Abstract
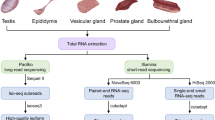
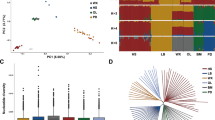
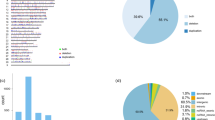
Sex reversal has been studied extensively in vertebrate species, particularly in domestic goats, because polled intersex syndrome (PIS) has seriously affected their production efficiency. In the present study, we used histopathologically diagnosed cases of PIS to identify correlated genomic regions and variants using representative selection signatures and performed GWAS using Restriction-Site Associated Resequencing DNA. We identified 171 single-nucleotide polymorphisms (SNPs) that may have contributed to this phenotype, and 53 SNPs were determined to be located in coding regions using a general linear model. The transcriptome data sets of differentially expressed genes (DEGs) in the pituitary tissues of intersexual and nonintersexual goats were examined using high-throughput technology. A total of 10,063 DEGs and 337 long noncoding RNAs were identified. The DEGs were clustered into 56 GO categories and determined to be significantly enriched in 53 signaling pathways by KEGG analysis. In addition, according to qPCR results, PSPO2 and FSH were significantly more highly expressed in sexually mature pituitary tissues of intersexual goats compared to healthy controls (nonintersexual). These results demonstrate that certain novel potential genomic regions may be responsible for intersexual goats, and the transcriptome data indicate that the regulation of various physiological systems is involved in intersexual goat development. Therefore, these results provide helpful data for understanding the molecular mechanisms of intersex syndrome in goats.






Similar content being viewed by others
References
André GM, Martins Trevisan C, Pedruzzi IN, Fernandes RFM, Oliveira R, Christofolini DM, Bianco B, Barbosa CP (2018) The impact of FSHR gene polymorphisms Ala307Thr and Asn680Ser in the endometriosis development. DNA Cell Biol 37(6):584–591. https://doi.org/10.1089/dna.2017.4093
Antypa N, Souery D, Tomasini M, Albani D, Fusco F, Mendlewicz J, Serretti A (2016) Clinical and genetic factors associated with suicide in mood disorder patients. Eur Arch Psychiatry Clin Neurosci 266(2):181–193
Bongso TA, Thavalingam M, Mukherjee TK: Thavalingam, Mukherjee TK (1982) Intersexuality associated with XX/XY mosaicism in a horned goat. Cytogenet Cell Genet 34(4):315–319
Bouilly J, Beau I, Barraud S, Bernard V, Delemer B, Young J, Binart N (2017) R-spondin2, a novel target of NOBOX: identification of variants in a cohort of women with primary ovarian insufficiency. J Ovarian Res 10(1):51. https://doi.org/10.1186/s13048-017-0345-0
Boulanger L, Pannetier M, Gall L, Allais-Bonnet A, Elzaiat M, Le Bourhis D, Daniel N, Richard C, Cotinot C, Ghyselinck NB, Pailhoux E (2014) FOXL2 is a female sex-determining gene in the goat. Curr Biol 24(4):404–408
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Chen ZJ, Pudas R, Sharma S, Smart OS, Juffer AH, Hiltunen JK, Wierenga RK, Haapalainen AM (2008) Structural enzymological studies of 2-enoyl thioester reductase of the human mitochondrial FAS II pathway: new insights into its substrate recognition properties. J Mol Biol 379(4):830–844. https://doi.org/10.1016/j.jmb.2008.04.041
Chen D, Liu Y, Yang H, Chen D, Zhang X, Fermandes JC, Chen Y (2016) Connexin 43 promotes ossification of the posterior longitudinal ligament through activation of the ERK1/2 and p38 MAPK pathways. Cell Tissue Res 363(3):765–773. https://doi.org/10.1007/s00441-015-2277-6
Crisponi L, Deiana M, Loi A, Chiappe F, Uda M, Amati P, Bisceglia L, Zelante L, Nagaraja R, Porcu S, Ristaldi MS, Marzella R, Rocchi M, Nicolino M, Lienhardt-Roussie A, Nivelon A, Verloes A, Schlessinger D, Gasparini P, Bonneau D, Cao A, Pilia G (2001) The putative forkhead transcription factor FOXL2 is mutated in blepharophimosis/ptosis/epicanthus inversus syndrome. Nat Genet 27(2):159–166
