Abstract
The interpretation of biological data such as the ElectroCardioGram (ECG) signal gives clinical information and helps to assess the heart function. There are distinct ECG patterns associated with a specific class of arrhythmia. The convolutional neural network, inspired by findings in the study of biological vision, is currently one of the most commonly employed deep neural network algorithms for ECG processing. However, deep neural network models require many hyperparameters to tune. Selecting the optimal or the best hyperparameter for the convolutional neural network algorithm is a highly challenging task. Often, we end up tuning the model manually with different possible ranges of values until a best fit model is obtained. Automatic hyperparameters tuning using Bayesian Optimization (BO) and evolutionary algorithms can provide an effective solution to current labour-intensive manual configuration approaches. In this paper, we propose to optimize the Residual one Dimensional Convolutional Neural Network model (R-1D-CNN) at two levels. At the first level, a residual convolutional layer and one-dimensional convolutional neural layers are trained to learn patient-specific ECG features over which multilayer perceptron layers can learn to produce the final class vectors of each input. This level is manual and aims to limit the search space and select the most important hyperparameters to optimize. The second level is automatic and based on our proposed BO-based algorithm. Our optimized proposed architecture (BO-R-1D-CNN) is evaluated on two publicly available ECG datasets. Comparative experimental results demonstrate that our BO-based algorithm achieves an optimal rate of 99.95% for the MIT-BIH database to discriminate between five kinds of heartbeats, including normal heartbeats, left bundle branch block, atrial premature, right bundle branch block, and premature ventricular contraction. Moreover, experiments demonstrate that the proposed architecture fine-tuned with BO achieves a higher accuracy tested on the 10,000 ECG patients dataset compared to the other proposed architectures. Our optimized architecture achieves excellent results compared to previous works on the two benchmark datasets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The ECG is a non-invasive electrical recording of the heart. The signal provides a useful information about heart health and can tell more about individuals such as gender, age, biometry and emotion recognition. Researchers have explored this peripheral physiological signal to extract useful markers for future outcome research [1]. Several research works have been achieved in ECG analysis [2]. Challenges have been raised to provide an accurate ECG beats classification. In recent years, Deep Neural Network (DNN) is becoming increasingly an important research area. The structure of DNN tries to emulate the structure of human brain given a bulky dataset, fast enough processors and a sophisticated algorithm by constructing layers of artificial neurons that can receive and transmit information. The DNN offers accurate results with more training data. It is useful for unstructured data. Complex problems can be solved with a greater number of hidden layers. This structure makes possible to continuously adjust and make inferences. It outperforms the traditional machine learning in several applications such as the ElectroCardioGram (ECG) [3], ElectroEncephaloGraphy (EEG) [4] classification, more recently industry 4.0 [5] and COVID-19 detection [6, 7]. Recent researches provide many successful algorithms in DNN. The Convolutional Neural Network (with the acronyms CNN) is currently one of the commonly employed DNN algorithms for image recognition including detection of anomalies on ECG. Ebrahimi et al. [8] revealed that the CNN is dominantly found as the appropriate technique for feature extraction, observed in 52% of the studies about explainable DNN methods for ECG arrhythmia classification. The idea of CNN comes from the biological visual cortex. The cortex consists of small regions that are sensitive to specific areas of the visual field. Similarly, the CNN based on small regions inside of an object that perform specific tasks. The algorithm is a hierarchical neural network. It gets the input and processes it through a series of hidden layers. It is organized to automatically learn spatial hierarchies of features through backpropagation by employing different blocks, like convolution layers and pooling layers. The One-Dimension Convolutional Neural Network (1D-CNN) is a distinguished variant of CNN. It is typically used for the time series input with one direction x that represents the time axis. While they have achieved excellent results in working with a variety of hard problems [9, 10], the CNNs are usually exposed to overfitting or underfitting problems. Hence, the model fails to predict the output of unseen data or even the output of the training data. In fact, the noise introduced to the input signal slows the learning process. Various types of artifacts could lead to noisy ECG signals such as baseline wander, drift, powerline interference and muscle artifacts. The noisy signals lead to produce high false alarm rates thus the misclassification of ECG beats and misdiagnosis of cardiac arrhythmias. In addition, the ECG signals are non-stationary. Furthermore, the high number of parameters especially in fully connected layers makes the network prone to overfitting. Several previous works have proposed methods to boost classification results based on CNN hyperparameters and regularization. Regularizing the network structure or designing specific training schemes for stable and robust prediction is considered among the hottest topic for efficient and robust pattern recognition in the DNN [11]. A complex model may achieve a high performance on training data since all the inherent relations in seen data are memorized. However, the model is usually unable to perform well for unseen data including validation and test data. In order to solve this issue, different regularization methods were applied in the literature. Xu et al. [12] proposed SparseConnect to alleviate overfitting by sparsifying regularization on dense layers of CNNs. However, they raise, furthermore, the complexity of the model which in turn put the model harder to optimize. In our work, we choose to randomly dropping few nodes. Unlike conventional methods of tuning based on manual tries to choose the best hyperparameter value, our work proposes to use BO to select an optimal configuration of the dropout rate and the number of convolutional layers. In our proposal, two-level process has been established for building a robust Residual 1D-CNN (R-1D-CNN). The level one has the potential of reducing the search space of hyperparameters. The second level allows to test some configurations of the model. The innovative contributions associated with this work can be described as follows:
-
1.
