Abstract
Myocardial infarction (MI) is an important clinical problem because of its large contribution to mortality. The main causal and treatable risk factors for MI include hypertension, hypercholesterolemia or dyslipidemia, diabetes mellitus, and smoking. In addition to these risk factors, recent studies have shown the importance of genetic factors and interactions between multiple genes and environmental factors. Disease prevention is an important strategy for reducing the overall burden of MI, with the identification of markers for disease risk being key both for risk prediction and for potential intervention to lower the chance of future events. Although genetic linkage analyses of families and sib-pairs as well as candidate gene and genome-wide association studies have implicated several loci and candidate genes in predisposition to coronary heart disease (CHD) or MI, the genes that contribute to genetic susceptibility to these conditions remain to be identified definitively. In this review, we summarize both candidate loci for CHD or MI identified by linkage analyses and candidate genes examined by association studies. We also review in more detail studies that have revealed the association with MI or CHD of polymorphisms in MTHFR, LPL, and APOE by the candidate gene approach and those in LTA and at chromosomal region 9p21.3 by genome-wide scans. Such studies may provide insight into the function of implicated genes as well as into the role of genetic factors in the development of CHD and MI.
Similar content being viewed by others
Introduction
Recent progress in human genetics and genomics research, highlighted by completion of the nucleotide sequence of the human genome by the Human Genome Project (International Human Genome Sequencing Consortium 2004), has provided substantial benefits to clinical medicine, including facilitation of the characterization of disease pathogenesis at the molecular level and the development of panels of genetic markers for assessment of disease risk. In particular, determination of single nucleotide polymorphisms (SNPs) and haplotype blocks and the specification of tag SNPs in each haplotype block for four ethnic groups by the International HapMap Project (The International HapMap Consortium 2007) have led to increasingly effective approaches to the identification of genetic variation associated with various multifactorial diseases, providing new insight into the pathogenesis of these conditions. Furthermore, technological developments such as cDNA microarrays and SNP chips that provide huge amounts of genetic information have made possible the detection of genetic differences among individuals at the whole-genome level.
Selection of the most appropriate strategies for disease prevention or therapy on the basis of genetic information for a given individual is referred to as personalized or individualized medicine. In conventional medicine, medications are prescribed on the basis of the diagnosis and severity of the disease. However, the efficacy of drugs and the incidence of side effects vary among individuals. The goal of treatment based on genetic or genomic information is to be able to predict therapeutic outcome or side effects in an individual, thereby increasing the effectiveness and safety of therapy. In addition, the clarification of disease etiologies at the molecular level and the identification of genetic variants that confer disease susceptibility are likely to contribute both to disease prevention and to the development of new medicines.
Myocardial infarction (MI) is an important clinical problem because of its large contribution to mortality. In the United States, the total number of individuals affected by coronary heart disease (CHD) was 15.8 million in 2004, with nearly 450,000 patients dying annually from this condition (Rosamond et al. 2007). The annual incidence of MI was 565,000 new attacks and 300,000 recurrent attacks, with an annual mortality of 157,000 (Rosamond et al. 2007). As in the United States, CHD is the most common cause of death in the United Kingdom, where it is responsible for around 101,000 deaths each year (British Heart Foundation; http://www.heartstats.org/homepage.asp). In Japan, the total number of individuals affected by CHD was 910,000 in 2005, and ~50,000 people die annually from MI (Ministry of Health, Labor, and Welfare of Japan).
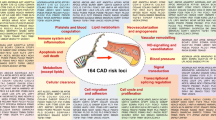
The main causal and treatable risk factors for MI include hypertension, hypercholesterolemia or dyslipidemia, diabetes mellitus, and smoking. In addition to these risk factors, recent studies have shown the importance of genetic factors and of interactions between multiple genes and environmental factors in this condition (Arnett et al. 2007; Kullo and Ding 2007; Topol et al. 2006). The common forms of CHD and MI are thus thought to be multifactorial and to be determined by many genes, each with a relatively small effect, working alone or in combination with modifier genes or environmental factors (or both). The “common disease, common variants hypothesis” proposes that genetic variants present in many normal individuals contribute to overall CHD risk. In addition, susceptibility to some common diseases may be conferred, in part, by rarer variants (Arnett et al. 2007).
Despite recent advances in therapy, such as drug-eluting stents (Marroquin et al. 2008), for acute coronary syndrome (ACS), CHD remains the leading cause of death in the US and UK and the second leading cause of death in Japan. Disease prevention is an important strategy for reducing the overall burden of CHD and MI, and the identification of biomarkers for disease risk is key both for risk prediction and for potential intervention to reduce the chance of future events.
Linkage analysis of MI, ACS, or CHD
Several genome-wide linkage analyses of families or sib-pairs have identified chromosomal loci linked to or genetic variations that confer susceptibility to MI, ACS, or CHD (Broeckel et al. 2002; Farrall et al. 2006; Francke et al. 2001; Harrap et al. 2002; Hauser et al. 2004; Helgadottir et al. 2004; Pajukanta et al. 2000; The BHF Family Heart Study Research Group 2005; Wang et al. 2003, 2004). The published results of genome-wide linkage analyses for these conditions are summarized in Table 1. Genomic regions identified in the published linkage studies as being correlated with MI or CHD are largely nonoverlapping, suggestive of genetic complexity in which multiple genes are responsible for the development of these conditions, although phenotypic heterogeneity could also have contributed to the nonreplicability of results.
The deCODE Genetics group (Helgadottir et al. 2004) performed linkage analysis with 1,068 microsatellite markers and found a linkage peak (LOD score of 2.86) at chromosomal region 13q12-q13 for 296 Icelandic families (713 individuals) enrolled on the basis of a history of MI. The researchers then genotyped an additional 120 microsatellite markers in this interval in 802 individuals with MI and 837 controls, and they found that a four-marker SNP haplotype spanning the arachidonate 5-lipoxygenase-activating protein gene (ALOX5AP) was associated with MI (odds ratio, 1.8) and stroke (odds ratio, 1.7). A subsequent study found that ALOX5AP was associated with CHD in British individuals and with stroke in Icelandic and Scottish populations (Helgadottir et al. 2005).
On the basis of the results of the same genome-wide scan, the deCODE Genetics group (Helgadottir et al. 2006) performed fine mapping to determine that a five- to seven-marker SNP haplotype of the leukotriene A4 hydrolase gene (LTA4H) accounted for a linkage peak at 12q22. Of particular interest with this haplotype was its ancestry-specific association with the incidence and risk of MI. In European-Americans, the relative risk for MI was only 1.2, with a population attributable risk of 4.6%, whereas among individuals of African ancestry, the relative risk was 3.5 and the population attributable risk was 14% (Helgadottir et al. 2006). Two different genes (ALOX5AP and LTA4H) in the same inflammation-related pathway of leukotriene B4 production were thus found to be associated with disease in a single genome-wide scan. This pathway had already been implicated in studies of murine experimental atherosclerosis as well as in human epidemiological and pathological studies (Dwyer et al. 2004; Mehrabian et al. 2002; Spanbroek et al. 2003). In addition, a small-molecule inhibitor of ALOX5AP was shown to reduce both leukotriene production and the plasma concentration of C-reactive protein (CRP), an important biomarker for CHD, in a pilot, placebo-controlled, randomized trial with individuals harboring the risk ALOX5AP or LTA4H haplotype (Hakonarson et al. 2005). Of note, LTA4H was the first MI-linked gene to show an ancestry-specific risk (Damani and Topol 2007; Topol et al. 2006).
Association studies of MI or CHD
Various association studies of unrelated individuals have identified genetic variations that confer susceptibility to MI or CHD. The published results for genes associated with these conditions are summarized in Table 2. Numerous candidate genes have been implicated, but those that show reproducible associations between risk alleles and CHD or MI in replication studies are few. The candidate gene approach has been widely applied to analysis of the possible association between genetic variants and disease, with genes selected on the basis of a priori hypotheses regarding their potential etiologic role. It is characterized as a hypothesis-testing approach because of the biological observation supporting a role for the proposed candidate gene. The candidate gene approach is not able, however, to identify disease-associated polymorphisms in unknown genes. The recent development of high-density genotyping arrays has improved the resolution of unbiased genome-wide scans for common variants associated with multifactorial diseases. Currently, the genome-wide association study (GWAS) makes use of high-throughput genotyping technologies that include about 1 million probes for SNPs and 1 million probes for copy number variations to examine their relation to clinical conditions or measurable traits. Since 2005, nearly 100 loci for as many as 40 common diseases or traits have been identified by GWASs, many in genes not previously suspected of having a role in the condition studied, and some in genomic regions containing no known genes. Although GWASs represent a substantial advance in the search for genetic variants that influence disease, they also have important limitations, including the potential for generating false-positive or false-negative results and for biases related to the selection of study participants and genotyping errors (Pearson and Manolio 2008).
Mendelian randomization
Mendelian randomization analysis is a relatively recent development in genetic epidemiology based on Mendel’s second law, which states that the inheritance of one trait is independent of that of other traits (Davey Smith and Ebrahim 2003; Keavney 2002). It relies on common genetic polymorphisms that are known to influence exposure patterns (such as the propensity to drink alcohol) or to have effects equivalent to those produced by modifiable exposures (such as an increased serum cholesterol concentration). Associations between genetic variants and outcomes are not generally confounded by behavioral or environmental exposures, with the result that observational studies of genetic variants have similar properties to intention-to-treat analyses in randomized controlled trials. The simplest way of appreciating the potential of Mendelian randomization analysis is to consider applications of the underlying principles. The inferences that can be drawn from Mendelian randomization studies depend on the different ways in which genetic variants can serve as a proxy for environmentally modifiable exposures (Davey Smith and Ebrahim 2005).
