y (c) and hA3A-BE3-Y132D (d) on each cytosine in indicated target sites. Cytosines are counted with the base distal to the PAM setting as position 1. These data show that Y130F, D131Y or Y132D mutation on hA3A module narrowed the major editing window (red box).
Supplementary Figure 7 C101S and C106S mutations of hA3A eliminated the editing activity of hA3A-BE3.
Base editing frequencies induced by hA3A-BE3, hA3A-BE3-C101S (a) and hA3A-BE3-C106S (b) on indicated cytosines at seven different genomic loci. These data show that C101S and C106S mutations of hA3A eliminated the editing activity of corresponding BEs. Means ± s.d. were from three independent experiments.
Supplementary Figure 8 Product purity and indel formation yielded by hA3A-BE3-Y130F and hA3A-BE3-Y132D.
(a) Comparison of product purity yielded by BE3, hA3A-BE3, hA3A-BE3-Y130F and hA3A-BE3-Y132D in genomic DNA regions with natively high levels of DNA methylation. Fractions of cytosine substitutions induced by BE3 and hA3A-BE3s were individually determined at indicated cytosines. (b) Statistical analysis of C-to-T fractions of base editing products induced by BE3, hA3A-BE3-Y130F and hA3A-BE3-Y132D shown in (a). n = 12 samples from three independent experiments. (c) Comparison of indel formation induced by BE3, hA3A-BE3, hA3A-BE3-Y130F and hA3A-BE3-Y132D in genomic DNA regions with natively high levels of DNA methylation. Indel frequencies were individually determined at indicated genomic loci under different conditions. (d) Statistical analysis of normalized indel frequencies induced by BE3, hA3A-BE3-Y130F and hA3A-BE3-Y132D in regions with natively high levels of DNA methylation shown in (c), setting the ones induced by BE3 as 100%. n = 9 samples from three independent experiments. (e) Base editing efficiencies induced by BE3, hA3A-BE3, hA3A-BE3-Y130F and hA3A-BE3-Y132D in genomic regions with natively low levels of DNA methylation. C-to-T editing frequencies of indicated cytosine were individually determined. Target site sequences are shown with the overlapped editing window (position 4-7, setting the base distal to the PAM as position 1) in lavender, PAM in cyan and CpG site in capital. (f) Comparison of product purity yielded by BE3, hA3A-BE3, hA3A-BE3-Y130F and hA3A-BE3-Y132D in genomic DNA regions with natively low levels of DNA methylation. Fractions of cytosine substitutions induced by BE3 and hA3A-BE3s were individually determined at indicated cytosines. (g) Statistical analysis of C-to-T fractions of base editing products induced by BE3, hA3A-BE3-Y130F and hA3A-BE3-Y132D shown in (f). n = 9 samples from three independent experiments. (h) Comparison of indel formation induced by BE3, hA3A-BE3, hA3A-BE3-Y130F and hA3A-BE3-Y132D in genomic DNA regions with natively low levels of DNA methylation. Indel frequencies were individually determined at indicated genomic loci under different conditions. Asterisk denotes an unusually high basal indel frequency (or amplification, sequencing or alignment artifact) at examined VEGFA-M-c site in NT. (i) Statistical analysis of normalized indel frequencies induced by BE3, hA3A-BE3-Y130F and hA3A-BE3-Y132D in regions with natively low levels of DNA methylation shown in (h), setting the ones induced by BE3 as 100%. n = 6 samples from three independent experiments.
(b,d,g,i) P value, one-tailed Student’s t test. The median and IQR are shown.
(c,e,h) Means ± s.d. were from three independent experiments.
Supplementary Figure 9 Additional free UGI further enhances base editing mediated by hA3A-BEs.