We build a novel BO-R-1D-CNN architecture to detect features of the ECG automatically. The proposed model of biological vision presents good performance.
-
2.
To solve the overfitting issue and give robust classification results in real time through automatic hyperparamters tuning, we develop an algorithm based on BO.
-
3.
We further explore two datasets for experimental study. Our proposal outperforms another technique of optimization and almost the previous works in ECG classification, which displays the performance of our proposed architecture.
The rest of this paper is organized as follows: In the “Background’’ section, we outline a short background of the CNN and the BO technique. In the “Methods’’ section, the proposed architecture is detailed. The demonstration and performance of the proposed architecture are indicated in the “Results’’ section. At last, the “Conclusion’’ section concludes the paper and highlights the future work.
Background
Convolutional Neural Networks
Much of the current research on DNN has focused on improving and validating existing DNN algorithms rather developing new algorithms. The CNN is one of the commonly employed DNN algorithms. A CNN learns different levels of abstraction about an input. The CNN performs well in image processing, including image recognition and image classification thanks to its hierarchical layers. The hierarchical property allows to increasingly learn a complex model. For instance, the model learns in the first time basic elements, then it learns later their parts. Another advantage of CNN is the automatic extraction of features with minimal pre-processing operations [13]. The input of CNN is an array of pixels in the format of HxWxD where H = Height, W =Width and D = dimension. The HxW constitutes the feature map and D is the number of channels. A grey image of size 32x32 pixels is represented by an array 32X32X1 while an RGB image of the same size is represented by an array of 32X32X3. The structure may include convolutional layers hence its name, pooling layers, Rectified Linear Unit (ReLU) layers and fully connected layers.
-
Convolutional layer: the main layer of CNN. It consists of a set of filters that exploits the local spatial correlation assuming that near pixels are more correlated than distant pixels. The size of the filter defines the size of each feature map and its depth defines the number of feature maps. All local regions share the same weights called weight sharing. Mathematically saying, a convolution acts as a mixer, mixing two functions to obtain a reduced data space while preserving the information. The model involves training a multilayer architecture without the explicit need of handcrafted input features and is able to extract automatically the features such as edge, blur and sharpen. It helps to remove noise.
-
Pooling layer: common use is the max-pooling, which implements a sliding window. The max-pooling operation slides over the layer and takes pixels of the maximum value of each region with a step of stride vertically and horizontally.
-
ReLU: is a non-linear activation function. It performs a threshold operation. The output takes the same value as the input for the positive values and zero otherwise. The function is used by default for many DNN algorithms since it performs well and avoids a vanishing problem.
-
Fully connected or dense layer: in a fully connected layer, every neuron is connected to every neuron in the next layer. A model may contain one or more fully connected layer. The dense layer can be the last layer for the classification.
Based on the input, different convolutional dimensions can be used. A 1D-CNN is typically used for the time series input with one direction x that represents the time axis. Common uses of 1D-CNN are proposed for ECG data classification and anomaly detection [14]. A 2D-CNN performs well for image recognition and classification as the input is an image of 2 dimensions. The convolution is calculated based on two directions (x,y). With the increasing number of dimensions, a 3D-CNN applies a three-dimensional filter. The filter moves in three directions (x, y, z). The model is helpful in drug discovery [15].
Bayesian Optimization
The effective use of machine learning algorithms is associated with hyperparameters tuning. They adjust the model to a specific database and avoid ongoing training costs. To get up speed on hyperparameters tuning, BO can be used. The technique is based on Bayes’ theorem [16] to select the best configuration of hyperparameter values. The Bayes’ theorem consists of calculating the conditional probability of an event. The Bayes’ theorem uses prior probability distributions to be able to produce posterior probabilities. A prior probability could be the probability of an event before new knowledge is collected. The probability of A conditional on B is defined as Eq. 1.
