Abstract
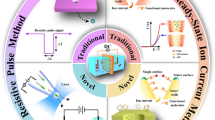
Solid-state nanopore (SSNP) or synthetic nanopore using semiconductor materials have established themselves as a single molecule bio-detection platform. Although biological nanopore with fixed dimension has been successfully utilized for many sensing applications, SSNP has unique characteristics of distinctly potent geometries and relaxation of modification. The most common method of molecular detection is to measure the temporal variations of the ionic current in the pore. In this review, the principles of the SSNP and the improvement of device performance for the molecular detection platform are discussed elaborately. Moreover, different experimental aspects of the SSNP are discussed in detail. For instance, the enhancement of spatial resolution, modification of temporal resolution with the difficulties of its analyte-detection, as well as reduction of the electrical noise for the improvement of device sensing functionalities, all are addressed in designated chapters for better conception. In addition, the typical and updated applications of SSNP including DNA, protein and virus are briefly discussed. Finally, this article offers the context needed to comprehend current research trends and promote molecular sensing through synthetic nanopores.









Similar content being viewed by others
References
Chen, Y., & Pepin, A. (2001). Nanofabrication: Conventional and nonconventional methods. Electrophoresis, 22(2), 187–207.
Zhu, L., Xu, Y., Ali, I., Liu, L., Wu, H., Lu, Z., & Liu, Q. (2018). Solid-state nanopore single-molecule sensing of DNAzyme cleavage reaction assisted with nucleic acid nanostructure. ACS Applied Materials and Interfaces, 10(31), 26555–26565.
Dong, G., Hou, J., Wang, J., Zhang, Y., Chen, V., & Liu, J. (2016). Enhanced CO2/N2 separation by porous reduced graphene oxide/Pebax mixed matrix membranes. Journal of Membrane Science, 520, 860–868.
Caglar, M., Silkina, I., Brown, B. T., Thorneywork, A. L., Burton, O. J., Babenko, V., Gilbert, S. M., Zettl, A., Hofmann, S., & Keyser, U. F. (2019). Tunable anion-selective transport through monolayer graphene and hexagonal boron nitride. ACS Nano, 14(3), 2729–2738.
Liu, Y., Zhang, Z., & Wang, S. (2020). Carbon nanopore-tailored reverse osmotic water desalination. ACS ES&T Water, 1(1), 34–47.
Cao, L., Wen, Q., Feng, Y., Ji, D., Li, H., Li, N., Jiang, L., & Guo, W. (2018). On the origin of ion selectivity in ultrathin nanopores: Insights for membrane-scale osmotic energy conversion. Advanced Functional Materials, 28(39), 1804189.
Feng, J., Graf, M., Liu, K., Ovchinnikov, D., Dumcenco, D., Heiranian, M., Nandigana, V., Aluru, N. R., Kis, A., & Radenovic, A. (2016). Single-layer MoS2 nanopores as nanopower generators. Nature, 536(7615), 197–200.
Lin, Y., Ying, Y. L., & Long, Y. T. (2018). Nanopore confinement for electrochemical sensing at the single-molecule level. Current Opinion in Electrochemistry, 7, 172–178.
Wanunu, M., Dadosh, T., Ray, V., Jin, J., McReynolds, L., & Drndić, M. (2010). Rapid electronic detection of probe-specific microRNAs using thin nanopore sensors. Nature nanotechnology, 5(11), 807–814.
Akeson, M., Branton, D., Kasianowicz, J. J., Brandin, E., & Deamer, D. W. (1999). Microsecond time-scale discrimination among polycytidylic acid, polyadenylic acid, and polyuridylic acid as homopolymers or as segments within single RNA molecules. Biophysical Journal, 77(6), 3227–3233.
Lee, M. H., Kumar, A., Park, K. B., Cho, S. Y., Kim, H. M., Lim, M. C., Kim, Y. R., & Kim, K. B. (2014). A low-noise solid-state nanopore platform based on a highly insulating substrate. Scientific Reports, 4(1), 1–7.
Yilmaz, D., Kaya, D., Kececi, K., & Dinler, A. (2021). Role of nanopore geometry in particle resolution by resistive-pulse sensing. ChemistrySelect, 6(1), 59–67.
Ivanov, A. P., Instuli, E., McGilvery, C. M., Baldwin, G., McComb, D. W., Albrecht, T., & Edel, J. B. (2011). DNA tunneling detector embedded in a nanopore. Nano Letters, 11(1), 279–285.
Gilboa, T., & Meller, A. (2015). Optical sensing and analyte manipulation in solid-state nanopores. The Analyst, 140(14), 4733–4747.
Crosland-Taylor, P. J. (1953). A device for counting small particles suspended in a fluid through a tube. Nature, 171(4340), 37–38.
Muthukumar, M., Plesa, C., & Dekker, C. (2015). Single-molecule sensing with nanopores. Physics Today, 68(8), 40.
Kasianowicz, J. J., Brandin, E., Branton, D., & Deamer, D. W. (1996). Characterization of individual polynucleotide molecules using a membrane channel. Proceedings of the National Academy of Sciences, 93(24), 13770–13773.
Faller, M., Niederweis, M., & Schulz, G. E. (2004). The structure of a mycobacterial outer-membrane channel. Science, 303(5661), 1189–1192.
Rhee, M., & Burns, M. A. (2007). Nanopore sequencing technology: Nanopore preparations. TRENDS in Biotechnology, 25(4), 174–181.
Liu, K., Pan, C., Kuhn, A., Nievergelt, A. P., Fantner, G. E., Milenkovic, O., & Radenovic, A. (2019). Detecting topological variations of DNA at single-molecule level. Nature Communications, 10(1), 1–9.
Storm, A. J., Chen, J. H., Ling, X. S., Zandbergen, H. W., & Dekker, C. (2003). Fabrication of solid-state nanopores with single-nanometre precision. Nature Materials, 2(8), 537–540.
Hall, A. R., Scott, A., Rotem, D., Mehta, K. K., Bayley, H., & Dekker, C. (2010). Hybrid pore formation by directed insertion of α-haemolysin into solid-state nanopores. Nature Nanotechnology, 5(12), 874–877.
Zhu, L., Zhang, Z., & Liu, Q. (2020). Deformation-mediated translocation of DNA origami nanoplates through a narrow solid-state nanopore. Analytical Chemistry, 92(19), 13238–13245.