The relations of polymorphisms of the CRP gene (CRP) to circulating CRP concentrations and the prevalence of CHD or MI have been examined by Mendelian randomization analysis. Pooled data from 4,659 Caucasian men in six studies revealed that individuals homozygous for the T allele of the 1444C→T polymorphism of CRP had a higher circulating CRP concentration than carriers of the C allele. However, men with the TT genotype were not at increased risk of nonfatal MI (Casas et al. 2006). This unbiased and nonconfounded estimate of the effect of CRP genotype on coronary events was smaller than estimates provided by previous studies. In two independent prospective cohort studies of 32,826 women and 18,225 men in the US, the minor alleles of 1919A→T and 4741G→C polymorphisms of CRP were associated with higher plasma CRP levels, and those of 2667G→C and 3872C→T polymorphisms of CRP were associated with lower plasma CRP levels. Two of the five common haplotypes of CRP were associated with lower CRP levels. However, neither the individual SNPs nor the common haplotypes were associated with risk of CHD in the direction that would be predicted by their association with CRP levels (Pai et al. 2008). These data suggest that the underlying inflammatory processes that predict coronary events cannot be captured solely by variation in CRP. The CRP CHD Genetics Collaboration is a consortium of investigators generating and pooling analyses of data on genetic determinants of circulating CRP levels and CHD. These data should help to clarify the likelihood and magnitude of any causal association between circulating CRP concentration and CHD. The collaboration is likely to advance understanding of the relevance of low-grade inflammation to CHD and indicate whether or not CRP itself should be prioritized as a therapeutic target for long-term prevention strategies (CRP CHD Genetics Collaboration 2008).
Candidate gene association studies for MI or CHD
Association studies based on the candidate gene approach have revealed many polymorphisms to be associated with the prevalence of MI or CHD (Table 2). In this section, we discuss the association of polymorphisms in MTHFR, LPL, and APOE with MI or CHD.
MTHFR
Homocysteine is a sulfur-containing amino acid that plays a pivotal role in methionine metabolism. 5,10-Methylenetetrahydrofolate reductase (MTHFR) catalyzes the reduction of 5,10-methylenetetrahydrofolate to 5-methylenetetrahydrofolate, a reaction that provides a substrate for the methylation of homocysteine to methionine catalyzed by methionine synthase. Individuals with the 677C→T (Ala222Val) substitution of MTHFR manifest reduced MTHFR activity and higher plasma homocysteine levels compared with those without it (Deloughery et al. 1996; Ma et al. 1996; Schwartz et al. 1997). Association of the 677C→T (Ala222Val) polymorphism of MTHFR with CHD or MI has been described by several groups, with the TT genotype being a risk factor for these conditions (Gallagher et al. 1996; Kluijtmans et al. 1996; Mager et al. 1999; Morita et al. 1997; Yamada et al. 2006). Other studies, however, did not support such an association (Folsom et al. 1998; Schwartz et al. 1997). These apparently contradictory results are attributable, at least in part, to differences in intake of folate and other B vitamins (Verhoef et al. 1998). A meta-analysis of the association of the 677C→T (Ala222Val) polymorphism of MTHFR with the risk of CHD in 11,162 cases and 12,758 controls from 40 studies revealed that individuals with the TT genotype had an odds ratio of 1.16 for CHD compared with those with the CC genotype (Klerk et al. 2002). These observations suggest that impaired folate metabolism, resulting in high homocysteine concentrations, is an important determinant of CHD. Another meta-analysis of the association of the 677C→T (Ala222Val) polymorphism of MTHFR with CHD in 26,000 cases and 31,183 controls from 80 studies yielded an overall odds ratio of 1.14 for the TT genotype versus the CC genotype; odds ratios for Europe, Australia, and North America were about 1.0, whereas those for the Middle East and Asia were 2.61 and 1.23, respectively (Lewis et al. 2005). These results indicate that the 677C→T (Ala222Val) polymorphism of MTHFR is associated with CHD in the Middle East and Asia, but not in Europe, North America, or Australia, with this geographic variability possibly reflecting higher folate intake in the latter regions (Lewis et al. 2005).
LPL
Lipoprotein lipase (LPL) is the rate-limiting enzyme in lipolysis of triglyceride-rich lipoproteins in the circulation. It is synthesized in parenchymal cells of adipose tissue as well as in skeletal and cardiac muscle, and it is then transferred to heparan sulfate-binding sites of the vascular endothelium (Kastelein et al. 2000). The hydrolytic function of LPL is important for the processing of triglyceride-rich chylomicrons and very low density lipoproteins to remnant particles as well as for the transfer of phospholipids and apolipoproteins to high density lipoproteins (HDLs). LPL also plays an important role in the receptor-mediated removal of lipoproteins from the circulation (Groenemeijer et al. 1997). LPL is polymorphic, with amino acid substitutions of the encoded protein affecting triglyceride and HDL-cholesterol levels, which are implicated in atherosclerosis risk (Wittrup et al. 1999). The 1595C→G (Ser447Stop) substitution of LPL results in carboxyl-terminal truncation of LPL by two amino acids. This change is thought to increase the binding affinity of the protein for receptors or to facilitate or otherwise affect its formation of dimers (Wittrup et al. 1999). The G (Stop) allele of the 1595C→G (Ser447Stop) polymorphism has also been shown to be related to decreased plasma triglyceride or increased HDL-cholesterol levels, or both (Groenemeijer et al. 1997; Jemaa et al. 1995; Kuivenhoven et al. 1997; Wittrup et al. 1999). In addition, the G (Stop) allele of this polymorphism was found to be associated with a reduced risk of CHD or MI (Wittrup et al. 1999; Yamada et al. 2006; Yang et al. 2004). Evidence suggests that the catalytic activity and stability of the truncated variant of LPL may be largely normal, but that it may be present at higher concentrations in the circulation, resulting in a higher level of LPL activity (Groenemeijer et al. 1997; Henderson et al. 1999; Humphries et al. 1998; Zhang et al. 1996).
APOE
Apolipoprotein E (ApoE) plays an important role in lipid transport and metabolism. Three common alleles (ε2, ε3, and ε4) of APOE encode the three major isoforms (E2, E3, and E4) of ApoE, which differ at amino acid positions 112 and 158. Allelic variation of APOE accounts for interindividual variability in total cholesterol and low density lipoprotein (LDL)–cholesterol concentrations, with studies in human populations demonstrating associations of the ε4 and ε2 alleles with increased and decreased LDL-cholesterol levels, respectively (Ehnholm et al. 1986; Sing and Davignon 1985; Xhignesse et al. 1991). The various ApoE isoforms interact differently with specific lipoprotein receptors, ultimately affecting circulating levels of cholesterol (Eichner et al. 2002). ApoE from very low density lipoprotein, chylomicrons, and chylomicron remnants binds to specific receptors on cells in the liver. Carriers of the ε2 allele of APOE are less efficient than carriers of the ε3 or ε4 alleles at synthesizing very low density lipoprotein and chylomicrons and at transferring them from plasma to the liver as a result of the binding properties of the ApoE2 isoform. Thus, compared with carriers of the ε3 or ε4 alleles, carriers of the ε2 allele are slower to clear dietary fat from their blood (Weintraub et al. 1987). The difference in uptake of postprandial lipoprotein particles results in differences in regulation of hepatic LDL receptors, which in turn contribute to genotypic differences in total and LDL-cholesterol levels (Davignon et al. 1988; Hallman et al. 1991; Schaefer et al. 1994).
The relation of APOE polymorphisms to CHD or MI has been extensively investigated in the last 2 decades. In many studies, the ε4 allele has been associated with CAD or MI (Lahoz et al. 2001; van Bokxmeer and Mamotte 1992; Wilson et al. 1994). A meta-analysis of 15,492 subjects with CHD and 32,965 controls pooled from 48 studies revealed that, compared with individuals with the ε3/ε3 genotype, carriers of the ε4 allele had a higher risk for CHD (odds ratio, 1.42), whereas the ε2 allele was not associated with CHD risk (Song et al. 2004). The ε4 allele of APOE is thus an important risk factor for CHD.
The -219G→T SNP of APOE has been associated with MI for men in France and Northern Ireland, with the T allele representing a risk factor for this condition (Lambert et al. 2000). Consistent with its location in the promoter region of APOE, the -219G→T SNP was shown to be associated with the plasma concentration of ApoE, with the T allele conferring a reduced ApoE concentration (Lambert et al. 2000). The deleterious influence of the T allele on MI therefore cannot be explained by its effect on the circulating level of ApoE. The T allele of this SNP was also shown to be a risk factor for CHD in low-risk Japanese men (Hirashiki et al. 2003).
Genome-wide association studies of MI or CHD
GWASs have identified susceptibility genes for various multifactorial diseases, including CHD and MI (Table 3).