Where:
-
P(A) = The probability of A occurring
-
P(B) = The probability of B occurring
-
P(A | B) = The probability of A given B
-
P(B | A) = The probability of B given A
-
P(A \(\bigcap\) B) = The probability of both A and B occurring
The BO provides a global optimal solution. By limiting the hyperparameters search space on ranges of values, the algorithm develops a probabilistic model of the objective function named the surrogate function. Mathematically saying, the algorithm is interested in solving Eq. 2:
This optimization method takes into account the problem of noise present in the evaluations of Eq. 3
where f is a black box and computationally expensive to evaluate.
Starting from default parameters, e.g. parameter ranges that are used in the literature, the performance evaluation calculated using a numeric score or a cost such as the accuracy rate. The aim is selecting a best configuration that maximizes or minimizes the cost. The best result achieved by a couple of hyperparameters would be used to construct the tuned model. Hence, the hyperparameters are assigned. For more details about hyperparameter optimization for machine learning models based on BO, please see [17]. The Table 1 presents different approaches worked on 1D-CNN and optimized by BO. The table presents mainly the selected hyperparameters, their search space and the obtained optimal values that are used in BO.
Methods
CNNs have been widely employed to improve the performance of ECG heartbeat classification [21]. Therefore, the proposed classifier is based on a CNN model. The first setting of hyperparameters is done manually. The process is iterative to accomplish an acceptable rate of accuracy. We add layers and nodes to the model gradually. The increasing layer number made the manual optimization harder. This configuration is given to the optimization algorithms as the default parameters and runs as the first iteration. By optimizing the neural network loss, the smoothing parameters are optimized to perform the prediction task. A novel BO-R-1D-CNN architecture is presented in our work. The optimization method is described below.
First Level: Architecture Building and Hyperparameters Selection
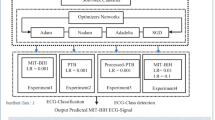
The primary hyperparameters employed for evaluation are the drop rate, the number of hidden layers, the learning rate and the Adam decay as they are more sensitive to variations in the classification performance. They are also widely used in many machine learning tasks [22]. In Table 2, a sample of models with different hyperparameters values by a large margin is displayed. The performance of the tested models is presented. In the iteration No. 3 where the classification performance is good in some parts, the F1 score of the proposed method is 80.94% while the accuracy and the recall are 86.06% and 86.06% respectively. Until this iteration, the different hyperparameters have been fixed except the number of hidden layers. As the network becomes deeper, residual connection [23] is introduced. Hence, a new level of depth is appeared in iteration No. 4. Deep residual learning comes with the benefit of solving the issue involved with vanishing/exploding gradients as well as the degradation problem. The residual network achieves this by employing skip connections, or shortcuts to leap across a number of layers. The number of residual blocks in our architecture is fixed to one after some trials. The skip connection is located in the position displayed as a residual block in Fig. 1. This new architecture allows an efficient training by including skip connections. In iteration No. 5, it is clear that the selected dropout rate hyperparameter has influence with the whole fixed hyperparameters. Moreover, the performance of the proposed method is more consistent with learning rate 0.001 than 0.000004 as displayed in iteration No. 6. The accuracy of iteration No. 7 is generally good throughout the records, therefore the corresponding values are selected.
Second Level: Architecture Optimization
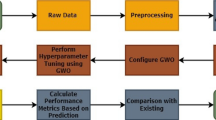
To enhance the architecture performance and avoid an overfitting problem, we choose to use the BO. The last is made using Bayesian inference and Gaussian process (GP). This approach is an appropriate algorithm to optimize hyperparameters of classification. By choosing which variables to optimize, and specifying the ranges to search in, the algorithm selects the optimal values. The GP is a well-known surrogate model for BO employed for approximating the objective function. It performs well in small dimensional spaces specifically when the number of features meets five features. The Table 3 illustrates the selecting hyperparameters, their type and their ranges. A DNN model is constructed according to the first level, and the most likely point to be maximized by the acquisition function is identified. Some hyperparameters that are very responsive to changes are chosen at the first level such as the learning rate. The BO enables fine-tuning of the model through the regularization of the penalty and determining the optimal number of layers.