Steinbock, L. J., Krishnan, S., Bulushev, R. D., Borgeaud, S., Blokesch, M., Feletti, L., & Radenovic, A. (2014). Probing the size of proteins with glass nanopores. Nanoscale, 6(23), 14380–14387.
Yusko, E. C., Bruhn, B. R., Eggenberger, O. M., Houghtaling, J., Rollings, R. C., Walsh, N. C., Nandivada, S., Pindrus, M., Hall, A. R., Sept, D., & Li, J. (2017). Real-time shape approximation and fingerprinting of single proteins using a nanopore. Nature Nanotechnology, 12(4), 360–367.
Houghtaling, J., Ying, C., Eggenberger, O. M., Fennouri, A., Nandivada, S., Acharjee, M., Li, J., Hall, A. R., & Mayer, M. (2019). Estimation of shape, volume, and dipole moment of individual proteins freely transiting a synthetic nanopore. ACS Nano, 13(5), 5231–5242.
Wei, R., Gatterdam, V., Wieneke, R., Tampé, R., & Rant, U. (2012). Stochastic sensing of proteins with receptor-modified solid-state nanopores. Nature Nanotechnology, 7(4), 257–263.
Storm, A. J., Chen, J. H., Zandbergen, H. W., & Dekker, C. (2005). Translocation of double-strand DNA through a silicon oxide nanopore. Physical Review E, 71(5), 051903.
Yanagi, I., Hamamura, H., Akahori, R., & Takeda, K. I. (2018). Two-step breakdown of a SiN membrane for nanopore fabrication: Formation of thin portion and penetration. Scientific Reports, 8(1), 1–13.
Yanagi, I., Ishida, T., Fujisaki, K., & Takeda, K. I. (2015). Fabrication of 3-nm-thick Si3N4 membranes for solid-state nanopores using the poly-Si sacrificial layer process. Scientific Reports, 5(1), 1–13.
Waduge, P., Hu, R., Bandarkar, P., Yamazaki, H., Cressiot, B., Zhao, Q., Whitford, P. C., & Wanunu, M. (2017). Nanopore-based measurements of protein size, fluctuations, and conformational changes. ACS Nano, 11(6), 5706–5716.
Ying, Y. L., Zhang, J., Gao, R., & Long, Y. T. (2013). Nanopore-based sequencing and detection of nucleic acids. Angewandte Chemie International Edition, 52(50), 13154–13161.
Feng, J., Liu, K., Bulushev, R. D., Khlybov, S., Dumcenco, D., Kis, A., & Radenovic, A. (2015). Identification of single nucleotides in MoS2 nanopores. Nature Nanotechnology, 10(12), 1070–1076.
Rusk, N. (2015). MinION takes center stage. Nature Methods, 12(1), 12–12.
Dekker, C. (2007). Solid-state nanopores. Nature Nanotechnology, 2, 209–215.
Venta, K., Shemer, G., Puster, M., Rodriguez-Manzo, J. A., Balan, A., Rosenstein, J. K., Shepard, K., & Drndic, M. (2013). Differentiation of short, single-stranded DNA homopolymers in solid-state nanopores. ACS Nano, 7(5), 4629–4636.
Rosenstein, J. K., Wanunu, M., Merchant, C. A., Drndic, M., & Shepard, K. L. (2012). Integrated nanopore sensing platform with sub-microsecond temporal resolution. Nature Methods, 9(5), 487–492.
Weber, M., Julbe, A., Ayral, A., Miele, P. & Bechelany, M. (2018). Atomic layer deposition for membranes: Basics, challenges, and opportunities. Chemistry of Materials, 30(21), 7368–7390.
Balan, A., Chien, C. C., Engelke, R., & Drndić, M. (2015). Suspended solid-state membranes on glass chips with sub 1-pF capacitance for biomolecule sensing applications. Scientific Reports, 5(1), 1–8.
Chen, P., Mitsui, T., Farmer, D. B., Golovchenko, J., Gordon, R. G., & Branton, D. (2004). Atomic layer deposition to fine-tune the surface properties and diameters of fabricated nanopores. Nano Letters, 4(7), 1333–1337.
Larkin, J., Henley, R., Bell, D. C., Cohen-Karni, T., Rosenstein, J. K., & Wanunu, M. (2013). Slow DNA transport through nanopores in hafnium oxide membranes. ACS Nano, 7(11), 10121–10128.
Banerjee, S., Wilson, J., Shim, J., Shankla, M., Corbin, E. A., Aksimentiev, A., & Bashir, R. (2015). Slowing DNA transport using graphene–DNA interactions. Advanced Functional Materials, 25(6), 936–946.
Waduge, P., Bilgin, I., Larkin, J., Henley, R. Y., Goodfellow, K., Graham, A. C., Bell, D. C., Vamivakas, N., Kar, S., & Wanunu, M. (2015). Direct and scalable deposition of atomically thin low-noise MoS2 membranes on apertures. ACS Nano, 9(7), 7352–7359.
Liu, S., Lu, B., Zhao, Q., Li, J., Gao, T., Chen, Y., Zhang, Y., Liu, Z., Fan, Z., & Yang, F. (2013). Boron nitride nanopores: Highly sensitive DNA single-molecule detectors. Advanced Materials, 25(33), 4549–4554.
Zhou, Z., Hu, Y., Wang, H., Xu, Z., Wang, W., Bai, X., Shan, X., & Lu, X. (2013). DNA translocation through hydrophilic nanopore in hexagonal boron nitride. Scientific Reports, 3(1), 1–5.
Danda, G., Masih Das, P., Chou, Y. C., Mlack, J. T., Parkin, W. M., Naylor, C. H., Fujisawa, K., Zhang, T., Fulton, L. B., Terrones, M., & Johnson, A. T. (2017). Monolayer WS2 nanopores for DNA translocation with light-adjustable sizes. ACS Nano, 11(2), 1937–1945.
Li, J., Stein, D., McMullan, C., Branton, D., Aziz, M. J., & Golovchenko, J. A. (2001). Ion-beam sculpting at nanometre length scales. Nature, 412(6843), 166–169.
Rigo, E., Dong, Z., Park, J. H., Kennedy, E., Hokmabadi, M., Almonte-Garcia, L., Ding, L., Aluru, N., & Timp, G. (2019). Measurements of the size and correlations between ions using an electrolytic point contact. Nature Communications, 10(1), 1–13.