LTA
Screening of 65,671 SNPs revealed that two polymorphisms of the lymphotoxin-α gene (LTA) were associated with susceptibility to MI in a study with 1,133 MI patients and 1,878 controls (Ozaki et al. 2002). Functional analysis in vitro indicated that the G allele of one of these two polymorphisms, 252A→G in intron 1 (rs909253), was associated with an increase in the transcriptional activity of LTA and that the A (Asn) allele of the second SNP, 804C→A (Thr26Asn) in exon 3 (rs1041981), was associated with increased expression of the genes for vascular cell adhesion molecule 1 and selectin E. Ozaki et al. (2002) thus suggested that variants of LTA are risk factors for MI and that they influence the vascular inflammation that underlies this condition. These researchers subsequently showed that the 3279C→T polymorphism in intron 1 of the lectin, galactoside-binding, soluble, two gene (LGALS2) was associated with the prevalence of MI (Ozaki et al. 2004). LGALS2 plays a role in the secretion of LTA from smooth muscle cells and macrophages, and the identified polymorphism affects the transcriptional activity of LGALS2. These results suggested that an LGALS2–LTA axis is important in the pathophysiology of coronary atherosclerosis and thrombosis.
The relation of seven SNPs (rs2071590, rs1800683, rs909253, rs746868, rs2857713, rs3093543, and rs1041981) distributed throughout LTA and of their corresponding haplotypes to risk of MI was examined in the International Study of Infarct Survival (ISIS) case–control study involving 6,928 cases of nonfatal MI and 2,712 unrelated controls (Clarke et al. 2006). The seven SNPs were in strong linkage disequilibrium with each other and formed six common haplotypes. None of the SNPs or haplotypes was associated with risk of MI. A meta-analysis of rs909253 or rs1041981 in six previously published studies and the ISIS study (Clarke et al. 2006) found no association with CHD risk in a recessive model (odds raio, 1.07) and only a moderate association in a dominant model (odds raio, 1.09). Overall, these studies suggest that these common polymorphisms of LTA are not associated with susceptibility to CHD or MI. Given that the effect of LTA variants on the development of MI might differ among ethnic groups or among individuals exposed to different environmental factors such as smoking, further investigation is warranted with large independent subject panels of different ethnic groups.
Chromosome 9p21.3
In 2007, independent GWASs based on the use of SNP chips identified four SNPs on chromosome 9p21.3 that were associated with CHD or MI in several white cohorts (Helgadottir et al. 2007; McPherson et al. 2007; Samani et al. 2007; Wellcome Trust Case Control Consortium 2007). McPherson et al. (2007) identified two susceptibility SNPs (rs10757274 and rs2383206) that were located within 20 kbp of each other on chromosome 9p21.3 and were associated with CHD in a Canadian population and five other white cohorts. Helgadottir et al. (2007) described an association between MI and two SNPs (rs2383207 and rs10757278) located in the same 9p21.3 region in an Icelandic population, and they replicated the finding in four white cohorts. The same genetic locus was also identified by a GWAS performed with 1,926 CHD cases and 3,000 controls from a British population (Wellcome Trust Case Control Consortium 2007), and the finding was replicated in a German population (Samani et al. 2007). Association of SNPs on chromosome 9p21.3 was also replicated for MI in an Italian population (Shen et al. 2008b) and for CHD in a Korean population (Shen et al. 2008a). Interestingly, the independent population-based case–control studies also identified several SNPs at 9p21.3 that were significantly associated with type 2 diabetes mellitus in white populations in England (Zeggini et al. 2007), Finland (Scott et al. 2007), and Sweden (Saxena et al. 2007). In addition to MI, SNP rs10757278 at this locus was found to be associated with abdominal aortic aneurysm and intracranial aneurysm (Helgadottir et al. 2008). Schunkert et al. (2008) genotyped a SNP (rs1333049) representing the 9p21.3 locus in seven case–control studies including a total of 4,645 subjects with MI or CHD and 5,177 controls. The risk allele (C) of this SNP was uniformly associated with MI or CHD in each study, with pooled analysis revealing the odds ratio per copy of the risk allele to be 1.29. Meta-analysis of rs1333049 in 12,004 cases and 28,949 controls provided further evidence for association of this SNP with MI or CHD, yielding an odds ratio of 1.24 per risk allele.
The prospective Northwick Park Heart Study II analyzed complete trait and genotype information available for 2,057 men (183 CHD events over 10.8 years). For a panel of selected genotypes for UCP2, APOE, LPL, APOA4, IL6, and PECAM1, CHD risk estimates incorporating conventional risk factors (age, triglyceride and cholesterol levels, systolic blood pressure, and smoking) and genetic risk interactions were more effective than those based on conventional risk factors alone (Humphries et al. 2007). In a study of the same cohort involving 2,742 men (270 CHD events over 15 years), although rs10757274 at 9p21.3 was associated with CHD, it did not add substantially to the usefulness of the Framingham risk score based on conventional risk factors alone for predicting future CHD events. However, it did improve reclassification of CHD risk and thus may be of clinical utility (Talmud et al. 2008).
Although this broad replication of the association with chromosome 9p21.3 provides important new information on the molecular genetics of CHD and MI, the underlying mechanism is as yet elusive. The region is defined by two flanking recombination hot spots and contains the coding sequences of genes for two cyclin-dependent kinase inhibitors, CDKN2A and CDKN2B. These genes play an important role in regulation of the cell cycle and belong to a family of genes that have been implicated in the pathogenesis of atherosclerosis as a result of their contribution to inhibition of cell growth by transforming growth factor-β1. However, the SNPs associated most strongly with MI or CHD lie considerably upstream of these genes, with the nearest being located 10 kbp upstream of CDKN2B. Although an effect mediated through one or both of these genes is possible, other explanations for the association of the 9p21.3 region with MI or CHD need to be considered (Schunkert et al. 2008).
The high-risk CHD haplotype at 9p21.3 [T (rs10116277)–T (rs6475606)–G (rs10738607)–T (rs10757272)–G (rs10757274)–G (4977574)–G (2891168)–G (1333042)–G (2383206)–G (2383207)–C (1333045)–G (10757278)–C (1333048)–C (1333049)] was recently shown to overlap with exons 13 to 19 of ANRIL (Broadbent et al. 2008) (Fig. 1), a newly annotated gene for a large antisense noncoding RNA that was identified by deletion analysis of an extended French family with hereditary melanoma–neural system tumors (Pasmant et al. 2007). Reverse transcription and polymerase chain reaction analysis showed that ANRIL is expressed in atheromatous human vessels (specimens of abdominal aortic aneurysm or carotid endarterectomy), which manifest a cell type profile similar to that of atherosclerotic coronary arteries. ANRIL was found to be expressed in vascular endothelial cells, monocyte-derived macrophages, and coronary smooth muscle cells (Broadbent et al. 2008), all of which contribute to atherosclerosis. Little is known of the function of ANRIL, as is typical of most genes for noncoding RNAs, which in general are thought to participate in transcriptional control (Mattick and Makunin 2006). A survey of the dbSNP database revealed no SNPs that map within the exons of ANRIL that colocalize with the risk haplotype. However, multiple SNPs coupled to the high-risk haplotype map to intronic or downstream sequences of this gene; these variants are plausible candidates for determinants of the level of ANRIL expression. The targets of ANRIL action remain to be discovered, as do any interactions with neighboring genes (Broadbent et al. 2008). Clarification of the functional relevance of SNPs at 9p21.3 to CHD and MI may provide insight into the pathogenesis of these conditions as well as into the role of genetic factors in their development.
Conclusion
There has been a growing effort to find genetic variants that confer risk for CHD and MI as a means to understand the underlying biological events of these conditions. Such studies may ultimately lead to the personalized prevention of MI (Yamada 2006). It may thus become possible to predict the future risk for MI in each individual on the basis of conventional laboratory examinations and genetic analyses. It should also be possible to assess how the risk level of an individual will decrease if treatable risk factors, including hypertension, diabetes mellitus, hypercholesterolemia or dyslipidemia, and smoking, are ameliorated or eliminated. Furthermore, it may be possible to prevent an individual from undergoing MI by medical intervention based on his or her genotype for specific polymorphisms. In the future, we may have the ability to use specific therapeutic agents individualized on the basis of certain genetic susceptibility factors, thereby increasing the efficacy and limiting the toxicity of treatment (Damani and Topol 2007). Identification of disease susceptibility genes will thus contribute to the prevention, early diagnosis, and treatment of CHD and MI.
