Abstract
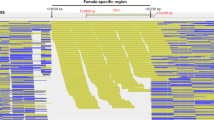
A male-specific marker of a DNA sequence for the Pacific bluefin tuna Thunnus orientalis, Male delta 6 (Md6), was identified by comparative study of the sequences obtained by F3-cultured male and female AFLP-selective DNA amplification products, followed by high-throughput DNA sequencing. Md6 was characteristic in continuous 6-bp nucleotide deletions compared to redundant sequences that could be seen in both sexes. The PCR genotyping primers were developed based on the Md6 sequence, which showed that 29 of 32 F3 males (90.6 %) and 9 of 13 (69.2 %) F2 parental males were genotyping positive, whereas in females, the genotyping test was negative in parental F2 (n = 12), and 1 of 32 (3.1 %) F3 female was Md6-positive. Parentage allocation tests suggested that Md6-positive F2 males were more attendant in spawning than Md6-negative males, suggesting that Md6 was passed from male parents to male progeny under aquaculture conditions.




Similar content being viewed by others
References
Fromentin JM, Powers JE (2005) Atlantic bluefin tuna: population dynamics, ecology, fisheries and management. Fish Fish 6:281–306
Anonymous (2011) Conservation and management measure for Pacific bluefin tuna. WCPFC 7th regular session available online (http://www.wcpfc.int/node/3407)
Miyashita S, Murata O, Sawada Y, Okada T, Kubo Y, Ishitani Y, Seoka M, Kumai H (2000) Maturation and spawning of cultured bluefin tuna, Thunnus thynnus. Suisanzoshoku 48:475–488 (in Japanese, with English abstract)
Sawada Y, Okada T, Miyashita S, Murata O, Kumai H (2005) Completion of the Pacific bluefin tuna Thunnus orientalis (Temminck et Schlegel) life cycle. Aquacult Res 36:413–421
Normile D (2009) Persevering researchers make a splash with farm-bred tuna. Science 324:1260–1261
Masuma S, Takebe T, Sakakura Y (2010) A review of the broodstock management and larviculture of the Pacific northern bluefin tuna in Japan. Aquaculture 315:2–8
Ida H, Oka N, Terashima H, Hayashizaki K (1993) Karyotypes and cellular DNA contents of three species of the family Scombridae from Japan. Nippon Suisan Gakkaishi 59:1319–1323
Susca V, Corriero A, Bridges CR, De Metrio G (2001) Study of the sexual maturity of female bluefin tuna: purification and partial characterization of vitellogenin and its use in an enzyme-linked immunosorbent assay. J Fish Biol 58:815–831
Pousis C, De Giorgi C, Mylonas CC, Bridges CR, Zupa R, Vassallo-Agius R, de la Gándara F, Dileo C, De Metrio G, Corriero A (2011) Comparative study of liver vitellogenin gene expression and oocyte yolk accumulation in wild and captive Atlantic bluefin tuna (Thunnus thynnus L.). Anim Reprod Sci 123:98–105
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Agawa Y, Komiya T, Honryo T, Kurata M, Okada T, Murata O, Kumai H, Sawada Y (2011) Screening of the male characteristic DNA fragment using AFLP analysis in Pacific bluefintuna, Thunnus orientalis. Nippon Suisan Gakkaishi 77:639–646 (in Japanese, with English abstract)
Kumai H, Miyashita S, Sakamoto W, Ono S (eds) (2012) Full-life cycle aquaculture of the Pacific bluefin tuna. Agriculture and Forestry Statistics Publishing Inc, Tokyo, pp 6–24
Seoka M, Kurata M, Tamagawa R, Biswas AK, Biswas BK, Yong ASK, Kim YS, Ji SC, Takii K, Kumai H (2008) Dietary supplementation of salmon roe phospholipid enhances the growth and survival of Pacific bluefin tuna Thunnus orientalis larvae and juveniles. Aquaculture 275:225–234
Adachi K, Kato K, Yamamoto M, Ishimaru K, Kobayashi T, Murata O, Kumai H (2008) Pulsed expression of growth hormone mRNA in the pituitary of juvenile Pacific bluefin tuna under aquacultured conditions. Aquaculture 281:158–161
Takagi M, Okamura T, Chow S, Taniguchi N (1999) PCR primers for microsatellite loci in tuna species of the genus Thunnus and its application for population genetic study. Fish Sci 65:571–576