Kwok, H., Briggs, K., & Tabard-Cossa, V. (2014). Nanopore fabrication by controlled dielectric breakdown. PLoS ONE, 9(3), e92880.
Yanagi, I., Akahori, R., Hatano, T., & Takeda, K. I. (2014). Fabricating nanopores with diameters of sub-1 nm to 3 nm using multilevel pulse-voltage injection. Scientific Reports, 4(1), 1–7.
Waugh, M., Briggs, K., Gunn, D., Gibeault, M., King, S., Ingram, Q., Jimenez, A. M., Berryman, S., Lomovtsev, D., Andrzejewski, L., & Tabard-Cossa, V. (2020). Solid-state nanopore fabrication by automated controlled breakdown. Nature Protocols, 15(1), 122–143.
Harrell, C. C., Choi, Y., Horne, L. P., Baker, L. A., Siwy, Z. S., & Martin, C. R. (2006). Resistive-pulse DNA detection with a conical nanopore sensor. Langmuir, 22(25), 10837–10843.
Yamazaki, H., Hu, R., Zhao, Q., & Wanunu, M. (2018). Photothermally assisted thinning of silicon nitride membranes for ultrathin asymmetric nanopores. ACS Nano, 12(12), 12472–12481.
Sriram, G., Patil, P., Bhat, M. P., Hegde, R. M., Ajeya, K. V., Udachyan, I., Bhavya, M. B., Gatti, M. G., Uthappa, U. T., Neelgund, G. M., & Jung, H. Y. (2016). Current trends in nanoporous anodized alumina platforms for biosensing applications. Journal of Nanomaterials, 2016, 1–14.
De Vreede, L. J., Muniz, M. S., van den Berg, A., & Eijkel, J. C. (2016). Nanopore fabrication in silicon oxynitride membranes by heating Au-particles. Journal of Micromechanics and Microengineering, 26(3), 037001.
Deng, T., Chen, J., Li, M., Wang, Y., Zhao, C., Zhang, Z., & Liu, Z. (2013). Controllable shrinking of inverted-pyramid silicon nanopore arrays by dry-oxygen oxidation. Nanotechnology, 24(50), 505303.
Wei, C., Bard, A. J., & Feldberg, S. W. (1997). Current rectification at quartz nanopipet electrodes. Analytical Chemistry, 69(22), 4627–4633.
Sun, L., Shigyou, K., Ando, T., & Watanabe, S. (2019). Thermally driven approach to fill sub-10-nm pipettes with batch production. Analytical Chemistry, 91(21), 14080–14084.
Steinbock, L. J., Otto, O., Chimerel, C., Gornall, J., & Keyser, U. F. (2010). Detecting DNA folding with nanocapillaries. Nano Letters, 10(7), 2493–2497.
Yuan, Z., Wang, C., Yi, X., Ni, Z., Chen, Y., & Li, T. (2018). Solid-state nanopore. Nanoscale Research Letters, 13(1), 1–10.
Ma, J., Li, K., Li, Z., Qiu, Y., Si, W., Ge, Y., Sha, J., Liu, L., Xie, X., Yi, H., & Ni, Z. (2019). Drastically reduced ion mobility in a nanopore due to enhanced pairing and collisions between dehydrated ions. Journal of the American Chemical Society, 141(10), 4264–4272.
Yanagi, I., Akahori, R., & Takeda, K. I. (2019). Stable fabrication of a large nanopore by controlled dielectric breakdown in a high-pH solution for the detection of various-sized molecules. Scientific Reports, 9(1), 1–15.
Hayashi, T., Arima, K., Yamashita, N., Park, S., Ma, Z., Tabata, O., & Kawai, K. (2018). Nanopore fabrication of two-dimensional materials on SiO2 membranes using he ion microscopy. IEEE Transactions on Nanotechnology, 17(4), 727–730.
Zahid, O. K., Wang, F., Ruzicka, J. A., Taylor, E. W., & Hall, A. R. (2016). Sequence-specific recognition of microRNAs and other short nucleic acids with solid-state nanopores. Nano Letters, 16(3), 2033–2039.
Henley, R. Y., Ashcroft, B. A., Farrell, I., Cooperman, B. S., Lindsay, S. M., & Wanunu, M. (2016). Electrophoretic deformation of individual transfer RNA molecules reveals their identity. Nano Letters, 16(1), 138–144.
Langecker, M., Ivankin, A., Carson, S., Kinney, S. R., Simmel, F. C., & Wanunu, M. (2015). Nanopores suggest a negligible influence of CpG methylation on nucleosome packaging and stability. Nano Letters, 15(1), 783–790.
Singer, A., Rapireddy, S., Ly, D. H., & Meller, A. (2012). Electronic barcoding of a viral gene at the single-molecule level. Nano Letters, 12(3), 1722–1728.
Yu, J. S., Hong, S. C., Wu, S., Kim, H. M., Lee, C., Lee, J. S., Lee, J. E., & Kim, K. B. (2019). Differentiation of selectively labeled peptides using solid-state nanopores. Nanoscale, 11(5), 2510–2520.
Zhao, X., Ma, R., Hu, Y., Chen, X., Dou, R., Liu, K., Cui, C., Liu, H., Li, Q., Pan, D., & Shan, X. (2019). Translocation of tetrahedral DNA nanostructures through a solid-state nanopore. Nanoscale, 11(13), 6263–6269.
Carlsen, A. T., Zahid, O. K., Ruzicka, J. A., Taylor, E. W., & Hall, A. R. (2014). Selective detection and quantification of modified DNA with solid-state nanopores. Nano Letters, 14(10), 5488–5492.
Plesa, C., Ruitenberg, J. W., Witteveen, M. J., & Dekker, C. (2015). Detection of individual proteins bound along DNA using solid-state nanopores. Nano Letters, 15(5), 3153–3158.
Squires, A., Atas, E., & Meller, A. (2015). Nanopore sensing of individual transcription factors bound to DNA. Scientific Reports, 5(1), 1–11.
Yu, J. S., Lim, M. C., Huynh, D. T. N., Kim, H. J., Kim, H. M., Kim, Y. R., & Kim, K. B. (2015). Identifying the location of a single protein along the DNA strand using solid-state nanopores. ACS Nano, 9(5), 5289–5298.
Asghar, W., Ilyas, A., Billo, J. A., & Iqbal, S. M. (2011). Shrinking of solid-state nanopores by direct thermal heating. Nanoscale Research Letters, 6(1), 1–6.