Dong X, Liao W, Zhang L, Tu X, Hu J, Chen T, Dai X, Xiong Y, Liang W, Ding C, Liu R, Dai J, Wang O, Lu L, Lu X (2017) RSPO2 suppresses colorectal cancer metastasis by counteracting the Wnt5a/Fzd7-driven noncanonical Wnt pathway. Cancer Lett 402:153–165
Fisher AD, Ristori J, Fanni E, Castellini G, Forti G, Maggi M (2016) Gender identity, gender assignment and reassignment in individuals with disorders of sex development: a major of dilemma. J Endocrinol Investig 39(11):1207–1224
Gonen N, Futtner CR, Wood S, Garcia-Moreno SA, Salamone IM, Samson SC, Sekido R, Poulat F, Maatouk DM, Lovell-Badge R (2018) Sex reversal following deletion of a single distal enhancer of Sox9. Science 360(6396):1469–1473. https://doi.org/10.1126/science.aas9408
Harris A, Siggers P, Corrochano S, Warr N, Sagar D, Grimes DT, Suzuki M, Burdine RD, Cong F, Koo BK, Clevers H, Stévant I, Nef S, Wells S, Brauner R, Ben Rhouma B, Belguith N, Eozenou C, Bignon-Topalovic J, Bashamboo A, McElreavey K, Greenfield A (2018) ZNRF3 functions in mammalian sex determination by inhibiting canonical WNT signaling. Proc Natl Acad Sci USA 115(21):5474–5479. https://doi.org/10.1073/pnas.1801223115
Hirohata S, Inagaki J, Ohtsuki T (2017) Diverse functions of a disintegrin and metalloproteinase with thrombospondin motif-1. Yakugaku Zasshi 137(7):811–814. https://doi.org/10.1248/yakushi.16-00236-4
Huang P, Zhou Z, Shi F, Shao G, Wang R, Wang J, Wang K, Ding W (2016) Effects of the IGF-1/PTEN/Akt/FoxO signaling pathway on male reproduction in rats subjected to water immersion and restraint stress. Mol Med Rep 14(6):5116–5124
Hullinger R, Li M, Wang J, Peng Y, Dowell JA, Bomba-Warczak E, Mitchell HA, Burger C, Chapman ER, Denu JM, Li L, Puglielli L (2016) Increased expression of AT-1/SLC33A1 causes an autistic-like phenotype in mice by affecting dendritic branching and spine formation. J Exp Med 213(7):1267–1284
Ilmer M, Boiles AR, Regel I, Yokoi K, Michalski CW, Wistuba II, Rodriguez J, Alt E, Vykoukal J (2015) RSPO2 enhances canonical Wnt signaling to confer stemness-associated traits to susceptible pancreatic cancer cells. Cancer Res 75(9):1883–1896
Joshi NA, Fass JN (2011) Sickle: a sliding-window, adaptive, quality-based trimming tool for FastQ files (Version 1.33). https://github.com/najoshi/sickle
Juniarto AZ, van der Zwan YG, Santosa A, Ariani MD, Eggers S, Hersmus R, Themmen AP, Bruggenwirth HT, Wolffenbuttel KP, Sinclair A, White S, Looijenga LH, de Jong FH, Faradz SM, Drop SL (2016) Hormonal evaluation in relation to phenotype and genotype in 286 patients with a disorder of sex development from Indonesia. Clin Endocrinol (Oxf) 85(2):247–257. https://doi.org/10.1111/cen.13051
Lapierre M, Docquier A, Castet-Nicolas A, Gitenay D, Jalaguier S, Teyssier C, Cavaillès V (2015) The emerging role of the transcriptional coregulator RIP140 in solid tumors. Biochim Biophys Acta 1856(1):144–150. https://doi.org/10.1016/j.bbcan.2015.06.006
Li H, Durbin R (2009) Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics 25:1754–1760
Maoz R, Garfinkel BP, Soreq H (2017) Alzheimer’s disease and ncRNAs. Adv Exp Med Biol 978:337–361. https://doi.org/10.1007/978-3-319-53889-1_18 (Review)
Pailhoux E, Cribiu EP, Chaffaux S, Darre R, Fellous M, Cotinot C (1994) Molecular analysis of 60, XX pseudohermaphrodite polled goats for the presence of SRY and ZFY genes. J Reprod Fertil 100:491–496
Pailhoux E, Vigier B, Chaffaux S, Servel N, Taourit S, Furet JP, Fellous M, Grosclaude F, Cribiu EP, Cotinot C, Vaiman D (2001a) A 11.7-kb deletion triggers intersexuality and polledness in goats. Nat Genet 29(4):453–458
Pailhoux E, Vigier B, Vaiman D, Schibler L, Vaiman A, Cribiu E, Nezer C, Georges M, Sundrstrom J, Pelliniemi LJ, Fellous M, Cotinol C (2001b) Contribution of domestic animals to the identification of new genes involved in sex determination. J Exp Zool 290:700–708
