Abstract
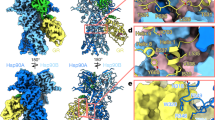
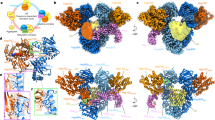
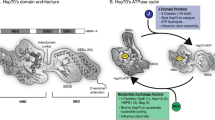
Extracellular heat shock protein 70 (HSP70) is recognized by receptors on the plasma membrane, such as Toll-like receptor 4 (TLR4), TLR2, CD14, and CD40. This leads to activation of nuclear factor-kappa B (NF-κB), release of pro-inflammatory cytokines, enhancement of the phagocytic activity of innate immune cells, and stimulation of antigen-specific responses. However, the specific characteristics of HSP70 binding are still unknown, and all HSP70 receptors have not yet been described. Putative models for HSP70 complexation to the receptor for advanced glycation endproducts (RAGEs), considering both ADP- and ATP-bound states of HSP70, were obtained through molecular docking and interaction energy calculations. This interaction was detected and visualized by a proximity fluorescence-based assay in A549 cells and further analyzed by normal mode analyses of the docking complexes. The interacting energy of the complexes showed that the most favored docking situation occurs between HSP70 ATP-bound and RAGE in its monomeric state. The fluorescence proximity assay presented a higher number of detected spots in the HSP70 ATP treatment, corroborating with the computational result. Normal-mode analyses showed no conformational deformability in the interacting interface of the complexes. Results were compared with previous findings in which oxidized HSP70 was shown to be responsible for the differential modulation of macrophage activation, which could result from a signaling pathway triggered by RAGE binding. Our data provide important insights into the characteristics of HSP70 binding and receptor interactions, as well as putative models with conserved residues on the interface area, which could be useful for future site-directed mutagenesis studies.







Similar content being viewed by others
References
Arnold-Schild D, Hanau D, Spehner D, Schmid C, Rammensee HG, de la Salle H, Schild H (1999) Cutting edge: receptor-mediated endocytosis of heat shock proteins by professional antigen-presenting cells. J Immunol 162(7):3757–3760
Asea A (2003) Chaperokine-induced signal transduction pathways. Exerc Immunol Rev 9:25–33
Asea A, Kabingu E, Stevenson MA, Calderwood SK (2000a) HSP70 peptidem bearing and peptide-negative preparations act as chaperokines. Cell Stress Chaperones 5(5):425–431
Asea A, Kraeft SK, Kurt-Jones EA, Stevenson MA, Chen LB, Finberg RW, Koo GC, Calderwood SK (2000b) HSP70 stimulates cytokine production through a CD14-dependant pathway, demonstrating its dual role as a chaperone and cytokine. Nat Med 6:435–442. doi:10.1038/74697
Asea A, Rehli M, Kabingu E, Boch JA, Bare O, Auron PE, Stevenson MA, Calderwood SK (2002) Novel signal transduction pathway utilized by extracellular HSP70: role of toll-like receptor (TLR) 2 and TLR4. J Biol Chem 277(17):15028–15034. doi:10.1074/jbc.M200497200
Bahar I, Lezon TR, Bakan A, Shrivastava IH (2010) Normal mode analysis of biomolecular structures: functional mechanisms of membrane proteins. Chem Rev 110(3):1463–1497. doi:10.1021/cr900095e
Bakan A, Meireles LM, Bahar I (2011) ProDy: protein dynamics inferred from theory and experiments. Bioinformatics 27(11):1575–1577. doi:10.1093/bioinformatics/btr168
Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA (2001) Electrostatics of nanosystems: application to microtubules and the ribosome. Proc Natl Acad Sci U S A 98(18):10037–10041. doi:10.1073/pnas.181342398
