Abstract
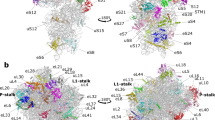
Ribosomes, the protein factories of living cells, translate genetic information carried by messenger RNAs into proteins, and are thus involved in virtually all aspects of cellular development and maintenance. The few available structures of the eukaryotic ribosome1,2,3,4,5,6 reveal that it is more complex than its prokaryotic counterpart7,8, owing mainly to the presence of eukaryote-specific ribosomal proteins and additional ribosomal RNA insertions, called expansion segments9. The structures also differ among species, partly in the size and arrangement of these expansion segments. Such differences are extreme in kinetoplastids, unicellular eukaryotic parasites often infectious to humans. Here we present a high-resolution cryo-electron microscopy structure of the ribosome of Trypanosoma brucei, the parasite that is transmitted by the tsetse fly and that causes African sleeping sickness. The atomic model reveals the unique features of this ribosome, characterized mainly by the presence of unusually large expansion segments and ribosomal-protein extensions leading to the formation of four additional inter-subunit bridges. We also find additional rRNA insertions, including one large rRNA domain that is not found in other eukaryotes. Furthermore, the structure reveals the five cleavage sites of the kinetoplastid large ribosomal subunit (LSU) rRNA chain, which is known to be cleaved uniquely into six pieces10,11,12, and suggests that the cleavage is important for the maintenance of the T. brucei ribosome in the observed structure. We discuss several possible implications of the large rRNA expansion segments for the translation-regulation process. The structure could serve as a basis for future experiments aimed at understanding the functional importance of these kinetoplastid-specific ribosomal features in protein-translation regulation, an essential step towards finding effective and safe kinetoplastid-specific drugs.




Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
The electron microscopy map has been deposited in the European Molecular Biology Laboratory (EMBL) European Bioinformatics Institute Electron Microscopy Data Bank (EMDB) under accession code EMD-2239. Coordinates of electron-microscopy-based model have been deposited in the RCSB Protein Data Bank under accession numbers 3ZEQ, 3ZEX, 3ZEY and 3ZF7.
References
Ben-Shem, A. et al. Crystal structure of the eukaryotic ribosome. Science 330, 1203–1209 (2010)
Ben-Shem, A. et al. The structure of the eukaryotic ribosome at 3.0 Å resolution. Science 334, 1524–1529 (2011)
Armache, J. P. et al. Localization of eukaryote-specific ribosomal proteins in a 5.5 Å cryo-EM map of the 80S eukaryotic ribosome. Proc. Natl Acad. Sci. USA 107, 19754–19759 (2010)
Armache, J. P. et al. Cryo-EM structure and rRNA model of a translating eukaryotic 80S ribosome at 5.5 Å resolution. Proc. Natl Acad. Sci. USA 107, 19748–19753 (2010)
Rabl, J., Leibundgut, M., Ataide, S. F., Haag, A. & Ban, N. Crystal structure of the eukaryotic 40S ribosomal subunit in complex with initiation factor 1. Science 331, 730–736 (2011)
Klinge, S., Voigts-Hoffmann, F., Leibundgut, M., Arpagaus, S. & Ban, N. Crystal structure of the eukaryotic 60S ribosomal subunit in complex with initiation factor 6. Science 334, 941–948 (2011)
Klinge, S., Voigts-Hoffmann, F., Leibundgut, M. & Ban, N. Atomic structures of the eukaryotic ribosome. Trends Biochem. Sci. 37, 189–198 (2012)
Wilson, D. N. & Doudna Cate, J. H. The structure and function of the eukaryotic ribosome. Cold Spring Harb. Perspect. Biol. 4, a011536 (2012)