Li, Q., Xie, S., Liang, Z., Meng, X., Liu, S., Girault, H. H., & Shao, Y. (2009). fast ion-transfer processes at nanoscopic liquid/liquid interfaces. Angewandte Chemie International Edition, 48(43), 8010–8013.
Hu, R., Rodrigues, J. V., Waduge, P., Yamazaki, H., Cressiot, B., Chishti, Y., Makowski, L., Yu, D., Shakhnovich, E., Zhao, Q., & Wanunu, M. (2018). Differential enzyme flexibility probed using solid-state nanopores. ACS Nano, 12(5), 4494–4502.
Darvish, A., Lee, J. S., Peng, B., Saharia, J., VenkatKalyana Sundaram, R., Goyal, G., Bandara, N., Ahn, C. W., Kim, J., Dutta, P., & Chaiken, I. (2019). Mechanical characterization of HIV-1 with a solid-state nanopore sensor. Electrophoresis, 40(5), 776–783.
McMullen, A., De Haan, H. W., Tang, J. X., & Stein, D. (2014). Stiff filamentous virus translocations through solid-state nanopores. Nature Communications, 5(1), 1–10.
Oukhaled, A., Cressiot, B., Bacri, L., Pastoriza-Gallego, M., Betton, J. M., Bourhis, E., Jede, R., Gierak, J., Auvray, L., & Pelta, J. (2011). Dynamics of completely unfolded and native proteins through solid-state nanopores as a function of electric driving force. ACS Nano, 5(5), 3628–3638.
Larkin, J., Henley, R. Y., Jadhav, V., Korlach, J., & Wanunu, M. (2017). Length-independent DNA packing into nanopore zero-mode waveguides for low-input DNA sequencing. Nature Nanotechnology, 12(12), 1169–1175.
Venkatesan, B. M., Dorvel, B., Yemenicioglu, S., Watkins, N., Petrov, I., & Bashir, R. (2009). Highly sensitive, mechanically stable nanopore sensors for DNA analysis. Advanced Materials, 21(27), 2771–2776.
Venkatesan, B. M., Estrada, D., Banerjee, S., Jin, X., Dorgan, V. E., Bae, M. H., Aluru, N. R., Pop, E., & Bashir, R. (2012). Stacked graphene–Al2O3 nanopore sensors for sensitive detection of DNA and DNA–protein complexes. ACS Nano, 6(1), 441–450.
Park, K. B., Kim, H. J., Kang, Y. H., Yu, J. S., Chae, H., Lee, K., Kim, H. M., & Kim, K. B. (2017). Highly reliable and low-noise solid-state nanopores with an atomic layer deposited ZnO membrane on a quartz substrate. Nanoscale, 9(47), 18772–18780.
Lee, J. S., Oviedo, J. P., Bandara, Y. M., Peng, X., Xia, L., Wang, Q., Garcia, K., Wang, J., Kim, M. J., & Kim, M. J. (2021). Detection of nucleotides in hydrated ssDNA via 2D h-BN nanopore with ionic-liquid/salt-water interface. Electrophoresis, 42(7–8), 991–1002.
Garaj, S., Hubbard, W., Reina, A., Kong, J., Branton, D., & Golovchenko, J. A. (2010). Graphene as a subnanometre trans-electrode membrane. Nature, 467(7312), 190–193.
Schneider, G. F., Xu, Q., Luik, S., Hage, S., Spoor, J. N., Malladi, S., Zandbergen, H. W., & Dekker, C. (2013). Tailoring the surface chemistry and hydrophobicity of graphene nanopores. Nature Communications, 4, 2619.
Harrell, C. C., Lee, S. B., & Martin, C. R. (2003). Synthetic single-nanopore and nanotube membranes. Analytical Chemistry, 75(24), 6861–6867.
Liu, K., Feng, J., Kis, A., & Radenovic, A. (2014). Atomically thin molybdenum disulfide nanopores with high sensitivity for DNA translocation. ACS Nano, 8(3), 2504–2511.
Park, K. B., Kim, H. J., Kim, H. M., Han, S. A., Lee, K. H., Kim, S. W., & Kim, K. B. (2016). Noise and sensitivity characteristics of solid-state nanopores with a boron nitride 2-D membrane on a pyrex substrate. Nanoscale, 8(10), 5755–5763.
Prabhu, A. S., Freedman, K. J., Robertson, J. W., Nikolov, Z., Kasianowicz, J. J., & Kim, M. J. (2011). SEM-induced shrinking of solid-state nanopores for single molecule detection. Nanotechnology, 22(42), 425302.
Storm, A. J., Storm, C., Chen, J., Zandbergen, H., Joanny, J. F., & Dekker, C. (2005). Fast DNA translocation through a solid-state nanopore. Nano letters, 5(7), 1193–1197.
Goyal, G., Lee, Y. B., Darvish, A., Ahn, C. W., & Kim, M. J. (2016). Hydrophilic and size-controlled graphene nanopores for protein detection. Nanotechnology, 27(49), 495301.
Beamish, E., Tabard-Cossa, V., & Godin, M. (2017). Identifying structure in short DNA scaffolds using solid-state nanopores. ACS Sensors, 2(12), 1814–1820.
Lin, Y., Ying, Y. L., Shi, X., Liu, S. C., & Long, Y. T. (2017). Direct sensing of cancer biomarkers in clinical samples with a designed nanopore. Chemical Communications, 53(84), 11564–11567.
Carlsen, A. T., Briggs, K., Hall, A. R., & Tabard-Cossa, V. (2017). Solid-state nanopore localization by controlled breakdown of selectively thinned membranes. Nanotechnology, 28(8), 085304.
Gilboa, T., Zrehen, A., Girsault, A., & Meller, A. (2018). Optically-monitored nanopore fabrication using a focused laser beam. Scientific Reports, 8(1), 1–10.
Gilboa, T., Zvuloni, E., Zrehen, A., Squires, A. H., & Meller, A. (2020). Automated, ultra-fast laser-drilling of nanometer scale pores and nanopore arrays in aqueous solutions. Advanced Functional Materials, 30(18), 1900642.
Briggs, K., Kwok, H., & Tabard-Cossa, V. (2014). Automated fabrication of 2-nm solid-state nanopores for nucleic acid analysis. Small (Weinheim an der Bergstrasse, Germany), 10(10), 2077–2086.