Becker T (2002) CD40, an extracellular receptor for binding and uptake of Hsp70-peptide complexes. J Cell Biol 158(7):1277–1285. doi:10.1083/jcb.200208083
Bertelsen EB, Chang L, Gestwicki JE, Zuiderweg ER (2009) Solution conformation of wild-type E. coli Hsp70 (DnaK) chaperone complexed with ADP and substrate. Proc Natl Acad Sci U S A 106(21):8471–8476. doi:10.1073/pnas.0903503106
Biovia, D. S. (2015). Discovery studio modeling environment (version 4.5). San Diego
Bucciarelli LG, Wendt T, Rong L, Lalla E, Hofmann MA, Goova MT, Taguchi A, Yan SF, Yan SD, Stern DM, Schmidt AM (2002) RAGE is a multiligand receptor of the immunoglobulin superfamily: implications for homeostasis and chronic disease. Cell Mol Life Sci 59(7):1117–1128
Chen YC (2015) Beware of docking! Trends Pharmacol Sci 36(2):78–95. doi:10.1016/j.tips.2014.12.001
Comeau SR, Gatchell DW, Vajda S, Camacho CJ (2004) ClusPro: a fully automated algorithm for protein-protein docking. Nucleic Acids Res 32(Web Server issue):W96–W99. doi:10.1093/nar/gkh354
Delneste Y, Magistrelli G, Gauchat J, Haeuw J, Aubry J, Nakamura K, Kawakami-Honda N, Goetsch L, Sawamura T, Bonnefoy J, Jeannin P (2002) Involvement of LOX-1 in dendritic cell-mediated antigen cross-presentation. Immunity 17(3):353–362
Dolinsky TJ, Nielsen JE, McCammon JA, Baker NA (2004) PDB2PQR: an automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucleic Acids Res 32(Web Server issue):W665–W667. doi:10.1093/nar/gkh381
Frappier V, Najmanovich RJ (2014) A coarse-grained elastic network atom contact model and its use in the simulation of protein dynamics and the prediction of the effect of mutations. PLoS Comput Biol 10(4):e1003569. doi:10.1371/journal.pcbi.1003569
Frappier V, Chartier M, Najmanovich RJ (2015) ENCoM server: exploring protein conformational space and the effect of mutations on protein function and stability. Nucleic Acids Res 43(W1):W395–W400. doi:10.1093/nar/gkv343
Fritz G (2011) RAGE: a single receptor fits multiple ligands. Trends Biochem Sci 36(12):625–632. doi:10.1016/j.tibs.2011.08.008
Fujiya A, Nagasaki H, Seino Y, Okawa T, Kato J, Fukami A, Himeno T, Uenishi E, Tsunekawa S, Kamiya H, Nakamura J, Oiso Y, Hamada Y (2014) The role of S100B in the interaction between adipocytes and macrophages. Obesity (Silver Spring) 22(2):371–379. doi:10.1002/oby.20532
Galloway E, Shin T, Huber N, Eismann T, Kuboki S, Schuster R, Blanchard J, Wong HR, Lentsch AB (2008) Activation of hepatocytes by extracellular heat shock protein 72. Am J Physiol Cell Physiol 295(2):C514–C520. doi:10.1152/ajpcell.00032.2008
Gelain DP, de Bittencourt Pasquali MA, Comin MC, Grunwald MS, Ritter C, Tomasi CD, Alves SC, Quevedo J, Dal-Pizzol F, Moreira JC (2011) Serum heat shock protein 70 levels, oxidant status, and mortality in sepsis. Shock 35(5):466–470. doi:10.1097/SHK.0b013e31820fe704
Gross C, Schmidt-Wolf IG, Nagaraj S, Gastpar R, Ellwart J, Kunz-Schughart LA, Multhoff G (2003) Heat shock protein 70-reactivity is associated with increased cell surface density of CD94/CD56 on primary natural killer cells. Cell Stress Chaperones 8(4):348–360
Grunwald MS, Pires AS, Zanotto-Filho A, Gasparotto J, Gelain DP, Demartini DR, Schöler CM, de Bittencourt PI Jr, Moreira JC (2014) The oxidation of HSP70 is associated with functional impairment and lack of stimulatory capacity. Cell Stress Chaperones 19(6):913–925. doi:10.1007/s12192-014-0516-5