Yokoyama, T. & Suzuki, T. Ribosomal RNAs are tolerant toward genetic insertions: evolutionary origin of the expansion segments. Nucleic Acids Res. 36, 3539–3551 (2008)
White, T. C., Rudenko, G. & Borst, P. Three small RNAs within the 10 kb trypanosome rRNA transcription unit are analogous to domain VII of other eukaryotic 28S rRNA. Nucleic Acids Res. 14, 9471–9489 (1986)
Cordingley, J. S. & Turner, M. J. 6.5 S RNA; preliminary characterization of unusual small RNAs in Trypanosoma brucei . Mol. Biochem. Parasitol. 1, 91–96 (1980)
Campbell, D. A., Kubo, K., Graham Clark, C. & Boothroyd, J. C. Precise identification of cleavage sites involved in the unusual processing of trypanosome ribosomal RNA. J. Mol. Biol. 196, 113–124 (1987)
Berriman, M. et al. The genome of the African trypanosome Trypanosoma brucei . Science 309, 416–422 (2005)
Gao, H., Juri Ayub, M., Levin, M. L. & Frank, J. The structure of the 80S ribosome from Trypanosoma cruzi reveals unique rRNA components. Proc. Natl Acad. Sci. USA 102, 10206–10211 (2005)
Clayton, C. & Shapira, M. Post-transcriptional regulation of gene expression in trypanosomes and leishmanias. Mol. Biochem. Parasitol. 156, 93–101 (2007)
Michaeli, S. Trans-splicing in trypanosomes: machinery and its impact on the parasite transcriptome. Future Microbiol. 6, 459–474 (2011)
Ciganda, M. & Williams, N. Characterization of a novel association between two trypanosome-specific proteins and 5S rRNA. PLoS ONE 7, e30029 (2012)
Ivens, A. C. et al. The genome of the kinetoplastid parasite, Leishmania major . Science 309, 436–442 (2005)
Zhang, Q., Bettadapura, R. & Bajaj, C. Macromolecular structure modeling from 3D EM using VolRover 2.0. Biopolymers 97, 709–731 (2012)
Pintilie, G., Zhang, J., Goddard, T., Chiu, W. & Gossard, D. Quantitative analysis of cryo-EM density map segmentation by watershed and scale-space filtering, and fitting of structures by alignment to regions. J. Struct. Biol. 170, 429–438 (2010)
Jossinet, F. & Westhof, E. Sequence to Structure (S2S): display, manipulate and interconnect RNA data from sequence to structure. Bioinformatics 21, 3320–3321 (2005)
Jossinet, F., Ludwig, T. E. & Westhof, E. Assemble: an interactive graphical tool to analyze and build RNA architectures at the 2D and 3D levels. Bioinformatics 26, 2057–2059 (2010)
Trabuco, L. G., Villa, E., Mitra, K., Frank, J. & Schulten, K. Flexible fitting of atomic structures into electron microscopy maps using molecular dynamics. Structure 16, 673–683 (2008)
Liao, H. Y. & Frank, J. Classification by bootstrapping in single particle methods. IEEE Int. Symp. Biom. Imaging 169–172 (2010)
Cannone, J. J. et al. The comparative RNA web (CRW) site: an online database of comparative sequence and structure information for ribosomal, intron, and other RNAs. Bioinformatics 3, 2 (2002)
Ayub, M. J., Atwood, J., Nuccio, A., Tarleton, R. & Levin, M. J. Proteomic analysis of the Trypanosoma cruzi ribosomal proteins. Biochem. Biophys. Res. Commun. 382, 30–34 (2009)
Meyuhas, O. Physiological roles of ribosomal protein S6: One of its kind. Int. Rev. Cell Mol. Biol. 268, 1–37 (2008)
Srivastava, S., Verschoor, A. & Frank, J. Eukaryotic initiation factor 3 does not prevent association through physical blockage of the ribosomal subunit–subunit interface. J. Mol. Biol. 226, 301–304 (1992)
Siridechadilok, B., Fraser, C. S., Hall, R. J., Doudna, J. A. & Nogales, E. structural roles for human translation factor eIF3 in initiation of protein synthesis. Science 310, 1513–1515 (2005)
