Abstract
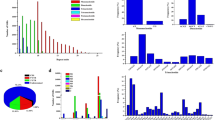
Opium poppy (Papaver somniferum L.) is an important pharmaceutical crop with very few genetic marker resources. To expand these resources, we sequenced genomic DNA using pyrosequencing technology and examined the DNA sequences for simple sequence repeats (SSRs). A total of 1,244,412 sequence reads were obtained covering 474 Mb. Approximately half of the reads (52 %) were assembled into 166,724 contigs representing 105 Mb of the opium poppy genome. A total of 23,283 non-redundant SSRs were identified in 18,944 contigs (11.3 % of total contigs). Trinucleotide and tetranucleotide repeats were the most abundant SSR repeats, accounting for 49.0 and 27.9 % of all SSRs, respectively. The AAG/TTC repeat was the most abundant trinucleotide repeat, representing 19.7 % of trinucleotide repeats. Other SSR repeat types were AT-rich. A total of 23,126 primer pairs (98.7 % of total SSRs) were designed to amplify SSRs. Fifty-three genomic SSR markers were tested in 37 opium poppy accessions and seven Papaver species for determination of polymorphism and transferability. Intraspecific polymorphism information content (PIC) values of the genomic SSR markers were intermediate, with an average 0.17, while the interspecific average PIC value was slightly higher, 0.19. All markers showed at least 88 % transferability among related species. This study increases sequence coverage of the opium poppy genome by sevenfold and the number of opium poppy-specific SSR markers by sixfold. This is the first report of the development of genomic SSR markers in opium poppy, and the genomic SSR markers developed in this study will be useful in diversity, identification, mapping and breeding studies in opium poppy.

Similar content being viewed by others
References
Abdelkrim J, Robertson BC, Stanton JA, Gemmell NJ (2009) Fast cost-efective development of species-specific microsatalite markers by genomic sequencing. Biotechniques 46:185–192. doi:10.2144/000113084
Acharya HS, Sharma V (2009) Molecular characterization of opium poppy (Papaver somniferum) germplasm. Am J Infect Dis 5:148–153
Bennett MD, Smith JB (1976) Nuclear DNA amounts in angiosperms. Philos Trans R Soc Lond B Biol Sci 274:227–274
Carolan JC, Hook ILI, Chase MW, Kadereit JW, Hodkinson TR (2006) Phylogenetics of Papaver and related genera based on DNA sequences from ITS nuclear ribosomal DNA and plastid trnL intron and trnL-F intergenic spacers. Ann Bot 98:141–155. doi:10.1093/aob/mcl079
Cavagnaro PF, Douglas AS, Yang L, Simon PW, Harkins TT, Kodira CD, Huang S, Weng Y (2010) Genome-wide characterization of simple sequence repeats in cucumber (Cucumis sativus L.). BMC Genom 11:569. doi:10.1186/1471-2164-11-569
Chabane K, Ablett GA, Cordeiro GM, Valkoun J, Henry RJ (2005) EST versus genomic derived microsatellite markers for genotyping wild and cultivated barley. Gen Resour Crop Evol 52:903–909. doi:10.1007/s10722-003-6112-7
Chevreux B, Pfisterer T, Drescher B, Driesel AJ, Müller WEG, Wetter T, Suhai S (2004) Using the miraEST assembler for reliable and automated mRNA transcript assembly and SNP detection in sequenced ESTs. Genome Res 14:1147–1159. doi:10.1101/gr.1917404
Dittbrenner A, Lohwasser U, Mock HP, Börner A (2008) V International symposium on the taxonomy of cultivated plants. In: Groendijk-Wilders N, Alexander C, Berg RGVD, Hetterscheid WLA (eds) Molecular and phytochemical studies of Papaver somniferum in the context of infraspecific classification. ISHS Acta Horticulturae, Wageningen, pp 81–88
Doyle JJ, Doyle JE (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Ellis JR, Burke JM (2007) EST-SSRs as a resource for population genetic analyses. Heredity 99:125–132. doi:10.1038/sj.hdy.6801001
Facchini PJ, De Luca V (2008) Opium poppy and Madagascar periwinkle: model non-model systems to investigate alkaloid biosynthesis in plants. Plant J Cell Mol Biol 54:763–784. doi:10.1111/j.1365-313X.2008.03438.x
Gumuscu A, Arslan N (2008) Researches on heterosis on yield and yield components of some poppy. Tarım Bilimleri Dergisi 14:365–373
Jones N, Ougham H, Thomas H, Pasakinskiene I (2009) Markers and mapping revisited: finding your gene. New Phytol 183:935–966. doi:10.1111/j.1469-8137.2009.02933.x
Kumar S, Blaxter ML (2010) Comparing de novo assemblers for 454 transcriptome data. BMC Genom 11:571. doi:10.1186/1471-2164-11-571
Lavania UC, Srivastava S (1999) Quantitative delineation of karyotype variation in Papaver as an measure of phylogenetic differentiation origin. Curr Sci 77:429–435
Lee RM, Thimmapuram J, Thinglum KA, Gong G, Hernandez AG, Wright CL, Kim RW, Mikel MA, Tranel PJ (2009) Sampling the waterhemp (Amaranthus tuberculatus) genome using pyrosequencing technology. Weed Sci 57:463–469. doi:http://dx.doi.org/10.1614/WS-09-021.1
