Abstract
Aplastic anaemia (AA) is frequently associated with other disorders of clonal haemopoiesis such as paroxysmal nocturnal haemoglobinuria (PNH), myelodysplastic syndrome (MDS) and T-large granular lymphocytosis. Certain clones may escape the immune attack within the bone marrow environment and proliferate and attain a survival advantage over normal haemopoietic stem cells, such as trisomy 8, loss of heterozygosity of short arm of chromosome 6 and del13q clones. Recently acquired somatic mutations (SM), excluding PNH clones, have been reported in around 20–25 % of patients with AA, which predispose to a higher risk of later malignant transformation to MDS/acute myeloid leukaemia. Furthermore, certain SM, such as ASXL1 and DNMT3A are associated with poor survival following immunosuppressive therapy, whereas PIGA, BCOR/BCORL1 predict for good response and survival. Further detailed and serial analysis of the immune signature in AA is needed to understand the pathogenetic basis for the presence of clones with SM in a significant proportion of patients.
Similar content being viewed by others
Introduction
Acquired aplastic anaemia (AA) is largely an immune mediated disorder that may present concurrently with clonal haemopoietic stem cell (HSC) disorders, most commonly paroxysmal nocturnal haemoglobinuria (PNH). It may later evolve to myelodysplastic syndrome (MDS) in up to 15–20 % of patients [1–3]. Furthermore, there is overlap between AA and MDS in the form of the entity hypocellular MDS which is often difficult to distinguish from AA on morphological criteria, especially when AA is of the non-severe sub-type [4]. Because AA may be associated with an abnormal cytogenetic clone in up to 12 % of patients, the finding of an abnormal cytogenetic clone does not always help in differentiating AA from hypocellular MDS, although the finding of monosomy 7 usually indicates MDS instead of AA [5]. There are specific acquired somatic mutations (SMs) that characterise these overlapping bone marrow failure (BMF) disorders, in addition to mutations of PIGA that occur in PNH. Acquired STAT3 mutations occur in 40 % of patients with T-large granular lymphocytosis (T-LGL) but can also be detected in 7 % of AA and 3 % of MDS patients with unsuspected T-LGL, that is, with subclinical T-LGL clones. [6] Lastly, SMs that typify MDS and acute myeloid leukaemia (AML) have recently been reported in AA, and which are the main focus of this mini review [7, 8].
Expansion of clones that escape immune attack in aplastic anaemia
Underlying this clonal haemopoiesis is an important interaction with the immune response that occurs in AA resulting in expansion and a proliferative advantage of certain clones that can evade the immune attack, and contribute to haemopoiesis [1–3]. This is best exemplified by PIGA mutated HSC. Data supporting an intrinsic survival advantage of PNH HSCs are lacking. Instead, the expansion of PNH clones in AA is more likely due to an extrinsic factor, for example, immune attack, whereby PNH HSCs are thought to escape the immune attack that occurs in AA because they lack expression of the glycosyl phosphatidylinositol (GPI) anchor (GPI-AP-), and that the target for the immune attack of normal HSCs is the GPI-anchor itself [9, 10]. Other theories include targeting of non-PNH HSCs by LGL clones [11]. Alternatively, NK cell mediated cytotoxicity may play a role; immunoglobulin-like receptors (KIR) may be differentially expressed in PNH compared to normal, resulting in cytotoxicity of normal HSCs [12]. The presence of GPI-specific, CD1d restricted T cells in PNH supports the hypothesis that PNH HSCs preferentially expands due to immune escape [13].