Appleyard SA, Grewe PM, Innes BH, Ward RD (2001) Population structure of yellowfin tuna (Thunnus albacares) in the western Pacific Ocean, inferred from microsatellite loci. Mar Biol 139:383–393
Nomura S, Kobayashi T, Agawa Y, Margulies D, Scholey V, Sawada Y, Yagishita N (2014) Genetic population structure of the Pacific bluefin tuna Thunnus orientalis and the yellowfin tuna Thunnus albacares in the North Pacific Ocean. Fish Sci. doi:10.1007/s12562-014-0789-8
Sekino M, Kakehi S (2012) PARFEX v1.0: an EXCEL™-based software package for parentage allocation. Conservation Genet Resour 4:275–278
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Nakamura Y, Mori K, Saitoh K, Oshima K, Mekuchi M, Sugaya T, Shigenobu Y, Ojima N, Muta S, Fujiwara A, Yasuike M, Oohara I, Hirakawa H, Chowdhury VS, Kobayashi T, Nakajima K, Sano M, Wada T, Tashiro K, Hattori M, Ikeo K, Kuhara S, Gojobori T, Inouye K (2013) Evolutionary changes of multiple visual pigment genes in the complete genome of Pacific bluefin tuna. Proc Natl Acad Sci USA 110(27):11061–11066
Chini V, Cattaneo AG, Rossi F, Bernardini G, Terova G, Saroglia M, Gornati R (2008) Genes expressed in Blue Fin Tuna (Thunnus thynnus) liver and gonads. Gene 29 410(1):207–213
Gangury A, Rock MJ, Prockop DJ (1993) Conformation-sensitive gel electrophoresis for rapid detection of single-base differences in double- stranded PCR products and DNA fragments: evidence for solvent-induced bends in DNA heteroduplexes. Proc Natl Acad Sci USA 90:10325–10329
Delwart EL, Sheppard HW, Walker BD, Goudsmit J, Mullins JI (1994) Human immunodeficiency virus type 1 evolution in vivo tracked by DNA heteroduplex mobility assays. J Virol 68:6672–6683
Takehana Y, Naruse K, Asada Y, Matsuda Y, Shin-I T, Kohara Y, Fujiyama A, Hamaguchi S, Sakaizumi M (2012) Molecular cloning and characterization of the repetitive DNA sequences that comprise the constitutive heterochromatin of the W chromosomes of medaka fishes. Chromosome Res 20:71–81
Sakamoto T, Danzmann RG, Gharbi K, Howard P, Ozaki A, Khoo SK, Woram RA, Okamoto N, Ferguson MM, Holm LE, Guyomard R, Hoyheim B (2000) A microsatellite linkage map of rainbow trout (Oncorhynchus mykiss) characterized by large sex-specific differences in recombination rates. Genetics 155:1331–1345
Chen SL, Li J, Deng SP, Tian YS, Wang QY, Zhuang ZM, Sha ZX, Xu JY (2007) Isolation of female-specific AFLP markers and molecular identification of genetic sex in half-smooth tongue sole (Cynoglossus semilaevis). Mar Biotechnol 9:273–280
Fuji K, Yoshida K, Hattori K, Ozaki A, Araki K, Okauchi M, Kubota S, Okamoto N, Sakamoto T (2010) Identifications of the sex-linked locus in yellowtail, Seriola quinqueradiata. Aquaculture 308:S51–S55
Kamiya T, Kai W, Tasumi S, Oka A, Matsunaga T, Mizuno N, Fujita M, Suetake H, Suzuki S, Hosoya S, Tohari S, Brenner S, Miyadai T, Venkatesh B, Suzuki Y, Kikuchi K (2012) A trans-species missense SNP in Amhr2 is associated with sex determination in the tiger pufferfish, Takifugu rubripes (Fugu). PLoS Genet 8(7):e1002798. doi:10.1371/journal.pgen.1002798
Acknowledgments
The authors thank Dr. E. Soeda and Ms. A. Kazama for assistance with AFLP analysis and PBF D-loop primer design. The authors also thank all the staff of the Oshima branch, especially Mr. S. Hayashi, Mr. N. Tsuro, Mr. Y. Hamaguchi, Mr. M. Nakatani, and Mr. T. Okada, for assistance with establishing the PBF blood and DNA specimen bank. This study was supported by a grant from the Japan Science and Technology Agency (JST) SATREPS program and Grant-in-Aid for Young Scientists B #25870937 provided from the Japan Society for the Promotion of Science (JSPS).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Agawa, Y., Iwaki, M., Komiya, T. et al. Identification of male sex-linked DNA sequence of the cultured Pacific bluefin tuna Thunnus orientalis . Fish Sci 81, 113–121 (2015). https://doi.org/10.1007/s12562-014-0833-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12562-014-0833-8