Lescrinier, E. M. H. P. et al. Structure of the pyrimidine-rich internal loop in the poliovirus 30′-UTR: The importance of maintaining pseudo-2-fold symmetry in RNA helices containing two adjacent non-canonical base-pairs. J. Mol. Biol. 331, 759–769 (2003)
Kaminsky, R., Beaudoin, E. & Cunningham, I. Cultivation of the life cycle stages of Trypanosoma brucei sspp. Acta Trop. 45, 33–43 (1988)
Gómez, E. B., Medina, G., Ballesta, J. P., Levin, M. J. & Téllez-Iñón, M. T. Acidic ribosomal P proteins are phosphorylated in Trypanosoma cruzi . Int. J. Parasitol. 31, 1032–1039 (2001)
Grassucci, R. A., Taylor, D. J. & Frank, J. Preparation of macromolecular complexes for cryo-electron microscopy. Nature Protocols 2, 3239–3246 (2007)
Dubochet, J. et al. Cryo-electron microscopy of vitrified specimens. Q. Rev. Biophys. 21, 129–228 (1988)
Wagenknecht, T., Frank, J., Boublik, M., Nurse, K. & Ofengand, J. Direct localization of the tRNA–anticodon interaction site on the Escherichia coli 30 S ribosomal subunit by electron microscopy and computerized image averaging. J. Mol. Biol. 203, 753–760 (1988)
Lei, J. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005)
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996)
Shaikh, T. R. et al. SPIDER image processing for single-particle reconstruction of biological macromolecules from electron micrographs. Nature Protocols 3, 1941–1974 (2008)
Scheres, S. H. W. et al. Disentangling conformational states of macromolecules in 3D-EM through likelihood optimization. Nature Methods 4, 27–29 (2007)
Scheres, S. H. W., Nuñez-Ramirez, R., Sorzano, C. O. S., Carazo, J. M. & Marabini, R. Image processing for electron microscopy single-particle analysis using Xmipp. Nature Protocols 3, 977–990 (2008)
Heymann, J. B. Bsoft: image and molecular processing in electron microscopy. J. Struct. Biol. 133, 156–169 (2001)
Heymann, J. B., Cardone, G., Winkler, D. C. & Steven, A. C. Computational resources for cryo-electron tomography in Bsoft. J. Struct. Biol. 161, 232–242 (2008)
Pintilie, G., Zhang, J., Goddard, T., Chiu, W. & Gossard, D. Quantitative analysis of cryo-EM density map segmentation by watershed and scale-space filtering, and fitting of structures by alignment to regions. J. Struct. Biol. 170, 427–438 (2010)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 13, 1605–1612 (2004)
Baker, M. L., Yu, Z., Chiu, W. & Bajaj, C. Automated segmentation of molecular subunits in electron cryomicroscopy density maps. J. Struct. Biol. 156, 432–441 (2006)
Yu, Z. & Bajaj, C. Automatic ultrastructure segmentation of reconstructed cryoEM maps of icosahedral viruses. IEEE Trans. Image Process. 14, 1324–1337 (2005)
Zhang, Q., Bettadapura, R. & Bajaj, C. Macromolecular structure modeling from 3D EM using VolRover 2.0. Biopolymers 97, 709–731 (2012)
Zeyen, Y. & Bajaj, C. Computational approaches for automatic structural analysis of large biomolecular complexes. IEEE/ACM Trans. Comput. Biol. Bioinform. 5, 568–582 (2008)
Penczek, P. A., Yang, C., Frank, J. & Spahn, C. M. Estimation of variance in single-particle reconstruction using the bootstrap technique. J. Struct. Biol. 154, 168–183 (2006)
Zhang, W., Kimmel, M., Spahn, C. M. & Penczek, P. A. Heterogeneity of large macromolecular complexes revealed by 3D cryo-EM variance analysis. Structure 16, 1770–1776 (2008)
Liao, H. Y. & Frank, J. Classification by bootstrapping in single particle methods. IEEE Int. Symp. Biom. Imaging 169–172 (2010)
Simonetti, A. et al. Structure of the 30S translation initiation complex. Nature 455, 416–420 (2008)