Wang, Y., Chen, Q., Deng, T., & Liu, Z. (2018). Self-aligned nanopore formed on a SiO2 pyramidal membrane by a multipulse dielectric breakdown method. The Journal of Physical Chemistry C, 122(21), 11516–11523.
Zhang, X., van Deursen, P. M., Fu, W., & Schneider, G. F. (2020). Facile and ultraclean graphene-on-glass nanopores by controlled electrochemical etching. ACS Sensors, 5(8), 2317–2325.
Kong, J., Zhu, J., Chen, K., & Keyser, U. F. (2019). Specific biosensing using DNA aptamers and nanopores. Advanced Functional Materials, 29(3), 1807555.
Loh, A. Y. Y., Burgess, C. H., Tanase, D. A., Ferrari, G., McLachlan, M. A., Cass, A. E. G., & Albrecht, T. (2018). Electric single-molecule hybridization detector for short DNA fragments. Analytical Chemistry, 90(23), 14063–14071.
Weckman, N. E., Ermann, N., Gutierrez, R., Chen, K., Graham, J., Tivony, R., Heron, A., & Keyser, U. F. (2019). Multiplexed DNA identification using site specific dCas9 barcodes and nanopore sensing. ACS Sensors, 4(8), 2065–2072.
Cai, S., Sze, J. Y., Ivanov, A. P., & Edel, J. B. (2019). Small molecule electro-optical binding assay using nanopores. Nature Communications, 10(1), 1–9.
Bafna, J. A., & Soni, G. V. (2016). Fabrication of low noise borosilicate glass nanopores for single molecule sensing. PLoS ONE, 11(6), e0157399.
Zhang, B., Zhang, Y., & White, H. S. (2004). The nanopore electrode. Analytical Chemistry, 76(21), 6229–6238.
Li, W., Bell, N. A., Hernández-Ainsa, S., Thacker, V. V., Thackray, A. M., Bujdoso, R., & Keyser, U. F. (2013). Single protein molecule detection by glass nanopores. ACS Nano, 7(5), 4129–4134.
Sze, J. Y., Ivanov, A. P., Cass, A. E., & Edel, J. B. (2017). Single molecule multiplexed nanopore protein screening in human serum using aptamer modified DNA carriers. Nature Communications, 8(1), 1–10.
Kubánková, M., Lin, X., Albrecht, T., Edel, J. B., & Kuimova, M. K. (2019). Rapid fragmentation during seeded lysozyme aggregation revealed at the single molecule level. Analytical Chemistry, 91(10), 6880–6886.
Bell, N. A., & Keyser, U. F. (2016). Digitally encoded DNA nanostructures for multiplexed, single-molecule protein sensing with nanopores. Nature Nanotechnology, 11(7), 645–651.
Henderson, P. (1978). Fleischer, (PB) Price, and (RM) Walker. Nuclear tracks in solids: Principles and applications. Berkeley and London (Univ. California Press), 1975. xxii+ 605 pp., 205 figs., I pl. Price. Mineralogical Magazine, 42(322), 306–307.
Siwy, Z., Dobrev, D., Neumann, R., Trautmann, C., & Voss, K. (2003). Electro-responsive asymmetric nanopores in polyimide with stable ion-current signal. Applied Physics A, 76(5), 781–785.
Wu, M. Y., Smeets, R. M., Zandbergen, M., Ziese, U., Krapf, D., Batson, P. E., Dekker, N. H., Dekker, C., & Zandbergen, H. W. (2009). Control of shape and material composition of solid-state nanopores. Nano Letters, 9, 479–484.
Van den Hout, M., Hall, A. R., Wu, M. Y., Zandbergen, H. W., Dekker, C., & Dekker, N. H. (2010). Controlling nanopore size, shape and stability. Nanotechnology, 21(11), 115304.
Emmrich, D., Beyer, A., Nadzeyka, A., Bauerdick, S., Meyer, J. C., Kotakoski, J., & Gölzhäuser, A. (2016). Nanopore fabrication and characterization by helium ion microscopy. Applied Physics Letters, 108(16), 163103.
Wu, M. Y., Smeets, R. M., Zandbergen, M., Ziese, U., Krapf, D., Batson, P. E., Dekker, N. H., Dekker, C., & Zandbergen, H. W. (2007). Sub-5 nm FIB direct patterning of nanodevices. Microelectronic Engineering, 84(5–8), 779–783.
Lo, C. J., Aref, T., & Bezryadin, A. (2006). Fabrication of symmetric sub-5 nm nanopores using focused ion and electron beams. Nanotechnology, 17(13), 3264.
Sawafta, F., Carlsen, A. T., & Hall, A. R. (2014). Membrane thickness dependence of nanopore formation with a focused helium ion beam. Sensors, 14(5), 8150–8161.
Chou, Y. C., Masih Das, P., Monos, D. S., & Drndić, M. (2020). Lifetime and stability of silicon nitride nanopores and nanopore arrays for ionic measurements. ACS Nano, 14(6), 6715–6728.
Zaraska, L., Czopik, N., Bobruk, M., Sulka, G. D., Mech, J., & Jaskuła, M. (2013). Synthesis of nanoporous tin oxide layers by electrochemical anodization. Electrochimica Acta, 104, 549–557.
Apel, P. Y., Korchev, Y. E., Siwy, Z., Spohr, R., & Yoshida, M. (2001). Diode-like single-ion track membrane prepared by electro-stopping. Nuclear Instruments and Methods in Physics Research Section B: Beam Interactions with Materials and Atoms, 184(3), 337–346.
Siwy, Z., & Fuliński, A. (2002). Fabrication of a synthetic nanopore ion pump. Physical Review Letters, 89(19), 198103.
Masuda, H., & Fukuda, K. (1995). Ordered metal nanohole arrays made by a two-step replication of honeycomb structures of anodic alumina. Science, 268(5216), 1466–1468.
Lee, W., & Park, S. J. (2014). Porous anodic aluminum oxide: Anodization and templated synthesis of functional nanostructures. Chemical Reviews, 114(15), 7487–7556.
Pandey, B., Thapa, P. S., Higgins, D. A., & Ito, T. (2012). Formation of self-organized nanoporous anodic oxide from metallic gallium. Langmuir, 28(38), 13705–13711.
Sulka, G. D., Kapusta-Kołodziej, J., Brzózka, A., & Jaskuła, M. (2013). Anodic growth of TiO2 nanopore arrays at various temperatures. Electrochimica Acta, 104, 526–535.