Results
Setup
We used the python and its data science library to implement our algorithms. We implemented the algorithms using the Keras of TensorFlow library version 2.5 on a Tesla P100 GPU and 25 GB that are provided by Google Colaboratory Notebooks. The training set contains 70% of randomly selected beats and the rest is divided into test and validation sets. Each set contains 50% of the remained beats.
Database
Our proposed model is trained on two publicly available datasets: (1) The MIT-BIH dataset [24] that includes 48 ECG recordings of 30 min duration of 47 subjects and 250 sampling rate. Each record is annotated by specialists and can be utilized as ground truth for training, validation and the test. The collected data is preprocessed. We build a new dataset that consists of 82813 segments. For beat segmentation, we consider a fixed window multiple of frequency. The raw ECG signal doesn’t require any pre-filtering technique or a feature extraction step as used in traditional machine learning algorithms. The database is relatively noise free. Furthermore, the CNN is robust to the noise and features are automatically extracted during the learning process. (2) The second dataset is 10,000 ECG patients dataset [25]. This dataset consists of 10646 subjects of 10 s duration and 500 Hz sampling rate.
Performance
The performance of our proposed model was evaluated using several experiments. In the beginning, experiments are used to build an architecture to fit the MIT-BIH dataset and achieved 99.70%. Figures 2 and 3 illustrate the accuracy and the loss obtained during the training phase. The model runs 100 epochs with early stopped enabled. While the size of the input vector and the number of the hidden layers is large, the model converges in an extremely small time (8 epochs).
However, the gap between the validation and the training is significant.
At level two, we introduce the BO. A form of pseudo-code is written to provide precise descriptions of what the BO does. The pseudo-code is presented in Algorithm 1. The performance of the algorithm has increased. The BO produced an improvement right after the 13th iteration. The numerical experiments are showing that the resulting accuracy for the optimization with a finite budget outperforms the accuracy of the baseline model.

Finally, we build the optimized model for the test. Figures 4 and 5 illustrate the accuracy and loss obtained during the training phase. The training accuracy and training loss are respectively close to the validation accuracy and validation loss at second level. The confusion matrix is displayed in Fig. 6. Typically, the diagonal elements present the rate of items that are well predicted. Off-diagonal items are mislabelled. The proposed BO-R-1D-CNN properly predicted ECG signals of five distinct classes with a high accuracy of 99.95%. By reviewing the individual performance ratios of the different classes, we notice that the minimum recognition rate belongs to Left bundle branch block class (L) with 98%. The highest recognition performance is in the Atrial premature (A) and Normal (N) classes with 100%. The Right bundle branch block (R) and the premature Ventricular contraction (V) achieve 99%. The remainder of experiments aims at testing the model on different benchmark datasets and using two different optimization techniques. Other metrics are used to evaluate the classification performance. The Recall, the Precision and the F1 score can be calculated respectively by Eqs. 4–6
The Table 4 displays the achieved results. According to the table, the BO outperforms the PSO. The model fits well for MIT-BIH and 10,000 patients databases.
Discussion
Both of our architectures at level one and level two present novelties. We demonstrate that the proposed R-1D-CNN architecture, fine-tuned by the BO method is efficient compared to the state-of-the-art architectures. In our experiments, we exploit the BO-R-1D-CNN to classify the ECG signal of two databases. The Table 5 displays the classification accuracy of different architectures. A prior study has shown that ECG signals have been successfully learned by the AutoEncoder Long Short-Term Memory (AE-LSTM) [26]. Other recently developed convolutional architectures have been trained on similar databases. Nevertheless, our proposal achieves the highest accuracy.
Compared to the previous works on the MIT-BIH database, the CNN architecture provides an automatic extraction of features and does not require a pre-selection feature step. Hence, it is more generic. The computational complexity of the convolutional layer is \(o(knd^{2})\) [31] with n represents the sequence length, d is the representation dimension and k is the kernel size of convolutions. These attractive features may facilitate the application of the model in more real-time context, such as the Internet of Things (IoT) data analysis.
Conclusion
In this paper, we address optimization challenges for the R-1D-CNN model and propose a novel architecture for ECG analysis. In addition, we develop an algorithm based on BO to produce robust classification results in real time through automatic hyperparameters tuning. Comparative experimental results performed on two publicly available ECG Datasets demonstrate that our BO-based algorithm can outperform the state-of-art approaches. The BO achieves for instance an optimal rate of 99.95% for the MIT-BIH database. In the future, we plan to test the algorithms on other databases, especially for a dialysis application. We will also introduce other type of layers and classifiers to manage and optimize the complexity of the network.