Pruesse, E. et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35, 7188–7196 (2007)
Cannone, J. J. et al. The Comparative RNA Web (CRW) Site: an online database of comparative sequence and structure information for ribosomal, intron, and other RNAs. Bioinformatics 3, 2 (2002)
Will, S., Joshi, T., Hofacker, I. L., Stadler, P. F. & Backofen, R. LocARNA-P: accurate boundary prediction and improved detection of structural RNAs. RNA 18, 900–914 (2012)
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201 (2006)
Kiefer, F., Arnold, K., Künzli, M., Bordoli, L. & Schwede, T. The SWISS-MODEL Repository and associated resources. Nucleic Acids Res. 37, D387–D392 (2009)
Kelley, L. A. & Sternberg, M. J. E. Protein structure prediction on the Web: a case study using the Phyre server. Nature Protocols 4, 363–371 (2009)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007)
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. Model. 1, 33–8–27-8 (1996)
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781 (2005)
Acknowledgements
This work is dedicated to the memory of Mariano Levin, who collaborated with J.F. and S.M.A. on the ribosomes from T. cruzi and T. brucei. We thank G. Cardone for assistance in the local resolution computation, and M. Thomas for her assistance with the preparation of figures. We wish to thank N. Williams for an useful discussion about the T. brucei LSU rRNA processing. This work was supported by the Howard Hughes Medical Institute (HHMI) and the National Institutes of Health (NIH) R01 GM29169 (to J.F.), L’Agence Nationale de la recherche (ANR) project AMIS ARN ANR-09-BLAN-0160 (E.W. and F.J.), as well as NIH R01-EB004873 and R01-GM074258 (to Q.Z. and C.B.). S.N.B. was supported by a Centers for Disease Control (CDC) Emerging Infectious Diseases (EID) fellowship program.
Author information
Authors and Affiliations
Contributions
Y.H., A.d.G., S.N.B., F.J., Q.Z., C.B., S.M.-A., E.W. and J.F. interpreted the data and wrote the manuscript. S.N.B. purified the T. brucei ribosomes. Y.H., J. Fu and R.A.G. carried out the cryo-EM experiments. H.Y.L. performed the three-dimensional variance estimation. Y.H., A.J. and Q.Z. performed the density-map segmentations. Y.H., A.d.G., J. Fu, A.J. and H.Y.L. carried out the cryo-EM data processing. Y.H. and F.J. modelled the rRNA. Y.H. and Q.Z. modelled the ribosomal proteins. J.F. directed research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Text, Supplementary References, Supplementary Figures 1-10 and Supplementary Tables 1-2. (PDF 5513 kb)
Rights and permissions
About this article
Cite this article
Hashem, Y., des Georges, A., Fu, J. et al. High-resolution cryo-electron microscopy structure of the Trypanosoma brucei ribosome. Nature 494, 385–389 (2013). https://doi.org/10.1038/nature11872
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11872
- Springer Nature Limited
This article is cited by
-
Role of the RNA-binding protein ZC3H41 in the regulation of ribosomal protein messenger RNAs in trypanosomes
Parasites & Vectors (2023)
-
A method for restoring signals and revealing individual macromolecule states in cryo-ET, REST
Nature Communications (2023)
-
A single pseudouridine on rRNA regulates ribosome structure and function in the mammalian parasite Trypanosoma brucei
Nature Communications (2023)
-
Typical structure of rRNA coding genes in diplonemids points to two independent origins of the bizarre rDNA structures of euglenozoans
BMC Ecology and Evolution (2022)
-
Structural basis for the inhibition of translation through eIF2α phosphorylation
Nature Communications (2019)