In both AA and MDS the presence of trisomy 8 (+8) is associated with a high response rate to immunosuppressive therapy (IST). Bone marrow (BM) haemopoietic progenitor cells (HPC) from patients with MDS and +8 show increased expression of WT1 antigen. This induces a specific T cell response to WT1 peptides, leading to suppression of non +8 HPC through a bystander effect by activated CD8 T-cells. In contrast, +8 HPCs survive this immune attack due to increased expression of anti-apoptotic proteins survivin, cyclin D1, and increased proliferation due to increased expression of c-myc, resulting in a proliferative advantage for the +8 HPC [14]. More recent work from Hosokawa and colleagues indicates that the presence of GPI-AP- HPC among patients with +8 may be an important factor associated with the high response to IST. In a series of 53 patients with AA (n = 22) and low risk MDS (n = 31), 26 % had a PNH clones, as defined by ≥0.005 % for red cells and ≥0.003 % granulocytes. Of 26 patients who received IST, the response rate was 88 % for those with a PNH clones compared to only 41 % without a PNH clone. Furthermore, the response rate to IST in patients with +8 was lower (56 %) than for patients with normal cytogenetics (81 %). Thus, patients with +8 and a PNH clones were more likely to respond to IST compared to those patients lacking a PNH clone [15].
Using SNPA karyotyping in more than 300 AA patients, Katagiri and colleagues showed that the most common genetic lesion was copy number neutral loss of heterozygosity involving the short arm of chromosome 6 (LOH6p) in 13 % of patients [16]. This is an acquired genetic event since it was not detected in CD3+ T-cells, and it involved multiple haemopoietic cell lineages as well as early (CD34+) BM HPCs. LOH commonly affected the HLA locus with loss of expression of HLA-A antigen. In AA, the targets of cytotoxic T-lymphocytes (CTLs) are HPC that present autoantigen expressed by class I HLA molecules, particularly certain HLA-A*02:01, A*02:06, A*03:01, and B*40:02. If there is LOH6p, then the HPC have lost the target for immune attack by CTLs and hence escape the immune attack, resulting in a growth advantage and clonal expansion over unaffected HPCs.
Del13q is closely associated with PIGA mutant clones. In a study from Japan, patients with isolated del13q all responded to IST and none progressed to MDS/AML. All had PNH clones, and using cytoFISH, the del13q cells were only present in the non-mutant GPI-AP+ cells. Following IST, expansion of del13q clone occurred more frequently than a decrease in the clone size. So in this situation, both the del13q HSCs and the PIGA mutant HSCs underwent preferential expansion and contributed to haematological recovery [17].
So, could a similar process of immune escape explain the existence of some of the other acquired SMs that have been recently described in some patients with AA?
Age-related clonal haemopoiesis: clonal haemopoiesis of indeterminate prognosis (CHIP)
Before considering further the significance of acquired SMs in AA, the recent observation that acquired SMs occur with increasing age in healthy individuals is highly. Several large population based cohort studies have demonstrated that between the ages of 70–79 years, SMs are detected in 10 % of people, and rising further with increasing age thereafter [18–20]. The most frequent genes mutated were DNMT3A, TET2, ASXL1 and less frequently TP53, JAK2, SF3B1, that is, genes that are frequently mutated in MDS/AML. The presence of a SM was associated with an increased risk of haematological malignancy, and increased mortality due to increase in cardiac, cerebral events and diabetes. Most frequently, there was only one SM per patient and the median allele burden (MAB) was low at 9 %, although for those individuals who later developed a haematological malignancy, the MAB was higher at around 20 %. The incidence of SMs, especially DNMT3A R882, rises with increased sensitivity of method used for detection, raising the important question of what clone size is definitely clinically relevant [21]. This entity of age-related clonal haemopoiesis has recently been termed ‘clonal haemopoiesis of indeterminate potential’ (‘CHIP’) [22].
Somatic mutations in AA
In recent years, a number of studies have reported the presence of acquired SMs in AA, often associated with low level clones (Table 1). Lane et al. screened for 219 genes in 39 patients, and found SMs in 9 (23 %), comprising ASXL1 (n = 2), DNMT3A (n = 1) and BCOR (n = 1). The MAB was <10 % in 7 patients [23]. Heuser et al. found 3 mutations in 2 of 38 patients (SLIT1, and SETBP1 with ASXL1) using a smaller panel of 42 genes and excluding SM with a MAB of <15 % [24]. However, the patient with SETBP1 and ASXL1 was tested at time of progression to MDS. In a small cohort of predominantly paediatric patients, SMs were detected in 72 %, most frequently involved in immune escape (PIGA, LOH6p) and signal transduction (STAT5B, CAMK2G), and MDS-associated SM were found in only 9 % of patients [25].