Abbreviations
- SNP:
-
Single nucleotide polymorphism
- MI:
-
Myocardial infarction
- CHD:
-
Coronary heart disease
- ACS:
-
Acute coronary syndrome
- CRP:
-
C-reactive protein
- GWAS:
-
Genome-wide association study
- HDL:
-
High density lipoprotein
- LDL:
-
Low density lipoprotein
References
Amisten S, Melander O, Wihlborg AK, Berglund G, Erlinge D (2007) Increased risk of acute myocardial infarction and elevated levels of C-reactive protein in carriers of the Thr-87 variant of the ATP receptor P2Y11. Eur Heart J 28:13–18. doi:10.1093/eurheartj/ehl410
Arnett DK, Baird AE, Barkley RA, Basson CT, Boerwinkle E et al (2007) Relevance of genetics and genomics for prevention and treatment of cardiovascular disease: a scientific statement from the American Heart Association Council on Epidemiology and Prevention, the Stroke Council, and the Functional Genomics and Translational Biology Interdisciplinary Working Group. Circulation 115:2878–2901. doi: 10.1161/CIRCULATIONAHA.107.183679
Bacci S, Ludovico O, Prudente S, Zhang YY, Di Paola R et al (2005) The K121Q polymorphism of the ENPP1/PC-1 gene is associated with insulin resistance/atherogenic phenotypes, including earlier onset of type 2 diabetes and myocardial infarction. Diabetes 54:3021–3025. doi:10.2337/diabetes.54.10.3021
Behague I, Poirier O, Nicaud V, Evans A, Arveiler D et al (1996) β Fibrinogen gene polymorphisms are associated with plasma fibrinogen and coronary artery disease in patients with myocardial infarction. The ECTIM Study. Etude Cas-Temoins sur l’Infarctus du Myocarde. Circulation 93:440–449
Broadbent HM, Peden JF, Lorkowski S, Goel A, Ongen H (2008) Susceptibility to coronary artery disease and diabetes is encoded by distinct, tightly linked SNPs in the ANRIL locus on chromosome 9p. Hum Mol Genet 17:806–814. doi: 10.1093/hmg/ddm352
Broeckel U, Hengstenberg C, Mayer B, Holmer S, Martin LJ et al (2002) A comprehensive linkage analysis for myocardial infarction and its related risk factors. Nat Genet 30:210–214. doi:10.1038/ng827
Cambien F, Poirier O, Lecerf L, Evans A, Cambou J-P et al (1992) Deletion polymorphism in the gene for angiotensin-converting enzyme is a potent risk factor for myocardial infarction. Nature 359:641–644. doi:10.1038/359641a0
Casas JP, Shah T, Cooper J, Hawe E, McMahon AD et al (2006) Insight into the nature of the CRP-coronary event association using Mendelian randomization. Int J Epidemiol 35:922–931. doi:10.1093/ije/dyl041
Cipollone F, Toniato E, Martinotti S, Fazia M, Iezzi A et al (2004) A polymorphism in the cyclooxygenase 2 gene as an inherited protective factor against myocardial infarction and stroke. JAMA 291:2221–2228. doi: 10.1001/jama.291.18.2221
Clarke R, Xu P, Bennett D, Lewington S, Zondervan K et al (2006) Lymphotoxin-alpha gene and risk of myocardial infarction in 6,928 cases and 2,712 controls in the ISIS case–control study. PLoS Genet 2:e107. doi: 10.1371/journal.pgen.0020107
Cohen JC, Boerwinkle E, Mosley TH Jr, Hobbs HH (2006) Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N Engl J Med 354:1264–1272. doi:10.1056/NEJMoa054013
Croft SA, Samani NJ, Teare MD, Hampton KK, Steeds RP et al (2001) Novel platelet membrane glycoprotein VI dimorphism is a risk factor for myocardial infarction. Circulation 104:1459–1463. doi:10.1161/hc3801.096397
CRP CHD Genetics Collaboration (2008) Collaborative pooled analysis of data on C-reactive protein gene variants and coronary disease: judging causality by Mendelian randomisation. Eur J Epidemiol 23:531–540. doi: 10.1007/s10654-008-9249-z
Damani SB, Topol EJ (2007) Future use of genomics in coronary artery disease. J Am Coll Cardiol 50:1933–1940. doi:10.1016/j.jacc.2007.07.062
Davey Smith G, Ebrahim S (2003) “Mendelian randomization”: Can genetic epidemiology contribute to understanding environmental causes of disease? Int J Epidemiol 32:1–22. doi:10.1093/ije/dyg070
Davey Smith G, Ebrahim S (2005) What can Mendelian randomization tell us about modifiable behavioural and environmental exposures? BMJ 330:1076–1079. doi:10.1136/bmj.330.7499.1076
Davignon J, Gregg RE, Sing CF (1988) Apolipoprotein E polymorphism and atherosclerosis. Arteriosclerosis 8:1–21
Deloughery TG, Evans A, Sadeghi A, McWilliams J, Henner WD et al (1996) Common mutation in methylenetetrahydrofolate reductase. Correlation with homocysteine metabolism and late-onset vascular disease. Circulation 94:3074–3078
Dugi KA, Brandauer K, Schmidt N, Nau B, Schneider JG et al (2001) Low hepatic lipase activity is a novel risk factor for coronary artery disease. Circulation 104:3057–3062. doi:10.1161/hc5001.100795
Dumont J, Zureik M, Cottel D, Montaye M, Ducimetiere P et al (2007) Association of arginase 1 gene polymorphisms with the risk of myocardial infarction and common carotid intima-media thickness. J Med Genet 44:526–553. doi:10.1136/jmg.2006.047449
Dwyer JH, Allayee H, Dwyer KM, Fan J, Wu H et al (2004) Arachidonate 5-lipoxygenase promoter genotype, dietary arachidonic acid, and atherosclerosis. N Engl J Med 350:29–37. doi:10.1056/NEJMoa025079
Edfeldt K, Bennet AM, Eriksson P, Frostegard J, Wiman B et al (2004) Association of hypo-responsive toll-like receptor 4 variants with risk of myocardial infarction. Eur Heart J 25:1447–1453. doi:10.1016/j.ehj.2004.05.004
Ehnholm C, Lukka M, Kuusi T, Nikkilä E, Utermann G (1986) Apolipoprotein E polymorphism in the Finnish population: gene frequencies and relation to lipoprotein concentrations. J Lipid Res 27:227–235
Eichner JE, Dunn ST, Perveen G, Thompson DM, Stewart KE et al (2002) Apolipoprotein E polymorphism and cardiovascular disease: a HuGE review. Am J Epidemiol 155:487–495. doi:10.1093/aje/155.6.487
Elrayess MA, Webb KE, Bellingan GJ, Whittall RA, Kabir J et al (2004) R643G polymorphism in PECAM–1 influences transendothelial migration of monocytes and is associated with progression of CHD and CHD events. Atherosclerosis 177:127–135. doi:10.1016/j.atherosclerosis.2004.06.009
Endler G, Mannhalter C, Sunder-Plassmann H, Lalouschek W, Kapiotis S et al (2001) Homozygosity for the C→T polymorphism at nucleotide 46 in the 5’ untranslated region of the factor XII gene protects from development of acute coronary syndrome. Br J Haematol 115:1007–1009. doi:10.1046/j.1365-2141.2001.03201.x
Eriksson P, Kallin B, van’t Hooft FM, Bavenholm P, Hamsten A (1995) Allele-specific increase in basal transcription of the plasminogen-activator inhibitor 1 gene is associated with myocardial infarction. Proc Natl Acad Sci USA 92:1851–1855. doi:10.1073/pnas.92.6.1851
Eriksson AL, Skrtic S, Niklason A, Hulten LM, Wiklund O et al (2004) Association between the low activity genotype of catechol-O-methyltransferase and myocardial infarction in a hypertensive population. Eur Heart J 25:386–391. doi:10.1016/j.ehj.2003.12.026
Farrall M, Green FR, Peden JF, Olsson PG, Clarke R et al (2006) Genome-wide mapping of susceptibility to coronary artery disease identifies a novel replicated locus on chromosome 17. PLoS Genet 2:755–761. doi:10.1371/journal.pgen.0020072
Folsom AR, Nieto FJ, McGovern PG, Tsai MY, Malinow MR et al (1998) Prospective study of coronary heart disease incidence in relation to fasting total homocysteine, related genetic polymorphisms, and B vitamins: the Atherosclerosis Risk in Communities (ARIC) Study. Circulation 98:204–210
Francke S, Manraj M, Lacquemant C, Lecoeur C, Leprêtre F et al (2001) A genome-wide scan for coronary heart disease suggests in Indo-Mauritians a susceptibility locus on chromosome 16p13 and replicates linkage with the metabolic syndrome on 3q27. Hum Mol Genet 10:2751–2765. doi:10.1093/hmg/10.24.2751
Fujimoto H, Taguchi JI, Imai Y, Ayabe S, Hashimoto H et al (2008) Manganese superoxide dismutase polymorphism affects the oxidized low-density lipoprotein-induced apoptosis of macrophages and coronary artery disease. Eur Heart J 29:1267–1274. doi:10.1093/eurheartj/ehm500
Fumeron F, Betoulle D, Nicaud V, Evans A, Kee F et al (2002) Serotonin transporter gene polymorphism and myocardial infarction: Etude Cas-Temoins de l’Infarctus du Myocarde (ECTIM). Circulation 105:2943–2945. doi:10.1161/01.CIR.0000022603.92986.99
Gallagher PM, Meleady R, Shields DC, Tan KS, McMaster D et al (1996) Homocysteine and risk of premature coronary heart disease. Evidence for a common gene mutation. Circulation 94:2154–2158
Georges JL, Loukaci V, Poirier O, Evans A, Luc G et al (2001) Interleukin-6 gene polymorphisms and susceptibility to myocardial infarction: the ECTIM study. J Mol Med 79:300–305. doi:10.1007/s001090100209
Georgopoulos A, Bloomfield H, Collins D, Brousseau ME, Ordovas JM et al (2007) Codon 54 polymorphism of the fatty acid binding protein (FABP) 2 gene is associated with increased cardiovascular risk in the dyslipidemic diabetic participants of the Veterans Affairs HDL Intervention Trial (VA-HIT). Atherosclerosis 194:169–174. doi:10.1016/j.atherosclerosis.2006.07.022
Gonzalez P, Alvarez R, Batalla A, Reguero JR, Alvarez V et al (2001) Genetic variation at the chemokine receptors CCR5/CCR2 in myocardial infarction. Genes Immun 2:191–195. doi:10.1038/sj.gene.6363760
Gonzalez-Conejero R, Corral J, Roldan V, Martinez C, Marin F et al (2002) A common polymorphism in the annexin V Kozak sequence (-1C>T) increases translation efficiency and plasma levels of annexin V, and decreases the risk of myocardial infarction in young patients. Blood 100:2081–2086
Groenemeijer BE, Hallman MD, Reymer PW, Gagné E, Kuivenhoven JA et al (1997) Genetic variant showing a positive interaction with β-blocking agents with a beneficial influence on lipoprotein lipase activity, HDL cholesterol, and triglyceride levels in coronary artery disease patients. The Ser447-stop substitution in the lipoprotein lipase gene. Circulation 95:2628–2635.