Availability of Data and Materials
The ECG signals are obtained from the MIT-BIH arrhythmia database and the 12-lead electrocardiogram database for arrhythmia research covering more than 10,000 patients. All the databases are public and available online: Link to the MIT-BIH arrhythmia database: https://physionet.org/content/mitdb/1.0.0/. The original publication is referenced by [24]. Link to the 12-lead electrocardiogram database: https://doi.org/10.6084/m9.figshare.c.4560497.v2. The original publication is referenced by [25].
References
Fki Z, Ammar B, Ayed MB. Machine learning with internet of things data for risk prediction: Application in ESRD. In: 2018 12th International Conference on Research Challenges in Information Science (RCIS). 2018. https://doi.org/10.1109/RCIS.2018.8406669.
Panigrahy D, Sahu PK, Albu F. Detection of ventricular fibrillation rhythm by using boosted support vector machine with an optimal variable combination. Comput Electr Eng. 2021;91. https://doi.org/10.1016/j.compeleceng.2021.107035.
Cai W, Chen Y, Guo J, Han B, Shi Y, Ji L, Wang J, Zhang G, Luo J. Accurate detection of atrial fibrillation from 12-lead ECG using deep neural network. Comput Biol Med. 2020;116. https://doi.org/10.1016/j.compbiomed.2019.103378.
Fourati R, Ammar B, Sanchez-Medina J, Alimi AM. Unsupervised learning in reservoir computing for EEG-based emotion recognition. IEEE Trans Affect Comput. 2020. https://doi.org/10.1109/TAFFC.2020.2982143.
Zamora-Hernandez M-A, Castro-Vargas JA, Azorin-Lopez J, Garcia-Rodriguez J. Deep learning-based visual control assistant for assembly in industry 4.0. Comput Ind. 2021;131. https://doi.org/10.1016/j.compind.2021.103485.
Alzubaidi M, Zubaydi HD, Bin-Salem AA, Abd-Alrazaq AA, Ahmed A, Househ M. Role of deep learning in early detection of COVID-19: Scoping review. Comput Methods Programs Biomed Update. 2021;1. https://doi.org/10.1016/j.cmpbup.2021.100025.
Fang Z, Ren J, MacLellan C, Li H, Zhao H, Hussain A, Fortino G. A novel multi-stage residual feature fusion network for detection of COVID-19 in chest x-ray images. IEEE Trans Mol Biol Multi-Scale Commun. 2021. https://doi.org/10.1109/TMBMC.2021.3099367.
Ebrahimi Z, Loni M, Daneshtalab M, Gharehbaghi A. A review on deep learning methods for ECG arrhythmia classification. Expert Syst Appl. 2020;7. https://doi.org/10.1016/j.eswax.2020.100033.
Li F, Yan R, Mahini R, Wei L, Wang Z, Mathiak K, Liu R, Cong F. End-to-end sleep staging using convolutional neural network in raw single-channel EEG. Biomed Signal Process Control. 2021;63. https://doi.org/10.1016/j.bspc.2020.102203.
Moitra D, Mandal RK. Classification of non-small cell lung cancer using one-dimensional convolutional neural network. Expert Syst Appl. 2020;159. https://doi.org/10.1016/j.eswa.2020.113564.
Bai X, Wang X, Liu X, Liu Q, Song J, Sebe N, Kim B. Explainable deep learning for efficient and robust pattern recognition: a survey of recent developments. Pattern Recogn. 2021;120. https://doi.org/10.1016/j.patcog.2021.108102.
Xu Q, Zhang M, Gu Z, Pan G. Overfitting remedy by sparsifying regularization on fully-connected layers of CNNS. Neurocomputing. 2019;328. https://doi.org/10.1016/j.neucom.2018.03.080.
Gao F, Huang T, Sun J, Wang J, Hussain A, Yang E. A new algorithm for SAR image target recognition based on an improved deep convolutional neural network. Cogn Comput. 2019;11. https://doi.org/10.1007/s12559-018-9563-z.
Chen S, Yu J, Wang S. One-dimensional convolutional neural network-based active feature extraction for fault detection and diagnosis of industrial processes and its understanding via visualization. ISA Trans. 2021. https://doi.org/10.1016/j.isatra.2021.04.042.
Rezaei MA, Li Y, Wu D, Li X, Li C. Deep learning in drug design: Protein-ligand binding affinity prediction. IEEE/ACM Trans Comput Biol Bioinform. 2020. https://doi.org/10.1109/TCBB.2020.3046945.
