Abstract
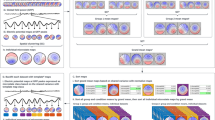
As the size of brain imaging data such as fMRI grows explosively, it provides us with unprecedented and abundant information about the brain. How to reduce the size of fMRI data but not lose much information becomes a more and more pressing issue. Recent literature studies tried to deal with it by dictionary learning and sparse representation methods, however, their computation complexities are still high, which hampers the wider application of sparse representation method to large scale fMRI datasets. To effectively address this problem, this work proposes to represent resting state fMRI (rs-fMRI) signals of a whole brain via a statistical sampling based sparse representation. First we sampled the whole brain’s signals via different sampling methods, then the sampled signals were aggregate into an input data matrix to learn a dictionary, finally this dictionary was used to sparsely represent the whole brain’s signals and identify the resting state networks. Comparative experiments demonstrate that the proposed signal sampling framework can speed-up by ten times in reconstructing concurrent brain networks without losing much information. The experiments on the 1000 Functional Connectomes Project further demonstrate its effectiveness and superiority.












Similar content being viewed by others
References
Abolghasemi, V., Ferdowsi, S., & Sanei, S. (2013). Fast and incoherent dictionary learning algorithms with application to fMRI. Signal, Image and Video Processing, 1–12.
Corbetta, M., Patel, G., & Shulman, G. L. (2008). The reorienting system of the human brain: from environment to theory of mind. Neuron, 58(3), 306–324. doi:10.1016/j.neuron.2008.04.017.
Daubechies, I., Roussos, E., Takerkart, S., Benharrosh, M., Golden, C., D’Ardenne, K., Richter, W., Cohen, J. D., & Haxby, J. (2009). Independent component analysis for brain fMRI does not select for independence. Proceedings of the National Academy of Sciences, 106(26), 10415–10422.
Duncan, J. (2010). The multiple-demand (MD) system of the primate brain: mental programs for intelligent behaviour. Trends in Cognitive Sciences, 14(4), 172–179. doi:10.1016/j.tics.2010.01.004.
Eavani, H., Filipovych, R., Davatzikos, C., Satterthwaite, T.D., Gur, R.E., & Gur, R.C. (2012). Sparse dictionary learning of resting state fMRI networks. International Workshop Pattern Recognition Neuroimaging, 73–76, doi:10.1109/PRNI.2012.25.
Gazzaniga, M.S. (2004). The cognitive neurosciences. MIT press.
Kalcher, K., Huf, W., Boubela, R. N., Filzmoser, P., Pezawas, L., Biswal, B., et al. (2012). Fully exploratory network independent component analysis of the 1000 functional connectomes database. Frontiers in Human Neuroscience, 6, 301. doi:10.3389/fnhum.2012.00301.
Lee, K., Tak, S., & Ye, J. C. (2011). A data-driven sparse GLM for fMRI analysis using sparse dictionary learning with MDL criterion. IEEE Transactions on Medical Imaging, 30(5), 1076–1089. doi:10.1109/Tmi.2010.2097275.
Li, Y., Namburi, P., Yu, Z., Guan, C., Feng, J., & Gu, Z. (2009). Voxel selection in fMRI data analysis based on sparse representation. Biomedical Engineering, IEEE Transactions on, 56(10), 2439–2451.
Li, K., Guo, L., Li, G., Nie, J., Faraco, C., Zhao, Q., et al. (2010). Cortical surface based identification of brain networks using high spatial resolution resting state FMRI data. In Biomedical Imaging: From Nano to Macro, 2010 I.E. International Symposium on, (pp. 656–659): IEEE.
Li, Y., Long, J., He, L., Lu, H., Gu, Z., & Sun, P. (2012). A sparse representation-based algorithm for pattern localization in brain imaging data analysis. PloS One, 7(12), e50332. doi:10.1371/journal.pone.0050332.
Lin, B., Li, Q., Sun, Q., Lai, M.-J., Davidson, L., Fan, W., et al. (2014). Stochastic coordinate coding and its application for drosophila gene expression pattern annotation. arXiv:1407.8147v2 [cs.LG].
Liu, T., Li, H., Wong, K., Tarokh, A., Guo, L., & Wong, S. T. (2007). Brain tissue segmentation based on DTI data. NeuroImage, 38(1), 114–123. doi:10.1016/j.neuroimage.2007.07.002.
Liu, T., Nie, J., Tarokh, A., Guo, L., & Wong, S. T. C. (2008). Reconstruction of central cortical surface from brain MRI images: method and application. NeuroImage, 40(3), 991–1002.
Lv, J., Jiang, X., Li, X., Zhu, D., Chen, H., Zhang, T., et al. (2014a). Sparse representation of whole-brain fMRI signals for identification of functional networks. Medical Image Analysis. doi:10.1016/j.media.2014.10.011.
Lv, J., Jiang, X., Li, X., Zhu, D., Zhang, S., Zhao, S., et al. (2014b). Holistic atlases of functional networks and interactions reveal reciprocal organizational architecture of cortical function. IEEE Transactions on Biomedical Engineering. doi:10.1109/TBME.2014.2369495.