Gruchala M, Ciecwierz D, Wasag B, Targonski R, Dubaniewicz W et al (2003) Association of the ScaI atrial natriuretic peptide gene polymorphism with nonfatal myocardial infarction and extent of coronary artery disease. Am Heart J 145:125–131. doi:10.1067/mhj.2003.52
Hakonarson H, Thorvaldsson S, Helgadottir A, Gudbjartsson D, Zink F et al (2005) Effects of a 5-lipoxygenase-activating protein inhibitor on biomarkers associated with risk of myocardial infarction. JAMA 293:2245–2256. doi:10.1001/jama.293.18.2245
Hallman DM, Boerwinkle E, Saha N, Sandholzer C, Menzel HJ et al (1991) The apolipoprotein E polymorphism: a comparison of allele frequencies and effects in nine populations. Am J Hum Genet 49:338–349
Harrap SB, Zammit KS, Wong ZY, Williams FM, Bahlo M et al (2002) Genome-wide linkage analysis of the acute coronary syndrome suggests a locus on chromosome 2. Arterioscler Thromb Vasc Biol 22:874–878. doi:10.1161/01.ATV.0000016258.40568.F1
Hauser ER, Crossman DC, Granger CB, Haines JL, Jones CJ et al (2004) A genomewide scan for early-onset coronary artery disease in 438 families: the GENECARD Study. Am J Hum Genet 75:436–447. doi:10.1086/423900
Hegele RA, Huang LS, Herbert PN, Blum CB, Buring JE et al (1986) Apolipoprotein B-gene DNA polymorphisms associated with myocardial infarction. N Engl J Med 315:1509–1515
Helgadottir A, Manolescu A, Thorleifsson G, Gretarsdottir S, Jonsdottir H et al (2004) The gene encoding 5-lipoxygenase activating protein confers risk of myocardial infarction and stroke. Nat Genet 36:233–239. doi:10.1038/ng1311
Helgadottir A, Gretarsdottir S, St Clair D, Manolescu A, Cheung J et al (2005) Association between the gene encoding 5-lipoxygenase-activating protein and stroke replicated in a Scottish population. Am J Hum Genet 76:505–509. doi:10.1086/428066
Helgadottir A, Manolescu A, Helgason A, Thorleifsson G, Thorsteinsdottir U et al (2006) A variant of the gene encoding leukotriene A4 hydrolase confers ethnicity-specific risk of myocardial infarction. Nat Genet 38:68–74. doi:10.1038/ng1692
Helgadottir A, Thorleifsson G, Manolescu A, Gretarsdottir S, Blondal T et al (2007) A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science 316:1491–1493. doi:10.1126/science.1142842
Helgadottir A, Thorleifsson G, Magnusson KP, Grétarsdottir S, Steinthorsdottir V et al (2008) The same sequence variant on 9p21 associates with myocardial infarction, abdominal aortic aneurysm and intracranial aneurysm. Nat Genet 40:217–224. doi:10.1038/ng.72
Henderson HE, Kastelein JJ, Zwinderman AH, Gagné E, Jukema JW et al (1999) Lipoprotein lipase activity is decreased in a large cohort of patients with coronary artery disease and is associated with changes in lipids and lipoproteins. J Lipid Res 40:735–743
Hirashiki A, Yamada Y, Murase Y, Suzuki Y, Kataoka H et al (2003) Association of gene polymorphisms with coronary artery disease in low- or high-risk subjects defined by conventional risk factors. J Am Coll Cardiol 42:1429–1437. doi:10.1016/S0735-1097(03)01062-3
Holmer SR, Hengstenberg C, Kraft HG, Mayer B, Poll M et al (2003) Association of polymorphisms of the apolipoprotein(a) gene with lipoprotein(a) levels and myocardial infarction. Circulation 107:696–701. doi:10.1161/01.CIR.0000048125.79640.77
Howell WM, Ali S, Rose-Zerilli MJ, Ye S (2005) VEGF polymorphisms and severity of atherosclerosis. J Med Genet 42:485–490. doi:10.1136/jmg.2004.025734
Hubacek JA, Rothe G, Pit’ha J, Skodova Z, Stanek V et al (1999) C(–260)→T polymorphism in the promoter of the CD14 monocyte receptor gene as a risk factor for myocardial infarction. Circulation 99:3218–3220
Humphries SE, Nicaud V, Margalef J, Tiret L, Talmud PJ (1998) Lipoprotein lipase gene variation is associated with a paternal history of premature coronary artery disease and fasting and postprandial plasma triglycerides: the European Atherosclerosis Research Study (EARS). Arterioscler Thromb Vasc Biol 18:526–534
Humphries SE, Cooper JA, Talmud PJ, Miller GJ (2007) Candidate gene genotypes, along with conventional risk factor assessment, improve estimation of coronary heart disease risk in healthy UK men. Clin Chem 53:8–16. doi:10.1373/clinchem.2006.074591
Iacoviello L, Di Castelnuovo A, De Knijff P, D’Orazio A, Amore C et al (1998) Polymorphisms in the coagulation factor VII gene and the risk of myocardial infarction. N Engl J Med 338:79–85. doi:10.1056/NEJM199801083380202
Iacoviello L, Di Castelnuovo A, Gattone M, Pezzini A, Assanelli D et al (2005) Polymorphisms of the interleukin-1β gene affect the risk of myocardial infarction and ischemic stroke at young age and the response of mononuclear cells to stimulation in vitro. Arterioscler Thromb Vasc Biol 25:222–227
Iakoubova OA, Tong CH, Chokkalingam AP, Rowland CM, Kirchgessner TG et al (2006) Asp92Asn polymorphism in the myeloid IgA Fc receptor is associated with myocardial infarction in two disparate populations: CARE and WOSCOPS. Arterioscler Thromb Vasc Biol 26:2763–2768. doi:10.1161/01.ATV.0000247248.76409.8b
Iakoubova OA, Tong CH, Rowland CM, Kirchgessner TG, Young BA et al (2008) Association of the Trp719Arg polymorphism in kinesin-like protein 6 with myocardial infarction and coronary heart disease in 2 prospective trials: the CARE and WOSCOPS trials. J Am Coll Cardiol 51:435–443. doi:10.1016/j.jacc.2007.05.057
Inoue N, Kawashima S, Kanazawa K, Yamada S, Akita H (1998) Polymorphism of the NADH/NADPH oxidase p22 phox gene in patients with coronary artery disease. Circulation 97:135–137
International Human Genome Sequencing Consortium (2004) Finishing the euchromatic sequence of the human genome. Nature 431:931–945. doi: 10.1038/nature03001
Iwai C, Akita H, Kanazawa K, Shiga N, Terashima M et al (2003) Arg389Gly polymorphism of the human β1-adrenergic receptor in patients with nonfatal acute myocardial infarction. Am Heart J 146:106–109. doi:10.1016/S0002-8703(03)00110-8
Jemaa R, Fumeron F, Poirier O, Lecerf L, Evans A et al (1995) Lipoprotein lipase gene polymorphisms: associations with myocardial infarction and lipoprotein levels, the ECTIM study. Etude Cas Temoin sur l’Infarctus du Myocarde. J Lipid Res 36:2141–2146
Juhan-Vague I, Morange PE, Aubert H, Henry M, Aillaud MF et al (2002) Plasma thrombin-activatable fibrinolysis inhibitor antigen concentration and genotype in relation to myocardial infarction in the north and south of Europe. Arterioscler Thromb Vasc Biol 22:867–873. doi:10.1161/01.ATV.0000015445.22243.F4
Kardys I, Klaver CC, Despriet DD, Bergen AA, Uitterlinden AG et al (2006) A common polymorphism in the complement factor H gene is associated with increased risk of myocardial infarction: the Rotterdam Study. J Am Coll Cardiol 47:1568–1575. doi:10.1016/j.jacc.2005.11.076
Kastelein JJ, Jukema JW, Zwinderman AH, Clee S, van Boven AJ et al (2000) Lipoprotein lipase activity is associated with severity of angina pectoris. Circulation 102:1629–1633
Katsuya T, Koike G, Yee TW, Sharpe N, Jackson R et al (1995) Association of angiotensinogen gene T235 variant with increased risk of coronary heart disease. Lancet 345:1600–1603. doi:10.1016/S0140-6736(95)90115-9
Keavney B (2002) Genetic epidemiological studies of coronary heart disease. Int J Epidemiol 31:730–736. doi:10.1093/ije/31.4.730
Klerk M, Verhoef P, Clarke R, Klerk M, Verhoef P et al (2002) MTHFR 677C→T polymorphism and risk of coronary heart disease: a meta-analysis. JAMA 288:2023–2031. doi:10.1001/jama.288.16.2023
Kluijtmans LA, van den Heuvel LP, Boers GH, Frosst P, Stevens EM et al (1996) Molecular genetic analysis in mild hyperhomocysteinemia: a common mutation in the methylenetetrahydrofolate reductase gene is a genetic risk factor for cardiovascular disease. Am J Hum Genet 58:35–41
Kohler HP, Stickland MH, Ossei-Gerning N, Carter A, Mikkola H et al (1998) Association of a common polymorphism in the factor XIII gene with myocardial infarction. Thromb Haemost 79:8–13
Koide S, Kugiyama K, Sugiyama S, Nakamura S, Fukushima H et al (2003) Association of polymorphism in glutamate-cysteine ligase catalytic subunit gene with coronary vasomotor dysfunction and myocardial infarction. J Am Coll Cardiol 41:539–545. doi:10.1016/S0735-1097(02)02866-8
