Abstract
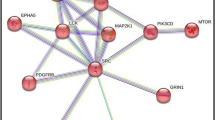
In this study, three kinds of phenothiazine drugs were analyzed to explore their potential antitumor mechanisms. First, target proteins that could interact with chlorpromazine, fluphenazine and trifluoperazine were predicted. Then, the target proteins of the three drugs were intersected. Cell signaling pathway enrichment and related disease enrichment were conducted for the intersected proteins to extract the enrichment categories associated with tumors. By regulation network analysis of the protein interactions, the mechanisms of action of these target proteins in tumor tissue were clarified, thus confirming the potential antitumor mechanisms of the phenothiazine drugs. The final results of cell signaling pathway enrichment and related disease enrichment showed that the categories with the highest score were all found in tumors. Target proteins belonging to the tumor category included signaling pathway members such as Wnt, MAPK and retinoic acid receptor. Moreover, another target protein, MAPK8, could indirectly act on target proteins CDK2, IGF1R, GSK3B, RARA, FGFR2 and MAPK10, thereby affecting tumor cell division and proliferation. Therefore, phenothiazine drugs may have potential antitumor effects, and tumor-associated target proteins play important roles in the process of cell signaling transduction cascades.
Article PDF
Similar content being viewed by others
References
Yde C W, Clausen M P, Bennetzen M V, et al. The antipsychotic drug chlorpromazine enhances the cytotoxic effect of tamoxifen in tamoxifen-sensitive and tamoxifen-resistant human breast cancer cells. Anticancer Drugs, 2009, 20: 723–735
Eriksson A, Yachnin J, Lewensohn R, et al. DNA-dependent protein kinase is inhibited by trifluoperazine. Biochem Biophys Res Commun, 2001, 283: 726–731
Liang W, Yang C Z. Effects of phenothiazine derivatives on protein kinase C and multidrug resistance of tumor cells. Chin Sci Bull, 1998, 43: 1179–1183
Lee M S, Johansen L, Zhang Y, et al. The novel combination of chlorpromazine and pentamidine exerts synergistic antiproliferative effects through dual mitotic action. Cancer Res, 2007, 67: 11359–11367
Wiklund E D, Catts V S, Catts S V, et al. Cytotoxic effects of antipsychotic drugs implicate cholesterol homeostasis as a novel chemotherapeutic target. Int J Cancer, 2010, 126: 28–40
Zhelev Z, Ohba H, Bakalova R, et al. Phenothiazines suppress proliferation and induce apoptosis in cultured leukemic cells without any influence on the viability of normal lymphocytes. Phenothiazines and leukemia. Cancer Chemother Pharmacol, 2004, 53: 267–275
Fond G, Macgregor A, Attal J, et al. Antipsychotic drugs: Pro-cancer or anti-cancer? A systematic review. Med Hypotheses, 2012, 79: 38–42
Burgess D J. Anticancer drugs: Antipsychotic to anticancer agent? Nat Rev Drug Discov, 2012, 11: 516
Barrett T, Suzek T O, Troup D B, et al. NCBI GEO: Mining millions of expression profiles-database and tools. Nucleic Acids Res, 2005, 33: D562–D566
Shi Q, Pavey E S, Carter R E. Bonferroni-based correction factor for multiple, correlated endpoints. Pharm Stat, 2012, 11: 300–309
Orlov Y L, Zhou J, Lipovich L, et al. Quality assessment of the Affymetrix U133A&B probesets by target sequence mapping and expression data analysis. In Silico Biol, 2007, 7: 241–260
Lamb J, Crawford E D, Peck D, et al. The Connectivity Map: Using gene-expression signatures to connect small molecules, genes, and disease. Science, 2006, 313: 1929–1935
Luo H, Chen J, Shi L, et al. DRAR-CPI: A server for identifying drug repositioning potential and adverse drug reactions via the chemical-protein interactome. Nucleic Acids Res, 2011, 39: W492–W498
Dennis G J, Sherman B T, Hosack D A, et al. DAVID: Database for annotation, visualization, and integrated discovery. Genome Biol, 2003, 4: P3
Warde-Farley D, Donaldson S L, Comes O, et al. The GeneMANIA prediction server: Biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res, 2010, 38: W214–W220
Hu Z, Snitkin E S, Delisi C. VisANT: An integrative framework for networks in systems biology. Brief Bioinform, 2008, 9: 317–325
Hu Z, Mellor J, Wu J, et al. VisANT: Data-integrating visual framework for biological networks and modules. Nucleic Acids Res, 2005, 33: W352–W357
Hu Z, Mellor J, Wu J, et al. VisANT: An online visualization and analysis tool for biological interaction data. BMC Bioinformatics, 2004, 5: 17
Ogino S, Shima K, Baba Y, et al. Colorectal cancer expression of peroxisome proliferator-activated receptor gamma (PPARG, PPARgamma) is associated with good prognosis. Gastroenterology, 2009, 136: 1242–1250
Girnun G. PPARG: A new independent marker for colorectal cancer survival. Gastroenterology, 2009, 136: 1157–1160
Clarke N, Germain P, Altucci L, et al. Retinoids: Potential in cancer prevention and therapy. Expert Rev Mol Med, 2004, 6: 1–23
Papi A, Rocchi P, Ferreri A M, et al. RXRgamma and PPARgamma ligands in combination to inhibit proliferation and invasiveness in colon cancer cells. Cancer Lett, 2010, 297: 65–74
Chang J T, Nevins J R. GATHER: A systems approach to interpreting genomic signatures. Bioinformatics, 2006, 22: 2926–2933
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Qi, L., Ding, Y. Potential antitumor mechanisms of phenothiazine drugs. Sci. China Life Sci. 56, 1020–1027 (2013). https://doi.org/10.1007/s11427-013-4561-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-013-4561-6