Abstract
Arising from: Q. Shi & R. W. King Nature 437, 1038–1042 (2005); Shi & King reply
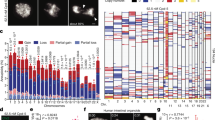
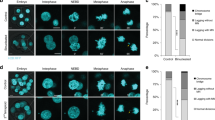
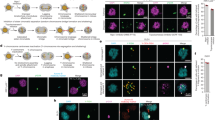
One simple, widely accepted mechanism for generating an aberrant chromosome number, or aneuploidy, is through nondisjunction — a chromosome distribution error that occurs during mitosis when both copies of a duplicated chromosome are deposited into one daughter cell and none into the other. Shi and King1 challenge this view, concluding that nondisjunction does not yield aneuploid cells directly, but instead gives rise to tetraploid cells that may subsequently become aneuploid through further division. Here we show that the direct result of chromosome nondisjunction is gain or loss of a single chromosome, which results in near-diploid aneuploidy, not tetraploidy. We suggest that chromatin trapped in the cytokinetic cleavage furrow is the more likely reason for furrow regression and tetraploidization.

Similar content being viewed by others
References
Shi, Q. & King, R. W. Nature 437, 1038–1042 (2005).
Weaver, B. A. A. et al. J. Cell Biol. 162, 551–563 (2003).
Babu, J. R. et al. . J. Cell Biol. 160, 341–353 (2003).
Matsuura, S. et al. Am. J. Med. Genet. A 140, 358–367 (2006).
Hanks, S. et al. . Nature Genet. 36, 1159–1161 (2004).
Limwongse, C., Schwartz, S., Bocian, M. & Robin, N. H. Am. J. Med. Genet. 82, 20–24 (1999).
Mullins, J. M. & Biesele, J. J. J. Cell Biol. 73, 672–684 (1977).
Andreassen, P. R., Lohez, O. D., Lacroix, F. B. & Margolis, R. L. Mol. Biol. Cell 12, 1315–1328 (2001).
Uetake, Y. & Sluder, G. J. Cell Biol. 165, 609–615 (2004).
Wong, C. & Stearns, T. BMC Cell Biol. 6, doi:10.1186/1471-2121-6-6 (2005).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Weaver, B., Silk, A. & Cleveland, D. Nondisjunction, aneuploidy and tetraploidy. Nature 442, E9–E10 (2006). https://doi.org/10.1038/nature05139
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05139
- Springer Nature Limited
This article is cited by
-
Kinesin-7 CENP-E regulates chromosome alignment and genome stability of spermatogenic cells
Cell Death Discovery (2020)
-
53BP1 can limit sister-chromatid rupture and rearrangements driven by a distinct ultrafine DNA bridging-breakage process
Nature Communications (2018)
-
Multiparameter DNA content analysis identifies distinct groups in primary breast cancer
British Journal of Cancer (2013)
-
Skp-cullin-F box E3 ligase component FBXL2 ubiquitinates Aurora B to inhibit tumorigenesis
Cell Death & Disease (2013)
-
Mosaicism for combined tetrasomy of chromosomes 8 and 18 in a dysmorphic child: A result of failed tetraploidy correction?
BMC Medical Genetics (2009)