Kuivenhoven JA, Groenemeyer BE, Boer JM, Reymer PW, Berghuis R et al (1997) Ser447stop mutation in lipoprotein lipase is associated with elevated HDL cholesterol levels in normolipidemic males. Arterioscler Thromb Vasc Biol 17:595–599
Kuivenhoven JA, Jukema JW, Zwinderman AH, de Knijff P, McPherson R et al (1998) The role of a common variant of the cholesterol ester transfer protein gene in the progression of coronary atherosclerosis. N Engl J Med 338:86–93. doi:10.1056/NEJM199801083380203
Kullo IJ, Ding K (2007) Mechanisms of disease: the genetic basis of coronary heart disease. Nat Clin Pract Cardiovasc Med 4:558–569. doi:10.1038/ncpcardio0982
Kumamoto T, Kawai Y, Arakawa K, Morikawa N, Kuribara J et al (2006) Association of Gln222Arg polymorphism in the deoxyribonuclease I (DNase I) gene with myocardial infarction in Japanese patients. Eur Heart J 27:2081–2087. doi:10.1093/eurheartj/ehl177
Ladenvall P, Johansson L, Jansson JH, Jern S, Nilsson TK et al (2002) Tissue-type plasminogen activator-7, 351C/T enhancer polymorphism is associated with a first myocardial infarction. Thromb Haemost 87:105–109
Lahoz C, Schaefer EJ, Cupples LA, Wilson PW, Levy D et al (2001) Apolipoprotein E genotype and cardiovascular disease in the Framingham Heart Study. Atherosclerosis 154:529–537. doi:10.1016/S0021-9150(00)00570-0
Lambert JC, Brousseau T, Defosse V, Evans A, Arveiler D et al (2000) Independent association of an APOE gene promoter polymorphism with increased risk of myocardial infarction and decreased APOE plasma concentrations—the ECTIM study. Hum Mol Genet 9:57–61. doi:10.1093/hmg/9.1.57
Lange LA, Carlson CS, Hindorff LA, Lange EM, Walston J et al (2006) Association of polymorphisms in the CRP gene with circulating C-reactive protein levels and cardiovascular events. JAMA 296:2703–2711. doi:10.1001/jama.296.22.2703
Lavergne E, Labreuche J, Daoudi M, Debré P, Cambien F et al (2005) Adverse associations between CX3CR1 polymorphisms and risk of cardiovascular or cerebrovascular disease. Arterioscler Thromb Vasc Biol 25:847–853. doi:10.1161/01.ATV.0000157150.23641.36
Ledmyr H, McMahon AD, Ehrenborg E, Nielsen LB, Neville M et al (2004) The microsomal triglyceride transfer protein gene –493T variant lowers cholesterol but increases the risk of coronary heart disease. Circulation 109:2279–2284. doi: 10.1161/01.CIR.0000130070.96758.7b
Lewis SJ, Ebrahim S, Davy Smith G (2005) Meta-analysis of MTHFR 677C→T polymorphism and coronary heart disease: Does totality of evidence support causal role for homocysteine and preventive potential of folate? BMJ 331:1053–1056. doi:10.1136/bmj.38611.658947.55
Liu PY, Li YH, Chao TH, Wu HL, Lin LJ et al (2007) Synergistic effect of cytochrome P450 epoxygenase CYP2J2*7 polymorphism with smoking on the onset of premature myocardial infarction. Atherosclerosis 195:199–206. doi:10.1016/j.atherosclerosis.2006.11.001
Lundberg GA, Kellin A, Samnegard A, Lundman P, Tornvall P et al (2005) Severity of coronary artery stenosis is associated with a polymorphism in the CXCL16/SR-PSOX gene. J Intern Med 257:415–422. doi:10.1111/j.1365-2796.2005.01469.x
Ma J, Stampfer MJ, Hennekens CH, Frosst P, Selhub J et al (1996) Methylenetetrahydrofolate reductase polymorphism, plasma folate, homocysteine, and risk of myocardial infarction in US physicians. Circulation 94:2410–2416
Mager A, Lalezari S, Shohat T, Birnbaum Y, Adler Y et al (1999) Methylenetetrahydrofolate reductase genotypes and early-onset coronary artery disease. Circulation 100:2406–2410
Mango R, Biocca S, del Vecchio F, Clementi F, Sangiuolo F et al (2005) In vivo and in vitro studies support that a new splicing isoform of OLR1 gene is protective against acute myocardial infarction. Circ Res 97:152–158. doi:10.1161/01.RES.0000174563.62625.8e
Marroquin OC, Selzer F, Mulukutla SR, Williams DO, Vlachos HA et al (2008) A comparison of bare-metal and drug-eluting stents for off-label indications. N Engl J Med 358:342–352. doi:10.1056/NEJMoa0706258
Marz W, Seelhorst U, Wellnitz B, Tiran B, Obermayer-Pietsch B et al (2007) Alanine to serine polymorphism at position 986 of the calcium-sensing receptor associated with coronary heart disease, myocardial infarction, all-cause and cardiovascular mortality. J Clin Endocrinol Metab 92:2363–2369. doi:10.1210/jc.2006-0071
Mattick JS, Makunin IV (2006) Non-coding RNA. Hum Mol Genet 15:R17–R29. doi:10.1093/hmg/ddl046
McDermott DH, Yang Q, Kathiresan S, Cupples LA, Massaro JM et al (2005) CCL2 polymorphisms are associated with serum monocyte chemoattractant protein-1 levels and myocardial infarction in the Framingham Heart Study. Circulation 112:1113–1120. doi:10.1161/CIRCULATIONAHA.105.543579
McPherson R, Pertsemlidis A, Kavaslar N, Stewart A, Roberts R et al (2007) A common allele on chromosome 9 associated with coronary heart disease. Science 316:1488–1491. doi:10.1126/science.1142447
Mehrabian M, Allayee H, Wong J, Shih W, Wang X-P et al (2002) Identification of 5-lipoxygenase as a major gene contributing to atherosclerosis susceptibility in mice. Circ Res 91:120–126. doi:10.1161/01.RES.0000028008.99774.7F
Morita H, Taguchi J, Kurihara H, Kitaoka M, Kaneda H et al (1997) Genetic polymorphism of 5, 10-methylenetetrahydrofolate reductase (MTHFR) as a risk factor for coronary artery disease. Circulation 95:2032–2036
Moshfegh K, Wuillemin WA, Redondo M, Lammle B, Beer JH et al (1999) Association of two silent polymorphisms of platelet glycoprotein Ia/IIa receptor with risk of myocardial infarction: a case–control study. Lancet 353:351–354. doi:10.1016/S0140-6736(98)06448-4
Muckian C, Fitzgerald A, O’Neill A, O’Byrne A, Fitzgerald DJ et al (2002) Genetic variability in the extracellular matrix as a determinant of cardiovascular risk: association of type III collagen COL3A1 polymorphisms with coronary artery disease. Blood 100:1220–1223. doi:10.1182/blood-2002-01-0283
Murata M, Matsubara Y, Kawano K, Zama T, Aoki N et al (1997) Coronary artery disease and polymorphisms in a receptor mediating shear stress-dependent platelet activation. Circulation 96:3281–3286
Naber CK, Husing J, Wolfhard U, Erbel R, Siffert W (2000) Interaction of the ACE D allele and the GNB3 825T allele in myocardial infarction. Hypertension 36:986–989
Nakamura S, Kugiyama K, Sugiyama S, Miyamoto S, Koide S et al (2002) Polymorphism in the 5’-flanking region of human glutamate-cysteine ligase modifier subunit gene is associated with myocardial infarction. Circulation 105:2968–2973. doi:10.1161/01.CIR.0000019739.66514.1E
Nakayama T, Soma M, Saito S, Honye J, Yajima J et al (2002) Association of a novel single nucleotide polymorphism of the prostacyclin synthase gene with myocardial infarction. Am Heart J 143:797–801. doi:10.1067/mhj.2002.122171
Ohashi K, Ouchi N, Kihara S, Funahashi T, Nakamura T et al (2004) Adiponectin I164T mutation is associated with the metabolic syndrome and coronary artery disease. J Am Coll Cardiol 43:1195–1200. doi:10.1016/j.jacc.2003.10.049
Olivieri O, Stranieri C, Bassi A, Zaia B, Girelli D et al (2002) ApoC-III gene polymorphisms and risk of coronary artery disease. J Lipid Res 43:1450–1457. doi:10.1194/jlr.M200145-JLR200
Ono K, Goto Y, Takagi S, Baba S, Tago N et al (2004) A promoter variant of the heme oxygenase-1 gene may reduce the incidence of ischemic heart disease in Japanese. Atherosclerosis 173:315–319. doi:10.1016/j.atherosclerosis.2003.11.021
Ortlepp JR, Vesper K, Mevissen V, Schmitz F, Janssens U et al (2003) Chemokine receptor (CCR2) genotype is associated with myocardial infarction and heart failure in patients under 65 years of age. J Mol Med 81:363–367. doi:10.1007/s00109-003-0471-6
Ott I, Koch W, von Beckerath N, de Waha R, Malawaniec A et al (2004) Tissue factor promotor polymorphism -603 A/G is associated with myocardial infarction. Atherosclerosis 177:189–191. doi:10.1016/j.atherosclerosis.2004.07.006
Ozaki K, Ohnishi Y, Iida A, Sekine A, Yamada R et al (2002) Functional SNPs in the lymphotoxin-α gene that are associated with susceptibility to myocardial infarction. Nat Genet 32:650–654. doi:10.1038/ng1047