Berger, S., Tsuchiya, H., & Schmuki, P. (2008). Transition from nanopores to nanotubes: Self-ordered anodic oxide structures on titanium–aluminides. Chemistry of Materials, 20(10), 3245–3247.
Nam, S. W., Rooks, M. J., Kim, K. B., & Rossnagel, S. M. (2009). Ionic field effect transistors with sub-10 nm multiple nanopores. Nano Letters, 9(5), 2044–2048.
Zeng, S., Wen, C., Solomon, P., Zhang, S. L., & Zhang, Z. (2019). Rectification of protein translocation in truncated pyramidal nanopores. Nature Nanotechnology, 14(11), 1056–1062.
Kim, M. J., McNally, B., Murata, K., & Meller, A. (2007). Characteristics of solid-state nanometre pores fabricated using a transmission electron microscope. Nanotechnology, 18(20), 205302.
Wang, R., Gilboa, T., Song, J., Huttner, D., Grinstaff, M. W., & Meller, A. (2018). Single-molecule discrimination of labeled DNAs and polypeptides using photoluminescent-free TiO2 nanopores. ACS Nano, 12(11), 11648–11656.
Mojtabavi, M., VahidMohammadi, A., Liang, W., Beidaghi, M., & Wanunu, M. (2019). Single-molecule sensing using nanopores in two-dimensional transition metal carbide (MXene) membranes. ACS Nano, 13(3), 3042–3053.
Chen, K., Zhu, J., Boskovic, F., & Keyser, U. F. (2020). Nanopore-based DNA hard drives for rewritable and secure data storage. Nano Letters, 20(5), 3754–3760.
Rodríguez-Manzo, J. A., Puster, M., Nicolaï, A., Meunier, V., & Drndic, M. (2015). DNA translocation in nanometer thick silicon nanopores. ACS Nano, 9(6), 6555–6564.
Traversi, F., Raillon, C., Benameur, S. M., Liu, K., Khlybov, S., Tosun, M., Krasnozhon, D., Kis, A., & Radenovic, A. (2013). Detecting the translocation of DNA through a nanopore using graphene nanoribbons. Nature Nanotechnology, 8(12), 939–945.
Schneider, G. F., Kowalczyk, S. W., Calado, V. E., Pandraud, G., Zandbergen, H. W., Vandersypen, L. M., & Dekker, C. (2010). DNA translocation through graphene nanopores. Nano Letters, 10(8), 3163–3167.
Geim, A. K. (2009). Graphene: Status and prospects. Science, 324(5934), 1530–1534.
Merchant, C. A., Healy, K., Wanunu, M., Ray, V., Peterman, N., Bartel, J., Fischbein, M. D., Venta, K., Luo, Z., Johnson, A. C., & Drndic, M. (2010). DNA translocation through graphene nanopores. Nano Letters, 10(8), 2915–2921.
Yuan, L., & Huang, L. (2015). Exciton dynamics and annihilation in WS2 2D semiconductors. Nanoscale, 7(16), 7402–7408.
Heerema, S. J., Vicarelli, L., Pud, S., Schouten, R. N., Zandbergen, H. W., & Dekker, C. (2018). Probing DNA translocations with inplane current signals in a graphene nanoribbon with a nanopore. ACS Nano, 12(3), 2623–2633.
Wanunu, M., Sutin, J., McNally, B., Chow, A., & Meller, A. (2008). DNA translocation governed by interactions with solid-state nanopores. Biophysical Journal, 95(10), 4716–4725.
Liang, L., Cui, P., Wang, Q., Wu, T., Ågren, H., & Tu, Y. (2013). Theoretical study on key factors in DNA sequencing with graphene nanopores. RSC Advances, 3(7), 2445–2453.
Luan, B., & Aksimentiev, A. (2010). Electric and electrophoretic inversion of the DNA charge in multivalent electrolytes. Soft Matter, 6(2), 243–246.
He, Y., Tsutsui, M., Fan, C., Taniguchi, M., & Kawai, T. (2011). Controlling DNA translocation through gate modulation of nanopore wall surface charges. ACS Nano, 5(7), 5509–5518.
Wanunu, M., Morrison, W., Rabin, Y., Grosberg, A. Y., & Meller, A. (2010). Electrostatic focusing of unlabelled DNA into nanoscale pores using a salt gradient. Nature Nanotechnology, 5(2), 160–165.
Tsutsui, M., He, Y., Furuhashi, M., Rahong, S., Taniguchi, M., & Kawai, T. (2012). Transverse electric field dragging of DNA in a nanochannel. Scientific Reports, 2(1), 1–7.
Kowalczyk, S. W., Wells, D. B., Aksimentiev, A., & Dekker, C. (2012). Slowing down DNA translocation through a nanopore in lithium chloride. Nano Letters, 12(2), 1038–1044.
Anderson, B. N., Muthukumar, M., & Meller, A. (2013). pH tuning of DNA translocation time through organically functionalized nanopores. ACS Nano, 7(2), 1408–1414.
Yeh, I. C., & Hummer, G. (2004). Nucleic acid transport through carbon nanotube membranes. Proceedings of the National Academy of Sciences, 101(33), 12177–12182.
Luan, B., Stolovitzky, G., & Martyna, G. (2012). Slowing and controlling the translocation of DNA in a solid-state nanopore. Nanoscale, 4(4), 1068–1077.
Akca, S., Foroughi, A., Frochtzwajg, D., & Postma, H. W. C. (2011). Competing interactions in DNA assembly on graphene. PLoS ONE, 6(4), e18442.
Yoshida, H., Goto, Y., Akahori, R., Tada, Y., Terada, S., Komura, M., & Iyoda, T. (2016). Slowing the translocation of single-stranded DNA by using nano-cylindrical passage self-assembled by amphiphilic block copolymers. Nanoscale, 8(43), 18270–18276.
Goto, Y., Haga, T., Yanagi, I., Yokoi, T., & Takeda, K. I. (2015). Deceleration of single-stranded DNA passing through a nanopore using a nanometre-sized bead structure. Scientific Reports, 5(1), 1–7.
Squires, A. H., Hersey, J. S., Grinstaff, M. W., & Meller, A. (2013). A nanopore–nanofiber mesh biosensor to control DNA translocation. Journal of the American Chemical Society, 135(44), 16304–16307.