From our King’s College Hospital database of 345 patients with idiopathic BMF, we were the first to describe the molecular profile in a large cohort of 150 AA patients with no morphological evidence of MDS and who had stored BM and skin/buccal mucosa samples. We excluded all patients with a known constitutional BMF disorder or a family history of BMF or cancer [26]. The first cohort of 57 patients was screened using a custom panel of 832 gene exons and for the second cohort of 93 patients, more targeted sequencing was performed for genes identified in the first cohort. We identified a subgroup (19 %) with pathogenetically relevant SMs in a relatively small number of genes (ASXL1 in 12 patients, DNMT3A in 8, BCOR in 6 and one each for SRSF2, U2AF1, TET2, MPL, IKZF1 and ERBB2. The MAB was 20 %, and for 41 % of SMs the MAB was <19 % clone. SMs, when examined together, predicted for risk of later evolution to MDS; the risk was 38 % compared to 6 % in the absence of a SM (p < 0.001), and if the disease duration of the AA was >6 months, the risk of MDS was even more significant at 40 % compared to 4 % without a SM (p < 0.0002) (see Table 2). ASXL1 and DNMT3A mutations were associated with evolution to monosomy 7 in 4 AA patients. We also showed that presence of a SM was associated with shorter telomere length compared to patients who lacked a SM. Patients in the first cohort of 57 patients were also screened for PIGA mutations. 23 of the 57 had a PNH clone by flow cytometry, and of these, 17 had a PNH clone size >10 %. In 7/17 a single PIGA mutation was found, double PIGA mutations were present in 6, and a PIGA mutation with another SM in 4 patients (BCOR in 2, and one each with ASXL1 and IKZF2). A PIGA mutation was not detected in any of the 6 patients where the PNH clone size was <10 %, indicating that flow cytometry is far more sensitive than PIGA sequencing at detecting small PNH clones [26].
Subsequently, a combined Japanese and USA study reported targeted sequencing of 106 genes in 439 AA patients, with whole exome sequencing in 52, and serial sampling in 82 patients [27]. The most frequently mutated genes were similar to the King’s College Hospital study with the exception of BCORL1 (which was not in the King’s panel) and PIGA (in the Kings study, PIGA was only screened for in the first cohort of 57 patients). SMs were found in 36 % of AA patients, and in 24 % of patients if PIGA SMs were excluded. SMs increased with increasing age (except for BCOR/BCORL1 and PIGA). Most of the SMs were present at a lower MAB at diagnosis compared to 6 months after IST. The NIH cohort identified so-called ‘favourable’ SMs, BCOR and PIGA, which were associated with better response to IST and better overall survival, in contrast to ‘unfavourable’ SMs (DNMT3A, ASXL1, TP53, RUNX1, JAK2, JAK3, or CSMD1, although predominantly DNMT3A and ASXL1) as a group were associated with poorer response to IST and worse survival (see Fig. 1) and progression to MDS/AML. They also showed that monosomy 7 detected at 6 months was associated with poor survival and progression to MDS. The impact of unfavourable SMs was even more significant in patients aged <60 years, raising the possibility that in future, factors such as ‘unfavourable’ SMs may help identify patients who might be considered for alternative treatment strategy such as early allogeneic haemopoietic stem cell transplantation (HSCT). Using whole exome sequencing to examine clonal architecture, highly variable patterns were seen. In most patients, clonal haemopoiesis originated from a minor clone present at diagnosis. In some cases, clones were stable over many years. When they examined the pattern of change in clone size for individual SMs, the ‘unfavourable’ clones (ASXL1 and DNMT3A) more often continued to enlarge, but not in all cases and some even disappeared over time. In contrast the ‘favourable’ SMs (BCOR, PIGA) were more likely to remain stable or decrease in size. The NIH group also highlighted in a separate study the contribution of increased telomere loss to the subsequent acquisition of SMs and emergence of monosomy 7 in a cohort of 13 SAA patients treated with IST and sampled serially post IST [28].