Ozaki K, Inoue K, Sato H, Iida A, Ohnishi Y et al (2004) Functional variation in LGALS2 confers risk of myocardial infarction and regulates lymphotoxin-α secretion in vitro. Nature 429:72–75. doi:10.1038/nature02502
Ozaki K, Sato H, Iida A, Mizuno H, Nakamura T et al (2006) A functional SNP in PSMA6 confers risk of myocardial infarction in the Japanese population. Nat Genet 38:921–925. doi:10.1038/ng1846
Pai JK, Mukamal KJ, Rexrode KM, Rimm EB (2008) C-reactive protein (CRP) gene polymorphisms, CRP levels, and risk of incident coronary heart disease in two nested case–control studies. PLoS One 3:e1395. doi: 10.1371/journal.pone.0001395
Pajukanta P, Cargill M, Viitanen L, Nuotio I, Kareinen A et al (2000) Two loci on chromosomes 2 and X for premature coronary heart disease identified in early- and late-settlement populations of Finland. Am J Hum Genet 67:1481–1493. doi:10.1086/316902
Pasmant E, Laurendeau I, Heron D, Vidaud M, Vidaud D et al (2007) Characterization of a germ-line deletion, including the entire INK4/ARF locus, in a melanoma-neural system tumor family: identification of ANRIL, an antisense noncoding RNA whose expression coclusters with ARF. Cancer Res 67:3963–3969. doi:10.1158/0008-5472.CAN-06-2004
Pearce E, Tregouet DA, Samnegard A, Morgan AR, Cox C et al (2005) Haplotype effect of the matrix metalloproteinase-1 gene on risk of myocardial infarction. Circ Res 97:1070–1076. doi:10.1161/01.RES.0000189302.03303.11
Pearson TA, Manolio TA (2008) How to interpret a genome-wide association study. JAMA 299:1335–1344. doi:10.1001/jama.299.11.1335
Podgoreanu MV, White WD, Morris RW, Mathew JP, Stafford-Smith M et al (2006) Inflammatory gene polymorphisms and risk of postoperative myocardial infarction after cardiac surgery. Circulation 114:I275–I281. doi: 10.1161/CIRCULATIONAHA.105.001032
Ridker PM, Cook NR, Cheng S, Erlich HA, Lindpaintner K et al (2003) Alanine for proline substitution in the peroxisome proliferator-activated receptor gamma-2 (PPARG2) gene and the risk of incident myocardial infarction. Arterioscler Thromb Vasc Biol 23:859–863. doi:10.1161/01.ATV.0000068680.19521.34
Rosamond W, Flegal K, Friday G, Furie K, Go A et al (2007) Heart disease and stroke statistics—2007 update: a report from the American Heart Association Statistics Committee and Stroke Statistics Subcommittee. Circulation 115:e69–e171. doi:10.1161/CIRCULATIONAHA.106.179918
Sabatine MS, Ploughman L, Simonsen KL, Iakoubova OA, Kirchgessner TG et al (2008) Association between ADAMTS1 matrix metalloproteinase gene variation, coronary heart disease, and benefit of statin therapy. Arterioscler Thromb Vasc Biol 28:562–567. doi:10.1161/ATVBAHA.107.156653
Sala G, Di Castelnuovo A, Cuomo L, Gattone M, Giannuzzi P et al (2001) The E27 β2-adrenergic receptor polymorphism reduces the risk of myocardial infarction in dyslipidemic young males. Thromb Haemost 85:231–233
Samani NJ, Erdmann J, Hall AS, Hengstenberg C, Mangino M et al (2007) Genomewide association analysis of coronary artery disease. N Engl J Med 357:443–453. doi: 10.1056/NEJMoa072366
Saxena R, Voight BF, Lyssenko V, Burtt NP, de Bakker PI et al (2007) Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 316:1331–1336. doi:10.1126/science.1142358
Schaefer EJ, Lamon-Fava S, Johnson S, Ordovas JM, Schaefer MM et al (1994) Effects of gender and menopausal status on the association of apolipoprotein E phenotype with plasma lipoprotein levels: results from the Framingham Offspring Study. Arterioscler Thromb Vasc Biol 14:1105–1113
Schunkert H, Götz A, Braund P, McGinnis R, Tregouet DA et al (2008) Repeated replication and a prospective meta-analysis of the association between chromosome 9p21.3 and coronary artery disease. Circulation 117:1675–1684. doi: 10.1161/CIRCULATIONAHA.107.730614
Schwartz SM, Siscovick DS, Malinow MR, Rosendaal FR, Beverly RK et al (1997) Myocardial infarction in young women in relation to plasma total homocysteine, folate, and a common variant in the methylenetetrahydrofolate reductase gene. Circulation 96:412–417
Scott LJ, Mohlke KL, Bonnycastle LL, Willer CJ, Li Y et al (2007) A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science 316:1341–1345. doi:10.1126/science.1142382
Senti M, Fernandez-Fernandez JM, Tomas M, Vazquez E, Elosua R et al (2005) Protective effect of the KCNMB1 E65K genetic polymorphism against diastolic hypertension in aging women and its relevance to cardiovascular risk. Circ Res 97:1360–1365. doi:10.1161/01.RES.0000196557.93717.95
Serrato M, Marian AJ (1995) A variant of human paraoxonase/arylesterase (HUMPONA) gene is a risk factor for coronary artery disease. J Clin Invest 96:3005–3008. doi:10.1172/JCI118373
Shearman AM, Cupples LA, Demissie S, Peter I, Schmid CH et al (2003) Association between estrogen receptor α gene variation and cardiovascular disease. JAMA 290:2263–2270. doi:10.1001/jama.290.17.2263
Shen GQ, Li L, Girelli D, Seidelmann SB, Rao S et al (2007) An LRP8 variant is associated with familial and premature coronary artery disease and myocardial infarction. Am J Hum Genet 81:780–791. doi:10.1086/521581
Shen GQ, Li L, Rao S, Abdullah KG, Ban JM et al (2008a) Four SNPs on chromosome 9p21 in a South Korean population implicate a genetic locus that confers high cross-race risk for development of coronary artery disease. Arterioscler Thromb Vasc Biol 28:360–365. doi:10.1161/ATVBAHA.107.157248
Shen GQ, Rao S, Martinelli N, Li L, Olivieri O et al (2008b) Association between four SNPs on chromosome 9p21 and myocardial infarction is replicated in an Italian population. J Hum Genet 53:144–150. doi:10.1007/s10038-007-0230-6
Shiffman D, Ellis SG, Rowland CM, Malloy MJ, Luke MM et al (2005) Identification of four gene variants associated with myocardial infarction. Am J Hum Genet 77:596–605. doi:10.1086/491674
Shiffman D, Rowland CM, Louie JZ, Luke MM, Bare LA et al (2006) Gene variants of VAMP8 and HNRPUL1 are associated with early-onset myocardial infarction. Arterioscler Thromb Vasc Biol 26:1613–1618. doi:10.1161/01.ATV.0000226543.77214.e4
Shiffman D, O’Meara ES, Bare LA, Rowland CM, Louie JZ et al (2008) Association of gene variants with incident myocardial infarction in the Cardiovascular Health Study. Arterioscler Thromb Vasc Biol 28:173–179. doi:10.1161/ATVBAHA.107.153981
Shimasaki Y, Yasue H, Yoshimura M, Nakayama M, Kugiyama K et al (1998) Association of the missense Glu298Asp variant of the endothelial nitric oxide synthase gene with myocardial infarction. J Am Coll Cardiol 31:1506–1510. doi:10.1016/S0735-1097(98)00167-3
Sing CF, Davignon J (1985) Role of apolipoprotein E genetic polymorphism in determining normal plasma lipid and lipoprotein variation. Am J Hum Genet 37:268–285
Song Y, Stampfer MJ, Liu S (2004) Meta-analysis: apolipoprotein E genotypes and risk for coronary heart disease. Ann Intern Med 141:137–147
Spanbroek R, Grabner R, Lotzer K, Hildner M, Urbach A et al (2003) Expanding expression of the 5-lipoxygenase pathway within the arterial wall during human atherogenesis. Proc Natl Acad Sci USA 100:1238–1243. doi:10.1073/pnas.242716099
Swanberg M, Lidman O, Padyukov L, Eriksson P, Akesson E et al (2005) MHC2TA is associated with differential MHC molecule expression and susceptibility to rheumatoid arthritis, multiple sclerosis and myocardial infarction. Nat Genet 37:486–494. doi:10.1038/ng1544
Talmud PJ, Martin S, Taskinen MR, Frick MH, Nieminen MS et al (2004) APOA5 gene variants, lipoprotein particle distribution, and progression of coronary heart disease: results from the LOCAT study. J Lipid Res 45:750–756. doi:10.1194/jlr.M300458-JLR200
Talmud PJ, Cooper JA, Palmen J, Lovering R, Drenos F et al (2008) Chromosome 9p21.3 coronary heart disease locus genotype and prospective risk of CHD in healthy middle-aged men. Clin Chem 54:467–474. doi: 10.1373/clinchem.2007.095489
The BHF Family Heart Study Research Group (2005) A genomewide linkage study of 1,933 families affected by premature coronary artery disease: The British Heart Foundation (BHF) Family Heart Study. Am J Hum Genet 77:1011–1020. doi:10.1086/498653