Lee EJ, Jin GN, Lee KL, Han MS, Lee YH, Yang MS (2011) Exploiting expressed sequence tag databases for the development and characterization of gene-derived simple sequence repeat markers in the opium poppy (Papaver somniferum L.) for forensic applications. J Forensic Sci 56:1131–1135. doi:10.1111/j.1556-4029.2011.01810.x
Lepais O, Bacles CFE (2011) De novo discovery and multiplexed amplification of microsatellite markers for black alder (Alnus glutinosa) and related species using SSR-enriched shotgun pyrosequencing. J Hered 102:627–632. doi:10.1093/jhered/esr062
Li YC, Korol AB, Fahima T, Beiles A, Nevo E (2002) Microsatellites: genomic distribution, putative functions and mutational mechanisms: a review. Mol Ecol 11:2453–2465. doi:10.1046/j.1365-294X.2002.01643.x
Li YC, Korol AB, Fahima T, Nevo E (2004) Microsatellites within genes: structure, function, and evolution. Mol Biol Evol 21:991–1007
Lipman DJ, Pearson WR (1985) Rapid and sensitive protein similarity searches. Science 227:1435–1441. doi:10.1126/science.2983426
Morgante M, Hanafey M, Powell W (2002) Microsatellites are preferentially associated with nonrepetitive DNA in plant genomes. Nat Genet 30:194–200. doi:10.1038/ng822
Parmaksiz I, Ozcan S (2011) Morphological, chemical and molecular analyses of Turkish Papaver accessions (Sect. Oxytona). Turk J Bot 35:1–16. doi:10.3906/bot-1003-39
Perrier X, Jacquemoud-Collet JP (2006) DARwin software. http://darwin.cirad.fr/darwin
Roldan-Ruiz I, Dendauw J, Bockstaele EV, Depicker A, Loose MD (2000) AFLP markers reveal high polymorphic rates in ryegrasses (Lolium spp.). Mol Breed 6:125–134. doi:10.1023/A:1009680614564
Rozen S, Skaletsky HJ (2000) Bioinformatics methods and protocols: methods in molecular biology. In: Krawetz S, Misener S (eds) Primer3 on the WWW for general users and for biologist programmers. Humana Press, Totowa, pp 365–386
Saunders JA, Pedroni MJ, Penrose LDJ, Fist AJ (2001) AFLP analysis of opium poppy. Crop Sci 41:1596–1601. doi:10.2135/cropsci2001.4151596x
Schulz H, Baranska M, Quilitzsch R, Schütze W (2004) Determination of alkaloids in capsules, milk and ethanolic extracts of poppy (Papaver somniferum L.) by ATR-FT-IR and FT-Raman spectroscopy. Analyst 129:917–920. doi:10.1039/B408930H
Selale H, Celik I, Gultekin V, Allmer J, Doganlar S, Frary A (2013) Development of EST-SSR markers for diversity and breeding studies in opium poppy (Papaver somniferum L.). Plant Breed 132:344–351. doi:10.1111/pbr.12059
Straka P, Nothnagel T (2002) A genetic map of Papaver somniferum L. based on molecular and morphological markers. J Herbs Spices Med Plants 9:235–241. doi:10.1300/J044v09n02_a
Tian CX, Liang XF, Yang M, Zheng HZ, Dou YQ, Cao L (2012) Isolation and characterization of novel genomic and EST-SSR markers in Coreoperca whiteheadi Boulenger and cross-species amplification. Int J Mol Sci 13:13203–13211. doi:10.3390/ijms131013203
Turkish Soil Product Office (2009) Opium poppy report. Turkish Soil Production Office, Ankara
Varshney RK, Graner A, Sorrells ME (2005) Genic microsatellite markers in plants: features and applications. Trends Biotech 23:48–55. doi:10.1016/j.tibtech.2004.11.005
Winzer T, Gazda V, He Z, Kaminski F, Kern M, Larson TR, Li Y, Meade F, Teodor R, Vaistij FE, Walker C, Bowser TA, Graham IA (2012) A Papaver somniferum 10-gene cluster for synthesis of the anticancer alkaloid noscapine. Sci Exp 336:1704–1708. doi:10.1126/science.1220757
Zalapa JE, Cuevas H, Zhu H, Steffan S, Senalik D, Zeldin E, Mccown B, Harbut R, Simon P (2012) Using next-generation sequencing approaches to isolate simple sequence repeat (SSR) loci in the plant sciences. Am J Bot 99:193–208. doi:10.3732/ajb.1100394
Zhu H, Senalik D, McCown BH, Zeldin EL, Speers J, Hyman J, Bassil N, Hummer K, Simon PW, Zalapa JE (2012) Mining and validation of pyrosequenced simple sequence repeats (SSRs) from American cranberry (Vaccinium macrocarpon Ait.). Theor Appl Genet 124:87–96. doi:10.1007/s00122-011-1689-2
Ziegler J, Facchini PJ, Geissler R, Schmidt J, Ammer C, Kramell R, Voigtländer S, Gesell A, Pienkny S, Brandt W (2009) Evolution of morphine biosynthesis in opium poppy. Phytochemistry 70:1696–1707. doi:10.1016/j.phytochem.2009.07.006
Acknowledgments
The authors thank Dr. Hüseyin Camci, Dr. Arzu Köse and Dr. Ferda Kosar from the Anatolia Agricultural Research Institute for providing plant material. We are also grateful to Ali Üncü and Ayşe Dülger Üncü for SSR validation by sequencing. This study was supported by The Scientific and Technological Research Council of Turkey (TUBITAK) project no. 109O797.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Celik, I., Gultekin, V., Allmer, J. et al. Development of genomic simple sequence repeat markers in opium poppy by next-generation sequencing. Mol Breeding 34, 323–334 (2014). https://doi.org/10.1007/s11032-014-0036-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-014-0036-0