Impact of somatic mutations on survival after immunosuppressive therapy (IST). Aplastic anaemia patients with so-called ‘unfavourable’ SMs (ASXL1, DNMT3A) show significantly worse response to and overall survival after IST compared to patients with ‘favourable’ SMs (PIGA, BCOR/BCORL1). The impact of ‘unfavourable ‘SMs was more marked in patients aged <60 years. Modified from Yoshizato et al. [27], reproduced with permission
Features of SMs in AA compared to MDS, clonal cytopenia of uncertain significance (CCUS) and normal individuals
Acquired SMs occur more frequently in AA than in older age healthy individuals, but less frequently than in MDS (see Table 3). Many of the SMs seen in AA are similar in type to MDS, clonal cytopenia of uncertain significance (CCUS) which comprises 35 % of patients with idiopathic cytopenia of uncertain significance (ICUS) and who have a SM, and healthy people, except that there are more cases of BCOR and PIGA and a lower frequency of splicosome mutations, RUNX1 and TET2 in AA. In AA, one can identify good and poor prognosis SMs as discussed above. The MAB is in most AA patients low and lower than in MDS or CCUS [29]. Lastly, the number of SMs per patient in AA is less than in MDS and more similar to the number in older aged healthy individuals.
Questions and future perspectives
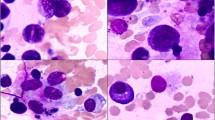
AA patients who are treated with IST, may recover normal HSC, or they may acquire sequentially SMs and dysplastic changes resulting in clonal expansion to MDS/AML. Complicating this picture is the so-called ‘overlap AA/hypocellular MDS’ syndrome of patients where it is not possible on morphological grounds to clearly distinguish AA from hypocellular MDS (see Fig. 2) [30]. Current data reviewed above now demonstrate that SMs (excluding PIGA) occur in around 20–24 % of patients who clearly have AA and with no morphological evidence of MDS [26, 27]. These patients have with an increased risk (40 % if the duration of AA is >6 months) of later developing MDS/AML [26]. Furthermore, patients with an ‘unfavourable’ SM (ASXL1, DNMT3A) have a lower response rate to IST and worse survival; in contrast, ‘favourable’ SMs (BCOR, PIGA) are associated with good response and survival after IST [27]. However, these exciting new data have raised several as yet unanswered and important questions (Table 3).
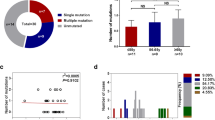
Mutational profile of aplastic anaemia and its evolution following immunosuppressive therapy, and frequency of 4 commonly mutated genes in aplastic anaemia. Reproduced with permission from Mufti et al. [31]
-
1.
What is the clonal architecture, mutational hierarchy and chronology of genetic events in AA? To help answer this, the European Blood and Marrow transplant (EBMT) Severe Aplastic Anaemia Working Party has established a prospective randomised trial for newly diagnosed SAA patients of standard IST with horse antithymocyte globulin (ATG) and ciclosporin with or without eltrombopag (‘RACE’ trial) [31]. The main outcome measure is complete response at 6 months. Patients will be closely monitored for morphological and cytogenetic evidence of later transformation to MDS/AML in view of the reported risk of MDS/AML (with high frequency of monosomy 7) in 18 % of patients when eltrombopag is used as a single agent in the treatment of refractory SAA [32–34]. As part of this clinical trial, blood and BM samples will be taken pre and at set time points after ATG for a key research study, the aims of which are to examine the evolving clonal architecture and mutational hierarchy at the genomic level with serial sampling, and to correlate results with the immune signature that predicts response and later risk of evolution to MDS/AML.
-
2.
Are SMs in AA related to ageing? Age-related clones occurring at low level may represent a predilection/founder stage that requires later cooperating mutations for clonal expansion and disease. In AA, the detection of both small and relatively large disease clone populations in AA likely indicates different stages of clonal evolution rather than normal ageing [26].
-
3.
