Abstract
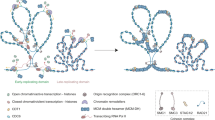
Current synthetic biology has witnessed a revolution that natural DNA molecule steps onto a broad scientific area by assembling a large variety of three-dimensional structures with the connectivity of polyhedra. A mathematical model of these biomolecules is crucial to clarify the biological self-assembly principle, and unravel a first-step understanding of biological regulation and controlling mechanisms. In this paper, mechanisms of two different enzymatic actions on DNA polyhedra are elucidated through theoretical models of polyhedral links: (1) topoisomerase that untangles DNA polyhedral links produces separated single-stranded DNA circles through the crossing change operation; (2) recombinase generates a class of polyhedral circular paths or polyhedral knots by applying the crossing smoothing operation. Furthermore, we also discuss the possibility of applying two theoretical operations in molecular design of DNA polyhedra. Thus, our research provides a new sight of how geometry and topology of DNA polyhedra can be manipulated and controlled by enzymes, as well as has implications for molecular design and structural analysis of structural genome organization.
Similar content being viewed by others
References
Adams, C. C. (1994). The knot book: an elementary introduction to the mathematical theory of knots. New York: Freeman.
Aldaye, F. A., Palmer, A. L., & Sleiman, H. F. (2008). Assembling materials with DNA as the guide. Science, 321, 1795–1799.
Andersen, F. F. et al. (2008). Assembly and structural analysis of a covalently closed nano-scale DNA cage. Nucleic Acids Res., 36, 1113–1119.
Angeleska, A., Jonoska, N., & Saito, M. (2009). DNA recombination through assembly graphs. Discrete Appl. Math., 157, 3020–3037.
Bates, A. D., & Maxwell, A. (2005). DNA topology (2nd ed.). Oxford: Oxford University Press.
Benham, C. J. et al. (2009). The IMA volumes in mathematics and its applications: Vol. 150. The mathematics of DNA structure, mechanics, and dynamics. New York: Springer.
Bhatia, D. et al. (2009). Icosahedral DNA nanocapsules by modular assembly. Angew. Chem., Int. Ed. Engl., 48, 1–5.
Buck, D., & Marcotte, C. V. (2005). Tangle solutions for a family of DNA-rearranging proteins. Math. Proc. Camb. Philos. Soc., 139, 59–80.
Buck, D., & Flapan, E. (2007a). Predicting knot or catenane type of site-specific recombination products. J. Mol. Biol., 374, 1186–1199.
Buck, D., & Flapan, E. (2007b). A topological characterization of knots and links arising from site-specific recombination. J. Phys. A, Math. Theor., 40, 12377–12395.
Buck, D., & Flapan, E. (2009). In Proceedings of symposia in applied mathematics: Vol. 66. Applications of knot theory. Providence: AMS.
Buck, D., & Marcotte, C. V. (2007). Classification of tangle solutions for integrases, a protein family that changes DNA topology. J. Knot Theory Ramif., 16, 969–995.
Cerf, C. (1997). Nullification writhe and chirality of alternating links. J. Knot Theory Ramif., 6, 621–632.
Cerf, C. (1998). A note on the tangle model for DNA recombination. Bull. Math. Biol., 60, 67–78.
Cerf, C., & Stasiak, A. (2000). A topological invariant to predict the three-dimensional writhe of ideal configurations of knots and links. Proc. Natl. Acad. Sci. USA, 97, 3795–3798.
Cerf, C., & Stasiak, A. (2003). Linear relations between writhe and minimal crossing number in Conway families of ideal knots and links. New J. Phys., 5, 87.1–87.5.
Chen, J., & Seeman, N. C. (1991). Synthesis from DNA of a molecule with the connectivity of a cube. Nature, 350, 631–633.
Cozzarelli, N. R., et al. (1984). A topological treatment of recombination and topoisomerases. Cold Spring Harbor Symp. Quant. Biol., 49, 383–400.
Darcy, I. K. (2008). Modeling protein-DNA complexes with tangle. Comput. Math. Appl., 55, 924–937.
Diao, Y., Ernst, C., & Stasiak, A. (2009). A partial ordering of knots through diagrammatic unknotting. J. Knot Theory Ramif., 6, 621–632.
Du, S. M., et al. (1995). Topological transformations of synthetic DNA knots. Biochemistry, 34, 673–682.
Erben, C. M., et al. (2007). A self-assembled DNA Bipyramid. J. Am. Chem. Soc., 129, 6992–6993.