Berrar D. Bayes’ theorem and naive Bayes classifier. In: Ranganathan S, Gribskov M, Nakai K, Schönbach C, editors. Encyclopedia of Bioinformatics and Computational Biology. 2019. https://doi.org/10.1016/B978-0-12-809633-8.20473-1.
Victoria AH, Maragatham G. Automatic tuning of hyperparameters using Bayesian optimization. Evol Syst. 2020;3. https://doi.org/10.1007/s12530-020-09345-2.
Sameen MI, Pradhan B, Lee S. Application of convolutional neural networks featuring Bayesian optimization for landslide susceptibility assessment. Catena. 2020;186. https://doi.org/10.1016/j.catena.2019.104249.
Shi D, Ye Y, Gillwald M, Hecht M. Designing a lightweight 1D convolutional neural network with Bayesian optimization for wheel flat detection using carbody accelerations. Int J Rail Transp. 2020. https://doi.org/10.1080/23248378.2020.1795942.
Ragab MG, Abdulkadir SJ, Aziz N, Alhussian H, Bala A, Alqushaibi A. An ensemble one dimensional convolutional neural network with Bayesian optimization for environmental sound classification. Appl Sci. 2021;11. https://doi.org/10.3390/app11104660.
Jangra M, Dhull SK, Singh KK. ECG beat classifiers: A journey from ANN to DNN. Prog Comput Sci. 2020;167. https://doi.org/10.1016/j.procs.2020.03.340. International Conference on Computational Intelligence and Data Science.
Tong Q, Liang G, Bi J. Calibrating the adaptive learning rate to improve convergence of ADAM. Neurocomputing. 2022;481. https://doi.org/10.1016/j.neucom.2022.01.014.
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. In: 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). 2016. https://doi.org/10.1109/CVPR.2016.90.
Moody GB, Mark RG. The impact of the MIT-BIH arrhythmia database. IEEE Eng Med Biol Mag. 2001;20. https://doi.org/10.1109/51.932724.
Jianwei Z, Jianming Z, Sidy D, Hai Y, Hangyuan G, Cyril R. A 12-lead electrocardiogram database for arrhythmia research covering more than 10,000 patients. Sci Data. 2020;7. https://doi.org/10.1038/s41597-020-0386-x.
Yildirim O, Baloglu UB, Tan R-S, Ciaccio EJ, Acharya UR. A new approach for arrhythmia classification using deep coded features and LSTM networks. Comput Methods Prog Biomed. 2019;176. https://doi.org/10.1016/j.cmpb.2019.05.004.
Nurmaini S, Tondas AE, Darmawahyuni A, Rachmatullah MN, Effendi J, Firdaus F, Tutuko B. Electrocardiogram signal classification for automated delineation using bidirectional long short-term memory. Inform Med Unlocked. 2021;22. https://doi.org/10.1016/j.imu.2020.100507.
Rath A, Mishra D, Panda G, Satapathy SC. Heart disease detection using deep learning methods from imbalanced ECG samples. Biomed Signal Process Control. 2021;68. https://doi.org/10.1016/j.bspc.2021.102820.
Murat F, Yildirim O, Talo M, Demir Y, Tan R-S, Ciaccio EJ, Acharya UR. Exploring deep features and ECG attributes to detect cardiac rhythm classes. Knowl-Based Syst. 2021;232. https://doi.org/10.1016/j.knosys.2021.107473.
Li Y, Qian R, Li K. Inter-patient arrhythmia classification with improved deep residual convolutional neural network. Comput Methods Prog Biomed. 2022;214. https://doi.org/10.1016/j.cmpb.2021.106582.
Zhang A, Lipton ZC, Li M, Smola AJ. Dive into deep learning. arXiv:2106.11342 [Preprint]. 2021. Available from http://arxiv.org/abs/2106.11342. https://d2l.ai/.
Funding
The research leading to these results has received funding from the Ministry of Higher Education and Scientific Research of Tunisia under the grant agreement number LR11ES48.
Author information
Authors and Affiliations
Contributions
Zeineb Fki: developed the method, collected the data, performed the experiments and drafted the manuscript. Boudour Ammar: revised the software, interpreted the results and supervised the project. Mounir Ben Ayed: conceived the study and supervised the project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed Consent
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fki, Z., Ammar, B. & Ayed, M.B. Towards Automated Optimization of Residual Convolutional Neural Networks for Electrocardiogram Classification. Cogn Comput (2023). https://doi.org/10.1007/s12559-022-10103-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12559-022-10103-6