What is the significance of low-level mutant clones? They may represent sub-populations of disease clones present at an early stage, but are they relevant to disease progression? Might they fluctuate over time in a similar manner to PNH clones or abnormal cytogenetic clones? Do they arise by selective protection from immune destruction analogous to PNH [9], LOH6p [16], +8 [14, 15] and del13q [17] clones? Other clones may be kept under control by process of immune surveillance leading to their elimination.
-
4.
Do SMs in AA indicate a diagnosis of hypocellular MDS rather than AA? It is evident that new diagnostic criteria are needed to help differentiate these two disorders, and molecular testing should now be incorporated into new diagnostic criteria.
-
5.
‘Unfavourable’ SMs, DNMT3A and ASXL1, help to identify patients with poor response to IST and worse survival, especially patients aged <60 years [27] and SMs in general predict for a high risk of later MDS/AML [25]. Should we now incorporate these results into other known poor prognostic factors such as short TL [35], low absolute lymphocyte and reticulocyte counts at the time of diagnosis? [36]. Should such patients now be considered for an alternative treatment strategy, specifically allogeneic HSCT? This is now timely in the light of recent improved outcomes of fludarabine-based HSCT, especially those using alemtuzumab, which is associated with a very low risk of GVHD [37–40].
-
6
Lastly, what is the correlation of the immune response with the emergence and clonal architecture of SMs in AA, and is there a specific immune signature that predicts for malignant transformation? Following an initial insult to the BM HSC, likely viral, an inflammatory immune response is initiated characterised by an increase in CD4+ T-helpers, Th1 (clonal expansion), and Th17 cells, and a reduction in Tregs which are also dysfunctional in terms of suppressing auto-reactive CD8+ T-cells, resulting in oligoclonal expansion of CD8+ CTLs [3]. The depleted stem cell pool and the increased proliferative pressure contribute to increased telomere attrition. A shift in the immune response from a state of immune surveillance through selection to escape with increased Tregs, with concurrent increasing genomic instability, results in emergence of abnormal clones. Repeated courses of IST [41], eltrombopag [33] and prolonged and high doses of G-CSF [42] are associated with increased risk of MDS/AML, especially with emergence of monosomy 7. Poor prognosis clones include monosomy 7, ASXL1 and DNMT3A, with increased risk of malignant transformation to MDS/AML, whereas good prognosis clones such as PNH clones, +8, del(13)q and BCOR/BCORL1 are not associated with increased malignant transformation (see Fig. 3). Finally, further understanding of the immune signature that occurs with the acquisition of SMs will be key to determining the pathogenetic mechanism for clonal transformation in SAA, using, for example, multi-dimensional mass cytometry, as recently reported by our group [43].
Fig. 3 Clonal haemopoiesis in acquired aplastic anaemia. An initial insult to the BM HSCs triggers an inflammatory immune response with an increase in CD4+ T-helpers and reduction in Tregs, which are dysfunctional in terms of suppressing auto-reactive CD8+ T-cells, resulting in oligoclonal expansion of CD8 + CTLs. The depleted stem cell pool and increased proliferative pressure contribute to increased telomere attrition. A shift in the immune response from immune surveillance through selection to escape with increased Tregs, and concurrent increasing genomic instability, result in emergence of abnormal clones. Repeated courses of IST, eltrombopag and prolonged and high doses of G-CSF are associated with increased risk of MDS/AML. Poor prognosis clones include monosomy 7, ASXL1 and DNMT3A, with increased risk of malignant transformation to MDS/AML, whereas good prognosis clones such as PNH clones, +8, del(13)q and BCOR/BCORL1 are not associated with increased malignant transformation. Tregs regulatory T-cells; CTLs cytotoxic T-lymphocytes
References
Young NS, Calado RT, Scheinberg P. Current concepts in the pathophysiology and treatment of aplastic anaemia. Blood. 2006;108:2509–19.
Young NS, Bacigalupo A, Marsh JCW. Aplastic anemia: pathophysiology and treatment. Biol Blood Marrow Transplant. 2010;16(1):S119–25.
Kordasti S, et al. Functional characterization of CD4+ T-cells in Aplastic Anemia. Blood. 2012;119:2033–43.
