Abstract
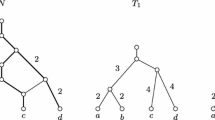
Suppose N is a phylogenetic network indicating a complicated relationship among individuals and taxa. Often of interest is a much simpler network, for example, a species tree T, that summarizes the most fundamental relationships. The meaning of a species tree is made more complicated by the recent discovery of the importance of hybridizations and lateral gene transfers. Hence, it is desirable to describe uniform well-defined procedures that yield a tree given a network N.
A useful tool toward this end is a connected surjective digraph (CSD) map φ:N→N′ where N′ is generally a much simpler network than N. A set W of vertices in N is “restricted” if there is at most one vertex u∉W from which there is an arc into W, thus yielding a bottleneck in N. A CSD map φ:N→N′ is “restricted” if the inverse image of each vertex in N′ is restricted in N. This paper describes a uniform procedure that, given a network N, yields a well-defined tree called the “restricted tree” of N. There is a restricted CSD map from N to the restricted tree. Many relationships in the tree can be proved to appear also in N.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bandelt, H.-J., & Dress, A. (1992). Split decomposition: a new and useful approach to phylogenetic analysis of distance data. Mol. Phylogenet. Evol., 1, 242–252.
Baroni, M., Semple, C., & Steel, M. (2004). A framework for representing reticulate evolution. Ann. Comb., 8, 391–408.
Baroni, M., Semple, C., & Steel, M. (2006). Hybrids in real time. Syst. Biol., 55, 46–56.
Cardona, G., Rosselló, F., & Valiente, G. (2009). Comparison of tree-child phylogenetic networks. IEEE/ACM Trans. Comput. Biol. Bioinform., 6(4), 552–569.
Dagan, T., Artzy-Randrup, Y., & Martin, W. (2008). Modular networks and cumulative impact of lateral transfer in prokaryote genome evolution. Proc. Natl. Acad. Sci. USA, 105, 10039–10044.
Degnan, J. H., & Rosenberg, N. A. (2006). Discordance of species trees with their most likely gene trees, PLos. Genetics, 2(5), e68.
Doolittle, W. F., & Bapteste, E. (2007). Pattern pluralism and the Tree of Life hypothesis. Proc. Natl. Acad. Sci. USA, 104, 2043–2049.
Dress, A., Moulton, V., Steel, M., & Wu, T. (2010). Species, clusters and the ‘tree of life’: a graph-theoretic perspective. J. Theoret. Biol., 265(4), 535–542.
Gusfield, D., Eddhu, S., & Langley, C. (2004). Optimal, efficient reconstruction of phylogenetic networks with constrained recombination. J. Bioinform. Comput. Biol., 2, 173–213.
Hahn, G., & Tardif, C. (1997). Graph homomorphisms: structure and symmetry. In G. Hahn & G. Sabidussi (Eds.), NATO Adv. Sci. Inst. Ser. C: Math. Phys. Sci.: Vol. 497. Graph symmetry: algebraic methods and applications (pp. 107–166). Dordrecht: Kluwer Academic.
Hell, P., & Nešetřil, J. (2004). Graphs and homomorphisms. London: Oxford University Press.
van Iersel, L. J. J., Keijsper, J. C. M., Kelk, S. M., Stougie, L., Hagen, F., & Boekhout, T. (2009). Constructing level-2 phylogenetic networks from triplets. IEEE/ACM Trans. Comput. Biol. Bioinform., 6(43), 667–681.
Jin, G., Nakhleh, L., Snir, S., & Tuller, T. (2007). Inferring phylogenetic networks by the maximum parsimony criterion: a case study. Mol. Biol. Evol., 24(1), 324–337.
Moret, B. M. E., Nakhleh, L., Warnow, T., Linder, C. R., Tholse, A., Padolina, A., Sun, J., & Timme, R. (2004). Phylogenetic networks: modeling, reconstructibility, and accuracy. IEEE/ACM Trans. Comput. Biol. Bioinform., 1, 13–23.
Morrison, D. A. (2009). Phylogenetic networks in systematic biology (and elsewhere). In R. M. Mohan (Ed.), Research advances in systematic biology (Global Research Network, Trivandrum, India) (pp. 1–48).
Nakhleh, L., Warnow, T., & Linder, C. R. (2004). Reconstructing reticulate evolution in species—theory and practice. In P. E. Bourne & D. Gusfield (Eds.), Proceedings of the eighth annual international conference on computational molecular biology (RECOMB’04, 27–31 March 2004, San Diego, California) (pp. 37–346). New York: ACM.
Rosenberg, N., & Tao, R. (2008). Discordance of species trees with their most likely gene trees: the case of five taxa. Syst. Biol., 57(1), 131–140.
Wang, L., Zhang, K., & Zhang, L. (2001). Perfect phylogenetic networks with recombination. J. Comput. Biol., 8, 69–78.
Willson, S. J. (2010), Relationships among phylogenetic networks (submitted). arXiv:1005.2108v1 [q-bio.PE].
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Willson, S.J. Restricted Trees: Simplifying Networks with Bottlenecks. Bull Math Biol 73, 2322–2338 (2011). https://doi.org/10.1007/s11538-010-9624-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-010-9624-2