Ma, P., Mahoney, M. W., & Yu, B. (2015). A statistical perspective on algorithmic leveraging. Journal of Machine Learning Research, 16, 861–911.
Mahoney, M. W. (2011). Randomized algorithms for matrices and data. Foundations and Trends® in Machine Learning, 3(2), 123–224.
Mairal, J., Bach, F., Ponce, J., & Sapiro, G. (2010). Online learning for matrix factorization and sparse coding. Journal of Machine Learning Research, 11, 19–60.
McKeown, M. J., et al. (1998). Spatially independent activity patterns in functional MRI data during the Stroop color-naming task. PNAS, 95(3), 803.
Meng, X., Saunders, M. A., & Mahoney, M. W. (2014). LSRN: a parallel iterative solver for strongly over-or underdetermined systems. SIAM Journal on Scientific Computing, 36(2), C95–C118.
Oikonomou, V. P., Blekas, K., & Astrakas, L. (2012). A sparse and spatially constrained generative regression model for fMRI data analysis. IEEE Transactions on Biomedical Engineering, 59(1), 58–67. doi:10.1109/TBME.2010.2104321.
Pessoa, L. (2012). Beyond brain regions: network perspective of cognition-emotion interactions. Behavioral and Brain Sciences, 35(3), 158–159. doi:10.1017/S0140525x11001567.
Rao, P. (2000). Sampling methodologies with applications. Chapman & Hall/CRC.
Smith, S. M., Fox, P. T., Miller, K. L., Glahn, D. C., Fox, P. M., Mackay, C. E., et al. (2009). Correspondence of the brain’s functional architecture during activation and rest. Proceedings of the National Academy of Sciences of the United States of America, 106(31), 13040–13045. doi:10.1073/pnas.0905267106.
Smith, S. M., Beckmann, C. F., Andersson, J., Auerbach, E. J., Bijsterbosch, J., Douaud, G., et al. (2013). Resting-state fMRI in the human connectome project. NeuroImage, 80, 144–168. doi:10.1016/j.neuroimage.2013.05.039.
Sotiropoulos, S. N., Moeller, S., Jbabdi, S., Xu, J., Andersson, J. L., Auerbach, E. J., et al. (2013). Effects of image reconstruction on fiber orientation mapping from multichannel diffusion MRI: reducing the noise floor using SENSE. Magnetic Resonance in Medicine, 70(6), 1682–1689. doi:10.1002/Mrm.24623.
Tillé, Y. (2011). Sampling algorithms. Springer.
Van Essen, D. C., Smith, S. M., Barch, D. M., Behrens, T. E., Yacoub, E., & Ugurbil, K. (2013). The WU-minn human connectome project: an overview. NeuroImage, 80, 62–79. doi:10.1016/j.neuroimage.2013.05.041.
Varoquaux, G., Gramfort, A., Pedregosa, F., Michel, V., & Thirion, B. (2011). Multi-subject dictionary learning to segment an atlas of brain spontaneous activity. Informaiton Processing Medical Imaging, 22, 562–573.
Wright, J., Ma, Y., Mairal, J., Sapiro, G., Huang, T. S., & Yan, S. C. (2010). Sparse representation for computer vision and pattern recognition. Proceedings of the Ieee, 98(6), 1031–1044. doi:10.1109/Jproc.2010.2044470.
Yan, C. G., Craddock, R. C., Zuo, X. N., Zang, Y. F., & Milham, M. P. (2013). Standardizing the intrinsic brain: towards robust measurement of inter-individual variation in 1000 functional connectomes. NeuroImage, 80, 246–262. doi:10.1016/j.neuroimage.2013.04.081.
Yates, D., Moore, D. S., & Starnes, D.S. (2002). The practice of statistics: TI-83/89 graphing calculator enhanced. Macmillan.
Zhu, D., Li, K., Guo, L., Jiang, X., Zhang, T., Zhang, D., et al. (2013). DICCCOL: dense individualized and common connectivity-based cortical landmarks. Cerebral Cortex, 23(4), 786–800. doi:10.1093/cercor/bhs072.
Acknowledgments
We thank all investigators contributing data to the 1000 Functional Connectomes project, without whom this analysis could not have been performed. T Liu was supported by NIH R01 DA-033393, NIH R01 AG-042599, NSF CAREER Award IIS-1149260, NSF CBET-1302089 and NSF BCS-1439051. B Ge was supported by NSFC 61403243, 2015JM6312, the Fundamental Research Funds for the Central Universities from China (No. GK201402008) and Interdisciplinary Incubation Project of Learning Science of Shaanxi Normal University. The authors would like to thank the anonymous reviewers for their constructive comments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
None of the authors has conflict of interest to declare.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Ge, B., Makkie, M., Wang, J. et al. Signal sampling for efficient sparse representation of resting state FMRI data. Brain Imaging and Behavior 10, 1206–1222 (2016). https://doi.org/10.1007/s11682-015-9487-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11682-015-9487-0