Bennett JM, Orazi A. Diagnostic criteria to distinguish hypocellular acute myeloid leukemia from hypocellular myelodysplastic syndromes and aplastic anemia: recommendations for a standardized approach. Haematologica. 2009;94:264–8.
Gupta V, et al. Clinical relevance of cytogenetics abnormalities in adult patients with acquired aplastic anaemia. Br J Haematol. 2006;134:95–9.
Jerez A, et al. STAT3 mutations indicate the presence of subclinical T-cell clones in a subset of aplastic anemia and myelodysplastic syndrome patients. Blood. 2013;122:2453–9.
Haferlach T, et al. Landscape of genetic lesions in 944 patients with myelodysplastic syndromes. Leukemia. 2014;28:241–7.
Papaemmanuil E, et al. Chronic Myeloid Disorders Working Group of the International Cancer Genome; Consortium. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood. 2013;122:3616–27.
Maciejewsk JP, et al. Impaired hematopoiesis in paroxysmal nocturnal haemoglobinuria/aplasticanemia is not associated with a selective proliferative defect in the glycosylphosphatidyl-inositol-anchored protein-deficient clone. Blood. 1997;89:1173–81.
Murakami Y, et al. Inefficient response of T lymphocytes to glycosylphosphatidyl-inositol-anchor-negative cells: implications for paroxysmal nocturnal hemoglobinuria. Blood. 2002;100:4116–22.
Risitano AM, et al. Large granular lymphocyte (LGL)- like clonal expansions in paroxysmal nocturnal hemoglobinuria (PNH) patients. Leukemia. 2005;19:217–22.
Va Bijnen ST, et al. T cells expressing the activating NK-cell receptors KIR2DS4, NKG2C and NKG2D are elevated in paroxysmal nocturnal hemoglobinuria and cytotoxic toward hematopoietic progenitor cell lines. Exp Hematol. 2011;39:751–62 (e1–3).
Gargiulo L, et al. Glycosylphosphatidylinositol-specific, CD1d-restricted T cells in paroxysmal nocturnal hemoglobinuria. Blood. 2013;121:2753–61.
Sloand EM, Pfannes L, Chen G, et al. CD34 cells from patients with trisomy 8 myelodysplastic syndrome (MDS) express early apoptotic markers but avoid programmed cell death by up-regulation of antiapoptotic proteins. Blood. 2007;109:2399–405.
Hosokawa K, et al. Increased glycosylphosphatidylinositol-anchored protein-deficient granulocytes define a benign subset of bone marrow failures in patients with trisomy 8. Eur J Hematol. 2015;95:230–8.
Katagiri T, Sato-Otsubo A, Kashiwase K, et al. Frequent loss of HLA alleles associated with copy number-neutral 6pLOH in acquired aplastic anemia. Blood. 2011;118:6601–9.
Hosokawa K, Katagiri T, Sugimori N, et al. Favorable outcome of patients who have 13q deletion: a suggestion for revision of the WHO ‘MDS-U’ designation. Haematologica. 2012;97:1845–9.
Jaiswal S, Fontanillas P, Flannick J, et al. Age-related clonal hematopoiesis associated with adverse outcomes. N Engl J Med. 2014;371:2488–98.
Genovese G, Kahler AK, Handsaker RE, et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N Engl J Med. 2014;371:2477–87.
Xie M, et al. Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat Med. 2014;20:1472–8.
Schlush LI, et al. Aging, clonal hematopoiesis and pre-leukemia: not just bad luck? Int J Hematol. 2015;102:513–22.
Steensma, et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndrome. Blood. 2015;126:9–16.
Lane AA, et al. Low frequency clonal mutations recoverable by deep sequencing in patients with aplastic anemia. Leukemia. 2013;27:968–71.
Heuser M, et al. Genetic characterization of acquired aplastic anemia by targeted sequencing. Haematologica. 2014;99(9):e165–7.
Babushok DV, et al. Emergence of clonal hematopoiesis in the majority of patients with acquired aplastic anemia. Cancer Genet. 2015;208:115–28.