Kindas-Mugge I, Hammerle AH, Frohlich I, Oismuller C, Micksche M, Trautinger F (1993) Granulocytes of critically ill patients spontaneously express the 72 kD heat shock protein. Circ Shock 39(4):247–252
Kityk R, Kopp J, Sinning I, Mayer MP (2012) Structure and dynamics of the ATP-bound open conformation of Hsp70 chaperones. Mol Cell 48(6):863–874. doi:10.1016/j.molcel.2012.09.023
Koch M, Chitayat S, Dattilo BM, Schiefner A, Diez J, Chazin WJ, Fritz G (2010) Structural basis for ligand recognition and activation of RAGE. Structure 18(10):1342–1352. doi:10.1016/j.str.2010.05.017
Kozakov D, Brenke R, Comeau SR, Vajda S (2006) PIPER: an FFT-based protein docking program with pairwise potentials. Proteins 65(2):392–406. doi:10.1002/prot.21117
Lopez-Blanco JR, Aliaga JI, Quintana-Orti ES, Chacon P (2014) iMODS: internal coordinates normal mode analysis server. Nucleic Acids Res 42(Web Server issue):W271–W276. doi:10.1093/nar/gku339
Ma W, Rai V, Hudson BI, Song F, Schmidt AM, Barile GR (2012) RAGE binds C1q and enhances C1q-mediated phagocytosis. Cell Immunol 274(1–2):72–82. doi:10.1016/j.cellimm.2012.02.001
Macindoe G, Mavridis L, Venkatraman V, Devignes MD, Ritchie DW (2010) HexServer: an FFT-based protein docking server powered by graphics processors. Nucleic Acids Res 38(Web Server issue):W445–W449. doi:10.1093/nar/gkq311
Mahajan S, Sanejouand YH (2015) On the relationship between low-frequency normal modes and the large-scale conformational changes of proteins. Arch Biochem Biophys 567:59–65. doi:10.1016/j.abb.2014.12.020
Musiani F, Ciurli S (2015) Evolution of macromolecular docking techniques: the case study of nickel and iron metabolism in pathogenic bacteria. Molecules 20(8):14265–14292. doi:10.3390/molecules200814265
Nakano N, Fukuhara-Takaki K, Jono T, Nakajou K, Eto N, Horiuchi S, Takeya M, Nagai R (2006) Association of advanced glycation end products with A549 cells, a human pulmonary epithelial cell line, is mediated by a receptor distinct from the scavenger receptor family and RAGE. J Biochem 139(5):821–829. doi:10.1093/jb/mvj092
Park H, Lee H, Seok C (2015) High-resolution protein-protein docking by global optimization: recent advances and future challenges. Curr Opin Struct Biol 35:24–31. doi:10.1016/j.sbi.2015.08.001
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF chimera—a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612. doi:10.1002/jcc.20084
Ritossa F (1962) A new puffing pattern induced by temperature shock and DNP in drosophila. Experientia 18(12):571–573. doi:10.1007/BF02172188
Ruan BH, Li X, Winkler AR, Cunningham KM, Kuai J, Greco RM, Nocka KH, Fitz LJ, Wright JF, Pittman DD, Tan XY, Paulsen JE, Lin LL, Winkler DG (2010) Complement C3a, CpG oligos, and DNA/C3a complex stimulate IFN-alpha pzcation end product-dependent manner. J Immunol 185(7):4213–4222. doi:10.4049/jimmunol.1000863
Schmidt AM, Hofmann M, Taguchi A, Yan SD, Stern DM (2000) RAGE: a multiligand receptor contributing to the cellular response in diabetic vasculopathy and inflammation. Semin Thromb Hemost 26(5):485–493. doi:10.1055/s-2000-13204
Schneidman-Duhovny D, Inbar Y, Nussinov R, Wolfson HJ (2005) PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res 33(Web Server issue):W363–W367. doi:10.1093/nar/gki481
Schymkowitz J, Borg J, Stricher F, Nys R, Rousseau F, Serrano L (2005) The FoldX web server: an online force field. Nucleic Acids Res 33(Web Server issue):W382–W388. doi:10.1093/nar/gki387
