Abstract
Purpose
To analyze BRCA1 and BRCA2 genes using a cost-effective and rapid approach based on next generation sequencing (NGS) technology.
Methods
A population of Spanish cancer patients with a personal or familial history of breast and/or ovarian cancer was analyzed for germline mutations in BRCA1 and BRCA2 genes. The methodology relies on a 5 multiplex PCR assay coupled to NGS.
Results
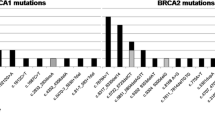
Ten pathogenic mutations (four in BRCA1 and six in BRCA2 gene) were identified in a Spanish population. The deletion c.1792delA, in exon 10, and the duplication c.5869dupA, in exon 11 of BRCA2 gene were not previously reported and should be considered as pathogenic due to its frameshift nature.
Conclusion
Two novel frameshift mutations in BRCA2 gene were detected using the multiplex PCR-based assay following by NGS.


Similar content being viewed by others
References
Ford D, Easton DF, Bishop DT, Narod SA, Goldgar DE. Risks of cancer in BRCA1-mutation carriers. Breast Cancer Link Consort Lancet. 1994;343:692–5.
The breast cancer linkage consortium. Cancer risks in BRCA2 mutation carriers. J Natl Cancer Inst. 1999;91:1310–6.
King MC, Marks JH, Mandell JB, New York Breast Cancer Study Group. Breast and ovarian cancer risks due to inherited mutations in BRCA1 and BRCA2. Science. 2003;302:643–6.
Peto J, Collins N, Barfoot R, Seal S, Warren W, Rahman N, et al. Prevalence of BRCA1 and BRCA2 gene mutations in patients with early-onset breast cancer. J Natl Cancer Inst. 1999;91:943–9.
Loman N, Johannsson O, Kristoffersson U, Olsson H, Borg A. Family history of breast and ovarian cancers and BRCA1 and BRCA2 mutations in a population-based series of early-onset breast cancer. J Natl Cancer Inst. 2001;93:1215–23.
Stratton MR, Wooster R. Hereditary predisposition to breast cancer. Curr Opin Genet Dev. 1996;6:93–7.
Rahman N, Stratton MR. The genetics of breast cancer susceptibility. Annu Rev Genet. 1998;32:95–121.
Díez O, Osorio A, Durán M, Martinez-Ferrandis JI, de la Hoya M, Salazar R, et al. Analysis of BRCA1 and BRCA2 genes in Spanish breast/ovarian cancer patients: a high proportion of mutations unique to Spain and evidence of founder effects. Hum Mutat. 2003;22:301–12.
Frank TS, Manley SA, Olopade OI, Cummings S, Garber JE, Bernhardt B, et al. Sequence analysis of BRCA1 and BRCA2: correlation of mutations with family history and ovarian cancer risk. J Clin Oncol. 1998;16:2417–25.
Gerhardus A, Schleberger H, Schlegelberger B, Gadzicki D. Diagnostic accuracy of methods for the detection of BRCA1 and BRCA2 mutations: a systematic review. Eur J Hum Genet. 2007;15:619–27.
Chan M, Ji SM, Yeo ZX, Gan L, Yap E, Yap YS, et al. Development of a next-generation sequencing method for BRCA mutation screening: a comparison between a high-throughput and a benchtop platform. J Mol Diagn. 2012;14:602–12.
Hernan I, Borràs E, de Sousa Dias M, Gamundi MJ, Mañé B, Llort G, et al. Detection of genomic variations in BRCA1 and BRCA2 genes by long-range PCR and next-generation sequencing. J Mol Diagn. 2012;14:286–93.
Michils G, Hollants S, Dehaspe L, Van Houdt J, Bidet Y, Uhrhammer N, et al. Molecular analysis of the breast cancer genes BRCA1 and BRCA2 using amplicon-based massive parallel pyrosequencing. J Mol Diagn. 2012;14:623–30.
Feliubadaló L, Lopez-Doriga A, Castellsagué E, del Valle J, Menéndez M, Tornero E, et al. Next-generation sequencing meets genetic diagnostics: development of a comprehensive workflow for the analysis of BRCA1 and BRCA2 genes. Eur J Hum Genet. 2013;21:864–70.
De Leeneer K, De Schrijver J, Clement L, Baetens M, Lefever S, De Keulenaer S, et al. Practical tools to implement massive parallel pyrosequencing of PCR prod-ucts in next generation molecular diagnostics. PLoS One. 2011;6:e25531.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hernan, I., Mañé, B., Borràs, E. et al. Two novel frameshift mutations in BRCA2 gene detected by next generation sequencing in a survey of Spanish patients of breast cancer. Clin Transl Oncol 17, 576–580 (2015). https://doi.org/10.1007/s12094-014-1271-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-014-1271-x