Kulasekararaj AG, et al. Somatic mutations identify a subgroup of aplastic anemia patients who progress to myelodysplastic syndrome. Blood. 2014;124:2698–704.
Yoshizato T, et al. Somatic mutations and clonal hematopoiesis in aplastic anaemia. N Eng J Med. 2015;373:35–47.
Dumitriu B, et al. Telomere attrition and candidate gene mutations preceding monosomy 7 in aplastic anemia. Blood. 2015;125:706–9.
Kwok B, et al. MDS-associated somatic mutations and clonal hematopoiesis are common in idiopathic cytopenias of undetermined significance. Blood. 2015;126:2355–61.
Mufti GJ, Kulasekararaj AG, Marsh JC. Somatic mutations and clonal hematopoiesis in Aplastic Anemia. N Engl J Med. 2015;373:1674–5.
Risitano AM, et al. The RACE study: a SAAWP prospective randomized multicenter study comparing horse antithymocyte globulin (hATG) + cyclosporine A (CsA) with or without eltrombopag as front-line therapy for severe aplastic anaemia patients (WP007). Bone Marrow Transplant. 2015;50(S1):pS99.
Olnes MJ, et al. Eltrombopag and improved hematopoiesis in refractory aplastic anemia. N Engl J Med. 2012;367:11–9.
Desmond, et al. Eltrombopag restores tri-lineage hematopoiesis in refractory severe aplastic anemia which can be sustained on discontinuation of drug. Blood. 2014;123:1818–25.
Marsh JC, Mufti GJ. Eltrombopag: a stem cell cookie? Blood. 2014;20(123):1774–5.
Scheinberg P, et al. Association of telomere length of peripheral blood leukocytes with hematopoietic relapse, malignant transformation, and survival in severe aplastic anemia. JAMA. 2010;304:1358–64.
Scheinberg P, et al. Predicting response to immunosuppressive therapy and survival in severe aplastic anaemia. Br J Haematol. 2009;144:206–16.
Bacigalupo A, et al, for the Aplastic Anemia Working Party of the European Group for Blood and Marrow Transplantation (WPSAA-EBMT). Current outcome of HLA identical sibling vs. unrelated donor transplants in severe aplastic anemia: an EBMT analysis. Haematologica. 2015; 100: 696–702.
Marsh JC, et al. Alemtuzumab with fludarabine and cyclophosphamide reduces chronic graft versus host disease after allogeneic stem cell transplantation for acquired aplastic anemia. Blood. 2011;118:2351–7.
Grimaldi, et al. King’s College Hospital FCC conditioning for severe aplastic anemia induces tolerance with mixed T-cell chimerism and extremely low incidence of Gvhd. Blood (Annual Scientific Meeting). 2014;124:1594.
Samarasinghe S, et al. Impact of different in vivo T cell depletion strategies on outcomes following hematopoietic stem cell transplantation for idiopathic aplastic anaemia: a study on behalf of the EBMT SAA Working Party. Blood (Annual Scientific Meeting). 2015;126(23):1210.
Socié G, et al. Malignant tumors occurring after treatment of aplastic anemia. European Bone Marrow Transplantation-Severe Aplastic Anaemia Working Party. N Engl J Med. 1993;329:1152–7.
Kojima S, et al. Japan Childhood Aplastic Anemia Study Group. Risk factors for evolution of acquired aplastic anemia into myelodysplastic syndrome and acute myeloid leukemia after immunosuppressive therapy in children. Blood. 2002;100(3):786–90.
Kordasti S, et al. High resolution mass cytometry (CyTOF) in aplastic anaemia (AA) can identify an aberrant Treg subset with pro-inflammatory properties, predicting poor response to immunosuppressive therapy. Blood. 2014;124:1600 (Annual Scientific Meeting of American Society of Hematology).
Acknowledgments
This work is supported by Bloodwise UK, and the AA&MDS International Foundation.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Marsh, J.C.W., Mufti, G.J. Clinical significance of acquired somatic mutations in aplastic anaemia. Int J Hematol 104, 159–167 (2016). https://doi.org/10.1007/s12185-016-1972-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12185-016-1972-8