Tang, Z., Liang, Z., Lu, B., Li, J., Hu, R., Zhao, Q., & Yu, D. (2015). Gel mesh as “brake” to slow down DNA translocation through solid-state nanopores. Nanoscale, 7(31), 13207–13214.
Plesa, C., Kowalczyk, S. W., Zinsmeester, R., Grosberg, A. Y., Rabin, Y., & Dekker, C. (2013). Fast translocation of proteins through solid state nanopores. Nano Letters, 13(2), 658–663.
Smoluchowski, M. V. (1917). Engineering biological structures of prescribed shape using self-assembled multicellular systems. Zeitschrift für Physikalische Chemie, 92, 129.
Pedone, D., Firnkes, M., & Rant, U. (2009). Data analysis of translocation events in nanopore experiments. Analytical Chemistry, 81(23), 9689–9694.
Larkin, J., Henley, R. Y., Muthukumar, M., Rosenstein, J. K., & Wanunu, M. (2014). High-bandwidth protein analysis using solid-state nanopores. Biophysical Journal, 106(3), 696–704.
Asandei, A., Schiopu, I., Chinappi, M., Seo, C. H., Park, Y., & Luchian, T. (2016). Electroosmotic trap against the electrophoretic force near a protein nanopore reveals peptide dynamics during capture and translocation. ACS Applied Materials and Interfaces, 8(20), 13166–13179.
Muthukumar, M. (2014). Communication: Charge, diffusion, and mobility of proteins through nanopores. The Journal of Chemical Physics, 141(8), 081104.
Kumar, A., Park, K. B., Kim, H. M., & Kim, K. B. (2013). Noise and its reduction in graphene based nanopore devices. Nanotechnology, 24(49), 495503.
Fragasso, A., Schmid, S., & Dekker, C. (2020). Comparing current noise in biological and solid-state nanopores. ACS Nano, 14(2), 1338–1349.
Waggoner, P. S., Kuan, A. T., Polonsky, S., Peng, H., & Rossnagel, S. M. (2011). Increasing the speed of solid-state nanopores. Journal of Vacuum Science & Technology B, Nanotechnology and Microelectronics: Materials, Processing, Measurement, and Phenomena, 29(3), 032206.
Uram, J. D., Ke, K., & Mayer, M. (2008). Noise and bandwidth of current recordings from submicrometer pores and nanopores. ACS Nano, 2(5), 857–872.
Smeets, R. M., Keyser, U. F., Wu, M. Y., Dekker, N. H., & Dekker, C. (2006). Nanobubbles in solid-state nanopores. Physical Review Letters, 97(8), 088101.
Marion, S., Macha, M., Davis, S. J., Chernev, A., & Radenovic, A. (2021). Wetting of nanopores probed with pressure. Physical Chemistry Chemical Physics, 23(8), 4975–4987.
Gravelle, S., Netz, R. R., & Bocquet, L. (2019). Adsorption kinetics in open nanopores as a source of low-frequency noise. Nano Letters, 19(10), 7265–7272.
Beamish, E., Kwok, H., Tabard-Cossa, V., & Godin, M. (2012). Precise control of the size and noise of solid-state nanopores using high electric fields. Nanotechnology, 23(40), 405301.
Steinbock, L. J., Bulushev, R. D., Krishnan, S., Raillon, C., & Radenovic, A. (2013). DNA translocation through low-noise glass nanopores. ACS Nano, 7(12), 11255–11262.
Balan, A., Machielse, B., Niedzwiecki, D., Lin, J., Ong, P., Engelke, R., Shepard, K. L., & Drndić, M. (2014). Improving signal-to-noise performance for DNA translocation in solid-state nanopores at MHz bandwidths. Nano Letters, 14(12), 7215–7220.
Fragasso, A., Pud, S. & Dekker, C. (2019). 1/f noise in solid-state nanopores is governed by access and surface regions. Nanotechnology, 30(39), 395202.
Saleh, O. A., & Sohn, L. L. (2001). Quantitative sensing of nanoscale colloids using a microchip coulter counter. Review of Scientific Instruments, 72(12), 4449–4451.
Varongchayakul, N., Song, J., Meller, A., & Grinstaff, M. W. (2018). Single-molecule protein sensing in a nanopore: A tutorial. Chemical Society Reviews, 47(23), 8512–8524.
Smeets, R. M., Keyser, U. F., Dekker, N. H., & Dekker, C. (2008). Noise in solid-state nanopores. Proceedings of the National Academy of Sciences, 105(2), 417–421.
Liu, L. F., Liu, C. C., & Alberts, B. M. (1980). Type II DNA topoisomerases: Enzymes that can unknot a topologically knotted DNA molecule via a reversible double-strand break. Cell, 19(3), 697–707.
Plesa, C., Verschueren, D., Pud, S., Van Der Torre, J., Ruitenberg, J. W., Witteveen, M. J., Jonsson, M. P., Grosberg, A. Y., Rabin, Y., & Dekker, C. (2016). Direct observation of DNA knots using a solid-state nanopore. Nature Nanotechnology, 11(12), 1093–1097.
Sharma, R. K., Agrawal, I., Dai, L., Doyle, P. S., & Garaj, S. (2019). Complex DNA knots detected with a nanopore sensor. Nature Communications, 10(1), 1–9.
Shim, J., Kim, Y., Humphreys, G. I., Nardulli, A. M., Kosari, F., Vasmatzis, G., Taylor, W. R., Ahlquist, D. A., Myong, S., & Bashir, R. (2015). Nanopore-based assay for detection of methylation in double-stranded DNA fragments. ACS Nano, 9(1), 290–300.
Charron, M., Briggs, K., King, S., Waugh, M., & Tabard-Cossa, V. (2019). Precise DNA concentration measurements with nanopores by controlled counting. Analytical Chemistry, 91(19), 12228–12237.
Bell, N. A., & Keyser, U. F. (2015). Specific protein detection using designed DNA carriers and nanopores. Journal of the American Chemical Society, 137(5), 2035–2041.
Han, A., Schürmann, G., Mondin, G., Bitterli, R.A., Hegelbach, N.G., de Rooij, N.F. & Staufer, U. (2006). Sensing protein molecules using nanofabricated pores. Applied Physics Letters, 88(9), 093901.
Talaga, D. S., & Li, J. (2009). Single-molecule protein unfolding in solid state nanopores. Journal of the American Chemical Society, 131(26), 9287–9297.