(a) Statistical analysis of C-to-T fractions of base editing products induced by hA3A-BE3-Y130F, hA3A-eBE-Y130F, hA3A-BE3-Y132D and hA3A-eBE-Y132D at cytosines of GpC sites in regions with natively low levels of DNA methylation shown in Figure 2h. n = 9 samples from three independent experiments. (b) Statistical analysis of normalized indel frequencies induced by hA3A-BE3-Y130F, hA3A-eBE-Y130F, hA3A-BE3-Y132D and hA3A-eBE-Y132D in regions with natively low levels of DNA methylation shown in Figure 2i, setting the ones induced by hA3A-BE3-Y130F (left) or hA3A-BE3-Y132D (right) as 100%. n = 6 samples from three independent experiments. (c) Comparison of product purity yielded by hA3A-BE3-Y130F, hA3A-eBE-Y130F, hA3A-BE3-Y132D and hA3A-eBE-Y132D in genomic DNA regions with natively high levels of DNA methylation. (d) Statistical analysis of C-to-T fractions of base editing products induced by hA3A-BE3-Y130F, hA3A-eBE-Y130F, hA3A-BE3-Y132D and hA3A-eBE-Y132D shown in (c). n = 6 samples from three independent experiments. (e) Comparison of indels induced by hA3A-BE3-Y130F, hA3A-eBE-Y130F, hA3A-BE3-Y132D and hA3A-eBE-Y132D in genomic DNA regions with natively high levels of DNA methylation. (f) Statistical analysis of normalized indel frequencies induced by hA3A-BE3-Y130F, hA3A-eBE-Y130F, hA3A-BE3-Y132D and hA3A-eBE-Y132D in regions with natively high levels of DNA methylation shown in (e), setting the ones induced by hA3A-BE3-Y130F (left) or hA3A-BE3-Y132D (right) as 100%. n = 6 samples from three independent experiments. (g) Comparison of C-to-T editing frequencies induced by hA3A-BE3-Y130F, hA3A-eBE-Y130F, hA3A-BE3-Y132D and hA3A-eBE-Y132D in genomic DNA regions with natively high levels of DNA methylation. (h) Statistical analysis of normalized C-to-T editing frequencies induced by hA3A-BE3-Y130F, hA3A-eBE-Y130F, hA3A-BE3-Y132D and hA3A-eBE-Y132D in regions with natively high levels of DNA methylation shown in (g), setting the ones induced by hA3A-BE3-Y130F (left) or hA3A-BE3-Y132D (right) as 100%. n = 6 samples from three independent experiments.
(a,b,d,f,h) P value, one-tailed Student’s t test. The median and IQR are shown.
(e,g) Means ± s.d. were from three independent experiments.
Supplementary Figure 10 Summary of BE3 and hA3A-BEs.
(a) Schematic diagrams illustrate configurations of BE3 and hA3A-BEs. (b) List of relevant features of BE3 and hA3A-BEs.
Supplementary Figure 11 Uncropped images of blots and gels.
(a) Uncropped blot images for Fig. 1d. (b) Uncropped blot images for Fig. 2e. (c) Uncropped gel images for Supplementary Fig. 2c. (d) Uncropped blot images for Supplementary Fig. 2d.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 (PDF 2729 kb)
Supplementary Items
Supplementary Tables 1 and 2 and Supplementary Note 1 (PDF 4239 kb)
Supplementary Table 3
Calculation of indels (XLSX 35 kb)
Supplementary Table 4
Calculation of base substitutions (XLSX 953 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Li, J., Wang, Y. et al. Efficient base editing in methylated regions with a human APOBEC3A-Cas9 fusion. Nat Biotechnol 36, 946–949 (2018). https://doi.org/10.1038/nbt.4198
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4198
- Springer Nature America, Inc.
This article is cited by
-
Base editing of organellar DNA with programmable deaminases
Nature Reviews Molecular Cell Biology (2024)
-
Whole-brain in vivo base editing reverses behavioral changes in Mef2c-mutant mice
Nature Neuroscience (2024)
-
Mesoscale DNA features impact APOBEC3A and APOBEC3B deaminase activity and shape tumor mutational landscapes
Nature Communications (2024)
-
Deep learning models incorporating endogenous factors beyond DNA sequences improve the prediction accuracy of base editing outcomes
Cell Discovery (2024)
-
Engineering APOBEC3A deaminase for highly accurate and efficient base editing
Nature Chemical Biology (2024)