Pailhoux E, Vigier B, Vaiman D, Servel N, Chaffaux S, Cribiu EP, Cotinot C (2002) Ontogenesis of female-to-male sex-reversal in XX polled goats. Dev Dyn 224(1):39–50
Pang XL, Long W, Li J, Yang J (2016) 45, XO/47, XXX/46, XX male sex reversal syndrome. A case report. J Reprod Med 61(9–10):510–512
Pannetier M, Renault L, Jolivet G, Cotinot C, Pailhoux E (2005) Ovarian-specific expression of a new gene regulated by the goat PIS region and transcribed by a FOXL2 bidirectional promoter. Genomics 85(6):715–726
Parl A, Mitchell SL, Clay HB, Reiss S, Li Z, Murdock DG (2013) The mitochondrial fatty acid synthesis (mtFASII) pathway is capable of mediating nuclear-mitochondrial cross talk through the PPAR system of transcriptional activation. Biochem Biophys Res Commun 441(2):418–424. https://doi.org/10.1016/j.bbrc.2013.10.072
Poth T, Breuer W, Walter B, Hecht W, Hermanns W (2010) Disorders of sex development in the dog—adoption of a new nomenclature and reclassification of reported cases. Anim Reprod Sci 121(3–4):197–207
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38(8):904–909
Schibler L, Cribiu EP, Oustry-Vaiman A, Furet JP, Vaiman D (2000) Fine mapping suggests that the goat polled intersex syndrome and the human blepharophimosis ptosis epicanthus syndrome map to a 100-kb homologous region. Genome Res 10:311–318
Sheridan R, Belludi C, Khoury J, Stanek J, Handwerger S (2015) FOXO1 expression in villous trophoblast of preeclampsia and fetal growth restriction placentas. Histol Histopathol 30(2):213–222
Silva SV, Lima MA, Cella N, Jaeger RG, Freitas VM (2016) ADAMTS-1 is found in the nuclei of normal and tumoral breast cells. PLoS One 11(10):e0165061
Szatkowska I, Zaborski D, Proskura WS, Tabor S (2015) Polledness intersex syndrome in goats—molecular and histological aspects. Turk J Vet Anim Sci 38(6):612–617
Tamburino L, La Vignera S, Tomaselli V, Condorelli RA, Cannarella R, Mongioì LM, Calogero AE (2017) The– 29G/A FSH receptor gene polymorphism is associated with higher FSH and LH levels in normozoospermic men. J Assist Reprod Genet 34(10):1289–1294
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739. https://doi.org/10.1093/molbev/msr121 (PMID: 21546353)
Vallenzasca C, Galli A (1990) Cytogenetical and histopathologic study of two cases of polled goats. Andrologia 22(3):289–290
Wei S, Shen X, Lai L, Liang H, Deng Y, Gong Z, Che T (2018) FSH receptor binding inhibitor impacts K-Ras and c-Myc of ovarian cancer and signal pathway. Oncotarget 9(32):22498–22508. https://doi.org/10.18632/oncotarget.25139
Wichura MJ (2006) The coordinate-free approach to linear models. Cambridge series in statistical and probabilistic mathematics. Cambridge University Press, Cambridge, pp 14–199 (ISBN 978-0-521-86842-6. MR 2283455)
Yang YX, Wei L, Zhang YJ, Hayano T, Piñeiro Pereda MDP, Nakaoka H, Li Q, Barragán Mallofret I, Lu YZ, Tamagnone L, Inoue I, Li X, Luo JY, Zheng K, You H (2018) Long non-coding RNA p10247, high expressed in breast cancer (lncRNA-BCHE), is correlated with metastasis. Clin Exp Metastasis. https://doi.org/10.1007/s10585-018-9901-2
Zhang Y (2014) Study on PIS region, PISRT1 and FoxL2 genes in polled intersex dairy goat (D). Agricultural University of Hebei, Shijiazhuang (in Chinese)
Acknowledgements
This work was supported by Fundamental Research Funds for the Central Universities (XDJK2018B014), National Natural Science Foundation of China (no. 31172195), the Characteristic Germplasm Resources Population Selection and Innovation on Mutton Sheeps and Goats (no. 2015BAD03B05), and Chongqing Research Program of Basic Research and Frontier Technology (cstc2018jcyjAX0153).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
E, GX., Jin, ML., Zhao, YJ. et al. Genome-wide analysis of Chongqing native intersexual goats using next-generation sequencing. 3 Biotech 9, 99 (2019). https://doi.org/10.1007/s13205-019-1612-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-019-1612-0