The International HapMap Consortium (2007) A second generation human haplotype map of over 3.1 million SNPs. Nature 449:851–861. doi:10.1038/nature06258
Tiret L, Bonnardeaux A, Poirier O, Ricard S, Marques-Vidal P et al (1994) Synergistic effects of angiotensin-converting enzyme and angiotensin-II type 1 receptor gene polymorphisms on risk of myocardial infarction. Lancet 344:910–913. doi:10.1016/S0140-6736(94)92268-3
Topol EJ, McCarthy J, Gabriel S, Moliterno DJ, Rogers WJ et al (2001) Single nucleotide polymorphisms in multiple novel thrombospondin genes may be associated with familial premature myocardial infarction. Circulation 104:2641–2644. doi:10.1161/hc4701.100910
Topol EJ, Smith J, Plow EF, Wang QK (2006) Genetic susceptibility to myocardial infarction and coronary artery disease. Hum Mol Genet 15:R117–R123. doi:10.1093/hmg/ddl183
Tregouet DA, Barbaux S, Escolano S, Tahri N, Golmard JL et al (2002) Specific haplotypes of the P-selectin gene are associated with myocardial infarction. Hum Mol Genet 11:2015–2023. doi:10.1093/hmg/11.17.2015
Tregouet DA, Ricard S, Nicaud V, Arnould I, Soubigou S et al (2004) In-depth haplotype analysis of ABCA1 gene polymorphisms in relation to plasma ApoA1 levels and myocardial infarction. Arterioscler Thromb Vasc Biol 24:775–781. doi:10.1161/01.ATV.0000121573.29550.1a
van Bokxmeer FM, Mamotte CDS (1992) Apolipoprotein ε4 homozygosity in young men with coronary heart disease. Lancet 340:879–880. doi:10.1016/0140-6736(92)93288-X
Vasku A, Goldbergova M, Izakovicova Holla L, Siskova L, Groch L et al (2004) A haplotype constituted of four MMP-2 promoter polymorphisms (-1575G/A, -1306C/T, -790T/G and -735C/T) is associated with coronary triple-vessel disease. Matrix Biol 22:585–591. doi:10.1016/j.matbio.2003.10.004
Vendrell J, Fernandez-Real JM, Gutierrez C, Zamora A, Simon I et al (2003) A polymorphism in the promoter of the tumor necrosis factor-α gene (−308) is associated with coronary heart disease in type 2 diabetic patients. Atherosclerosis 167:257–264. doi:10.1016/S0021-9150(02)00429-X
Verhoef P, Rimm EB, Hunter DJ, Chen J, Willett WC et al (1998) A common mutation in the methylenetetrahydrofolate reductase gene and risk of coronary heart disease: results among U.S. men. J Am Coll Cardiol 32:353–359. doi: 10.1016/S0735-1097(98)00244-7
Wang L, Fan C, Topol SE, Topol EJ, Wang Q (2003) Mutation of MEF2A in an inherited disorder with features of coronary artery disease. Science 302:1578–1581. doi:10.1126/science.1088477
Wang Q, Rao S, Shen GQ, Li L, Moliterno DJ et al (2004) Premature myocardial infarction novel susceptibility locus on chromosome 1p34–36 identified by genomewide linkage analysis. Am J Hum Genet 74:262–271. doi:10.1086/381560
Wang X, Ria M, Kelmenson PM, Eriksson P, Higgins DC et al (2005) Positional identification of TNFSF4, encoding OX40 ligand, as a gene that influences atherosclerosis susceptibility. Nat Genet 37:365–372. doi:10.1038/ng1524
Wang Y, Zhang W, Zhang Y, Yang Y, Sun L et al (2006) VKORC1 haplotypes are associated with arterial vascular diseases (stroke, coronary heart disease, and aortic dissection). Circulation 113:1615–1621. doi:10.1161/CIRCULATIONAHA.105.580167
Webb KE, Martin JF, Hamsten A, Eriksson P, Iacoviello L et al (2001) Polymorphisms in the thrombopoietin gene are associated with risk of myocardial infarction at a young age. Atherosclerosis 154:703–711. doi:10.1016/S0021-9150(00)00633-X
Weintraub MS, Eisenberg S, Breslow JL (1987) Dietary fat clearance in normal subjects is regulated by genetic variation in apolipoprotein E. J Clin Invest 80:1571–1577. doi:10.1172/JCI113243
Weiss EJ, Bray PF, Tayback M, Schulman SP, Kickler TS et al (1996) A polymorphism of a platelet glycoprotein receptor as an inherited risk factor for coronary thrombosis. N Engl J Med 334:1090–1094. doi:10.1056/NEJM199604253341703
Wellcome Trust Case Control Consortium (2007) Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447:661–678. doi: 10.1038/nature05911
Wilson PW, Myers RH, Larson MG, Ordovas JM, Wolf PA et al (1994) Apolipoprotein E alleles, dyslipidemia, and coronary heart disease. The Framingham Offspring Study. JAMA 272:1666–1671. doi:10.1001/jama.272.21.1666
Wittrup HH, Tybjærg-Hansen A, Nordestgaard BG (1999) Lipoprotein lipase mutations, plasma lipids and lipoproteins, and risk of ischemic heart disease. A meta-analysis. Circulation 99:2901–2907
Wu KK, Aleksic N, Ahn C, Boerwinkle E, Folsom AR et al (2001) Thrombomodulin Ala455Val polymorphism and risk of coronary heart disease. Circulation 103:1386–1389. doi: 10.1161/hc2301.091931
Xhignesse M, Lussier-Cacan S, Sing CF, Kessling AM, Davignon J (1991) Influences of common variants of apolipoprotein E on measures of lipid metabolism in a sample selected for health. Arterioscler Thromb 11:1100–1110
Yamada Y (2006) Identification of genetic factors and development of genetic risk diagnosis systems for cardiovascular diseases and stroke. Circ J 70:1240–1248. doi:10.1253/circj.70.1240
Yamada Y, Ichihara S, Fujimura T, Yokota M (1998) Identification of the G994→T missense mutation in exon 9 of the plasma platelet-activating factor acetylhydrolase gene as an independent risk factor for coronary artery disease in Japanese men. Metabolism 47:177–181. doi:10.1016/S0026-0495(98)90216-5
Yamada Y, Izawa H, Ichihara S, Takatsu F, Ishihara H et al (2002) Prediction of the risk of myocardial infarction from polymorphisms in candidate genes. N Engl J Med 347:1916–1923. doi:10.1056/NEJMoa021445
Yamada Y, Matsuo H, Segawa T, Watanabe S, Kato K et al (2006) Assessment of genetic risk for myocardial infarction. Thromb Haemost 96:220–227
Yamada Y, Kato K, Oguri M, Fujimaki T, Yokoi K et al (2008) Genetic risk for myocardial infarction determined by polymorphisms of candidate genes in Japanese individuals. J Med Genet 45:216–221. doi:10.1136/jmg.2007.054387
Yang Y, Ruiz-Narvaez E, Niu T, Xu X, Campos H (2004) Genetic variants of the lipoprotein lipase gene and myocardial infarction in the Central Valley of Costa Rica. J Lipid Res 45:2106–2109. doi:10.1194/jlr.M400202-JLR200
Ye S, Watts GF, Mandalia S, Humphries SE, Henney AM (1995) Preliminary report: genetic variation in the human stromelysin promoter is associated with progression of coronary atherosclerosis. Br Heart J 73:209–215. doi:10.1136/hrt.73.3.209
Yokota M, Ichihara S, Lin TL, Nakashima N, Yamada Y (2000) Association of a T29→C polymorphism of the transforming growth factor-β1 gene with genetic susceptibility to myocardial infarction in Japanese. Circulation 101:2783–2787
Yoshida M, Takano Y, Sasaoka T, Izumi T, Kimura A (2003) E-selectin polymorphism associated with myocardial infarction causes enhanced leukocyte-endothelial interactions under flow conditions. Arterioscler Thromb Vasc Biol 23:783–788. doi:10.1161/01.ATV.0000067427.40133.59
Zee RY, Cook NR, Cheng S, Erlich HA, Lindpaintner K et al (2004) Threonine for alanine substitution in the eotaxin (CCL11) gene and the risk of incident myocardial infarction. Atherosclerosis 175:91–94. doi:10.1016/j.atherosclerosis.2004.01.042
Zeggini E, Weedon MN, Lindgren CM, Frayling TM, Elliott KS et al (2007) Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science 316:1336–1341. doi: 10.1126/science.1142364
Zhang H, Henderson H, Gagne SE, Clee SM, Miao L et al (1996) Common sequence variants of lipoprotein lipase: standardized studies of in vitro expression and catalytic function. Biochim Biophys Acta 1302:159–166
Zhang B, Ye S, Herrmann SM, Eriksson P, de Maat M et al (1999) Functional polymorphism in the regulatory region of gelatinase B gene in relation to severity of coronary atherosclerosis. Circulation 99:1788–1794
Zwicker JI, Peyvandi F, Palla R, Lombardi R, Canciani MT et al (2006) The thrombospondin-1 N700S polymorphism is associated with early myocardial infarction without altering von Willebrand factor multimer size. Blood 108:1280–1283. doi:10.1182/blood-2006-04-015701
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Yamada, Y., Ichihara, S. & Nishida, T. Molecular genetics of myocardial infarction. HUGO J 2, 7–22 (2008). https://doi.org/10.1007/s11568-008-9025-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11568-008-9025-x