Sousa R (2012) A dancer caught midstep: the structure of ATP-bound Hsp70. Mol Cell 48(6):821–823. doi:10.1016/j.molcel.2012.12.008
Suhre K, Sanejouand YH (2004) ElNemo: a normal mode web server for protein movement analysis and the generation of templates for molecular replacement. Nucleic Acids Res 32(Web Server issue):W610–W614. doi:10.1093/nar/gkh368
Tang D, Loze MT, Zeh HJ, Kang R (2010) The redox protein HMGB1 regulates cell death and survival in cancer treatment. Autophagy 6(8):1181–1183. doi:10.4161/auto.6.8.13367
Tovchigrechko A, Vakser IA (2006) GRAMM-X public web server for protein-protein docking. Nucleic Acids Res 34(Web Server issue):W310–W314. doi:10.1093/nar/gkl206
Trautinger F, Kindas-Mugge I, Knobler RM, Honigsmann H (1996a) Stress proteins in the cellular response to ultraviolet radiation. J Photochem Photobiol B 35(3):141–148
Trautinger F, Knobler RM, Honigsmann H, Mayr W, Kindas-Mugge I (1996b) Increased expression of the 72-kDa heat shock protein and reduced sunburn cell formation in human skin after local hyperthermia. J Invest Dermatol 107(3):442–443
van Eden W (2015) Diet and the anti-inflammatory effect of heat shock proteins. Endocr Metab Immune Disord Drug Targets 15(1):31–36
Varecha M, Potesilova M, Matula P, Kozubek M (2012) Endonuclease G interacts with histone H2B and DNA topoisomerase II alpha during apoptosis. Mol Cell Biochem 363(1–2):301–307. doi:10.1007/s11010-011-1182-x
Vega VL, Rodriguez-Silva M, Frey T, Gehrmann M, Diaz JC, Steinem C, Multhoff G, Arispe N, De Maio A (2008) Hsp70 translocates into the plasma membrane after stress and is released into the extracellular environment in a membrane-associated form that activates macrophages. J Immunol 180(6):4299–4307
Yatime L, Andersen GR (2013) Structural insights into the oligomerization mode of the human receptor for advanced glycation end-products. FEBS J 280(24):6556–6568. doi:10.1111/febs.12556
Acknowledgements
This work was supported by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Fundação de Amparo à Pesquisa do Estado do Rio Grande do Sul (FAPERGS), CAPES, and Propesq-UFRGS. The authors thank the Electron Microscopy Center of the Federal University of Rio Grande do Sul for their assistance with the microscopy analyses. The authors also thank Mr. Henrique Biehl for his technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Author contribution
JCFM and HV conceived and coordinated the study. DPG helped conceive the idea. MSG designed, performed, and analyzed all the experiments, as well as wrote the manuscript and prepared all the figures. RLB provided technical assistance for normal mode analyses and contributed with the writing of the manuscript. CSS and LH provided technical assistance and contributed to the preparation of Fig. 2. All authors reviewed the results and approved the final version of the manuscript.
Electronic supplementary materials
ESM 1
(DOCX 4606 kb)
Rights and permissions
About this article
Cite this article
Grunwald, M.S., Ligabue-Braun, R., Souza, C.S. et al. Putative model for heat shock protein 70 complexation with receptor of advanced glycation end products through fluorescence proximity assays and normal mode analyses. Cell Stress and Chaperones 22, 99–111 (2017). https://doi.org/10.1007/s12192-016-0746-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12192-016-0746-9