Ernst, C., & Sumners, D. W. (1990). A calculus for rational tangles: applications to DNA recombination. Math. Proc. Camb. Philos. Soc., 108, 489–515.
Fuller, F. B. (1971). The writhing number of a space curve. Proc. Natl. Acad. Sci. USA, 68, 815–819.
Goodman, R. P., et al. (2005). Rapid chiral assembly of rigid DNA building blocks for molecular nanofabrication. Science, 310, 1661–1665.
Grayson, N. E., Taormina, A., & Twarock, R. (2009). DNA duplex cage structures with icosahedral symmetry. Theor. Comput. Sci., 410, 1440–1447.
Hu, G., & Qiu, W. Y. (2009). Extended Goldberg polyhedral links with even tangles. MATCH Commun. Math. Comput. Chem., 61, 737–752.
Hu, G. et al. (2009). The architecture of Platonic polyhedral links. J. Math. Chem., 46, 592–603.
Hu, G. et al. (2010). The complexity of Platonic and Archimedean polyhedral links. J. Math. Chem., 48, 401–412.
He, Y., et al. (2008). Hierarchical self-assembly of DNA into symmetric supramolecular polyhedra. Nature, 452, 198–202.
Jones, V. F. R. (1985). A polynomial invariant for knots via von Neumann algebras. Bull. Am. Math. Soc., 12, 103–111.
Jonoska, N., & Saito, M. (2002). Boundary components of thickened graphs. In N. Jonoska & N. C. Seeman (Eds.), LNCS: Vol. 2340. DNA 7 (pp. 70–81). Heidelberg: Springer.
Jonoska, N., & Twarock, R. (2008). Blueprints for dodecahedral DNA cages. J. Phys. A, Math. Theor., 41, 304043.
Qiu, W. Y. (2000). Knot theory, DNA topology, and molecular symmetry breaking. In D. Bonchev & D. H. Rouvray (Eds.), Mathematical chemistry series: Vol. 6. Chemical topology-applications and techniques (pp. 175–237). Amsterdam: Gordon and Breach (Chap. 3).
Qiu, W. Y., & Zhai, X. D. (2005). Molecular design of Goldberg polyhedral links. J. Mol. Struct., Theochem, 756, 163–166.
Qiu, W. Y., Wang, Z., & Hu, G. (2010). The chemistry and mathematics of DNA polyhedra. New York: NOVA.
Ramsing, N. B., & Jovin, T. M. (1988). Parallel stranded duplex DNA. Nucleic Acids Res., 16, 6659–6676.
Rothemund, P. W. K. (2006). Folding DNA to create nanoscale shapes and patterns. Nature, 440, 297–302.
Seeman, N. C. (1982). Nucleic acid junctions and lattices. J. Theor. Biol., 99, 237–247.
Seeman, N. C. (2000). In the Nick of Space: Generalized nucleic acid complementarily and the development of DNA nanotechnology. Synlett, 11, 1536–1548.
Shih, W. M., Quispe, J. D., & Joyce, G. F. (2004). A 1.7-kilobase single-stranded DNA that folds into a nanoscale octahedron. Nature, 427, 618–621.
Stasiak, A., et al. (1996). Electrophoretic mobility of DNA knots. Nature, 384, 122–122.
Sumners, D. W., et al. (1995). Analysis of the mechanism of DNA recombination using tangles. Q. Rev. Biophys., 28, 253–313.
White, J. H. (1969). Self-linking and the Gauss integral in higher dimensions. Am. J. Math., 91, 693–728.
White, J. H., & Cozzarelli, N. R. (1984). A simple topological method for describing stereoisomers of DNA catenanes and knots. Proc. Natl. Acad. Sci. USA, 81, 3322–3326.
White, J. H., Millett, K. C., & Cozzarelli, N. R. (1987). Description of the topological entanglement of DNA catenanes and knots by a powerful method involving strand passage and recombination. J. Mol. Biol., 197, 585–603.
Zhang, C., et al. (2008). Conformational flexibility facilitates self-assembly of complex DNA nanostructures. Proc. Natl. Acad. Sci. USA, 105, 10665–10669.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hu, G., Wang, Z. & Qiu, WY. Topological Analysis of Enzymatic Actions on DNA Polyhedral Links. Bull Math Biol 73, 3030–3046 (2011). https://doi.org/10.1007/s11538-011-9659-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-011-9659-z