Firnkes, M., Pedone, D., Knezevic, J., Doblinger, M., & Rant, U. (2010). Electrically facilitated translocations of proteins through silicon nitride nanopores: Conjoint and competitive action of diffusion, electrophoresis, and electroosmosis. Nano Letters, 10(6), 2162–2167.
Saharia, J., Bandara, Y. N. D., Goyal, G., Lee, J. S., Karawdeniya, B. I., & Kim, M. J. (2019). Molecular-level profiling of human serum transferrin protein through assessment of nanopore-based electrical and chemical responsiveness. ACS Nano, 13(4), 4246–4254.
Yusko, E. C., Prangkio, P., Sept, D., Rollings, R. C., Li, J., & Mayer, M. (2012). Single-particle characterization of Aβ oligomers in solution. ACS Nano, 6(7), 5909–5919.
Kaur, H., Nandivada, S., Acharjee, M. C., McNabb, D. S., & Li, J. (2018). Estimating RNA polymerase protein binding sites on λ DNA using solid-state nanopores. ACS Sensors, 4(1), 100–109.
Raillon, C., Cousin, P., Traversi, F., Garcia-Cordero, E., Hernandez, N., & Radenovic, A. (2012). Nanopore detection of single molecule RNAP–DNA transcription complex. Nano Letters, 12(3), 1157–1164.
Kong, J., Bell, N. A., & Keyser, U. F. (2016). Quantifying nanomolar protein concentrations using designed DNA carriers and solid-state nanopores. Nano Letters, 16(6), 3557–3562.
Uram, J. D., Ke, K., Hunt, A. J., & Mayer, M. (2006). Submicrometer pore-based characterization and quantification of antibody–virus interactions. Small (Weinheim an der Bergstrasse, Germany), 2(8–9), 967–972.
Freedman, K. J., Bastian, A. R., Chaiken, I., & Kim, M. J. (2013). Solid-state nanopore detection of protein complexes: Applications in healthcare and protein kinetics. Small (Weinheim an der Bergstrasse, Germany), 9(5), 750–759.
Zhou, K., Li, L., Tan, Z., Zlotnick, A., & Jacobson, S. C. (2011). Characterization of hepatitis B virus capsids by resistive-pulse sensing. Journal of the American Chemical Society, 133(6), 1618–1621.
Tsutsui, M., Yamazaki, T., Tatematsu, K., Yokota, K., Esaki, Y., Kubo, Y., Deguchi, H., Arima, A., Kuroda, S. I., & Kawai, T. (2019). High-throughput single nanoparticle detection using a feed-through channel-integrated nanopore. Nanoscale, 11(43), 20475–20484.
Arima, A., Tsutsui, M., Harlisa, I. H., Yoshida, T., Tanaka, M., Yokota, K., Tonomura, W., Taniguchi, M., Okochi, M., Washio, T., & Kawai, T. (2018). Selective detections of single-viruses using solid-state nanopores. Scientific Reports, 8(1), 1–7.
Shi, X., Gao, R., Ying, Y. L., Si, W., Chen, Y. F., & Long, Y. T. (2016). A scattering nanopore for single nanoentity sensing. ACS Sensors, 1(9), 1086–1090.
Shi, X., Verschueren, D. V., & Dekker, C. (2018). Active delivery of single DNA molecules into a plasmonic nanopore for label-free optical sensing. Nano Letters, 18(12), 8003–8010.
Jadhav, V., Hoogerheide, D. P., Korlach, J., & Wanunu, M. (2018). Porous Zero-Mode Waveguides for Picogram-Level DNA Capture. Nano Letters, 19(2), 921–929.
Garoli, D., Yamazaki, H., Maccaferri, N., & Wanunu, M. (2019). Plasmonic nanopores for single-molecule detection and manipulation: Toward sequencing applications. Nano Letters, 19(11), 7553–7562.
Ohshiro, T., Tsutsui, M., Yokota, K., Furuhashi, M., Taniguchi, M., & Kawai, T. (2014). Detection of post-translational modifications in single peptides using electron tunnelling currents. Nature Nanotechnology, 9(10), 835–840.
Dimitrov, V., Mirsaidov, U., Wang, D., Sorsch, T., Mansfield, W., Miner, J., Klemens, F., Cirelli, R., Yemenicioglu, S., & Timp, G. (2010). Nanopores in solid-state membranes engineered for single molecule detection. Nanotechnology, 21(6), 065502.
Tabard-Cossa, V., Trivedi, D., Wiggin, M., Jetha, N. N., & Marziali, A. (2007). Noise analysis and reduction in solid-state nanopores. Nanotechnology, 18(30), 305505.
Lee, K., Park, K. B., Kim, H. J., Yu, J. S., Chae, H., Kim, H. M., & Kim, K. B. (2018). Recent progress in solid-state nanopores. Advanced Materials, 30(42), 1704680.
Smeets, R. M. M., Dekker, N. H., & Dekker, C. (2009). Low-frequency noise in solid-state nanopores. Nanotechnology, 20(9), 095501.
Sherman-Gold, R. (Ed.). (1993). The Axon guide for electrophysiology and biophysics: Laboratory techniques. Axon Instruments.
Heerema, S. J., Schneider, G. F., Rozemuller, M., Vicarelli, L., Zandbergen, H. W., & Dekker, C. (2015). 1/f noise in graphene nanopores. Nanotechnology, 26(7), 074001.
Chang, H., Kosari, F., Andreadakis, G., Alam, M. A., Vasmatzis, G., & Bashir, R. (2004). DNA-mediated fluctuations in ionic current through silicon oxide nanopore channels. Nano Letters, 4(8), 1551–1556.
Smeets, R. M., Keyser, U. F., Krapf, D., Wu, M. Y., Dekker, N. H., & Dekker, C. (2006). Salt dependence of ion transport and DNA translocation through solid-state nanopores. Nano Letters, 6(1), 89–95.
Yang, L., & Yamamoto, T. (2016). Quantification of virus particles using nanopore-based resistive-pulse sensing techniques. Frontiers in Microbiology, 7, 1500.
Funding
This research was supported by the National Research Foundation of Korea (NRF) grants funded by the Korean government (Ministry of Science and ICT) (NRF-2020R1A2C300488512 and NRF-2020R1A4A200272812).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Haq, M.R., Lee, B.J. & Lee, J. Solid-State Nanopore for Molecular Detection. Int. J. Precis. Eng. Manuf. 22, 2001–2026 (2021). https://doi.org/10.1007/s12541-021-00590-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12541-021-00590-2