Abstract
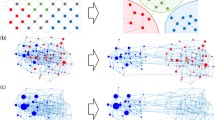
Our goal is to match some dynamical aspects of biological systems with that of networks of coupled logistic maps. With these networks we generate sequences of iterates, convert them to symbol sequences by coarse-graining, and count the number of times combinations of symbols occur. Comparison of this with the number of times these combinations occur in experimental data—a sequence of interbeat intervals for example—is a measure of the fitness of each network to describe the target data. The most fit networks provide a cartoon that suggests a decomposition of the experimental data into a component that may be produced by a simple dynamical subsystem, and a residual component, the result of detailed, particular characteristics of the system that generated the target data. In the space of all network parameters, each point corresponds to a particular network. We construct a fitness landscape when we assign a fitness to each point. Because the parameters are distributed continuously over their ranges, and because fitnesses are estimated numerically, any plot of the landscape involves a finite sample of parameter values. We’ll investigate how the local landscape geometry changes when the array of sample parameters is refined, and use the landscape geometry to explore complex relations between local fitness maxima.























Similar content being viewed by others
References
Wright, S.: Evolution in Mendelian populations. Genetics 16, 97–159 (1931)
Wright, S.: The role of mutation, inbreeding, crossbreeding, and selection in evolution. Proc. Sixth Int. Cong. Genetics 1, 356–366 (1932)
Nowak, M.: Evolutionary Dynamics: Exploring the Equations of Life. Harvard University Press (2006)
Waddington, C.: Principles of Development and Differentiation. Macmillian (1966)
Waddington, C.: Strategy of the Genes. A Discussion of Some Aspects of Theoretical Biology. George Allen & Unwin (1957)
Thom, R.: Structural Stability and Morphogenesis. An Outline of a General Theory of Models. Benjamin (1975)
Frauenfelder, H., Sligar, S., Wolynes, P.: The energy landscape and motions of proteins. Science 254, 1589–1603 (1991)
Frauenfelder, H., et al.: A unified model of protein dynamics. Proc. Nat. Acad. Sci. U.S.A. 106, 5129–5134 (2009)
Frauenfelder, H., Leeson, D.: The energy landscape in non-biological and biological molecules. Nat. Struct. Biol. 5, 757–759 (1998)
Frauenfelder, H.: Energy landscape and dynamics of biomolecules. J. Biological Physics 31, 413–416 (2005)
Jeffrey, H.: Chaos game representation of gene structure. Nucl. Acid Res. 18, 2163–2170 (1990)
Jeffrey, H.: Chaos game visualization of sequences. Comput. Graph. 16, 25–33 (1992)
Lind, D., Marcus, B.: An Introduction to Symbolic Dynamics and Coding. Cambridge University Press (1995)
Kitchens, B.: Symbolic Dynamics: One-Sided, Two-Sided, and Countable State Markov Shifts. Springer (1998)
Blanchard, F., Maass. A., Nogueira, A.: Topics in Symbolic Dynamics and Applications. Cambridge University Press (2000)
Hutchinson, J.: Fractals and self-similarity. Indiana Univ. J. Math. 30, 713–747 (1981)
Barnsley, M.: Fractals Everywhere. 2nd ed., Academic Press (1993)
Stewart, I.: Order within the Chaos Game? Dynamics Newsletter 3, 4–9 (1989)
May, R.: Simple mathematical models with very complicated dynamics. Nature 261, 459–467 (1976)
Devaney, R.: An Introduction to Chaotic Dynamical Systems. 2nd ed., Addison-Wesley (1989)
Rasband, N.: Chaotic Dynamics of Nonlinear Systems. Wiley-Interscience (1990)
Hindmarsh, J., Rose, M.: A model of neuronal bursting using using three coupled first order differential equations. Proc. Roy. Soc. London. Biol. Sci. 221, 87–102 (1984)
Storace, M., Linaro, D., de Lange, E.: The Hindmarsh-Rose neuron model: bifurcation analysis and piecewise-linear approximations. Chaos 18, 033128 (2008)
Sorkin, G.: Efficient simulated annealing on fractal energy landscapes. Algorithmica 6, 367–418 (1991)
Adami, C.: Introduction to Artificial Life. Springer (1997)
Weinberger, E., Stadler, P.: Why some fitness landscapes are fractal. J. Theoret. Biol. 163, 255–275 (1993)
Bahar, S.: The Essential Tension: Competition, Cooperation and Multilevel Selection in Evolution. Springer Nature (2018)
Strogatz, S.: Sync: The Emerging Science of Spontaneous Order. Hyperion Books (2003)
http://physionet.org/challenge/2002/event-2.shtml#downloading-the-challenge-dataset Accessed 10 Feb 2013 (2013)
Wessel, N., et al.: Classifying simulated and physiological heart rate variability signals. Computers in Cardiology 29, 133–135 (2002)
Accardo, A., et al.: Use of the fractal dimension for the analysis of electroencephalographic time series. Biol. Cybernetics 77, 339–350 (1997)
Kulish, V., Sourin, A., Sourina, O.: Human encephalograms seen as fractal time series: Mathematical analysis and visualization. Comput. Biol. Med. 36, 291–302 (2006)
Schalk, G., et al.: BCI2000: A general-purpose brain-computer interface (BCI) system. IEEE Trans. Biomed. Eng. 51, 1034–1043 (2004)
Frame, M., Urry, A.: Fractal Worlds: Grown, Built, and Imagined. Yale University Press (2016)
Falconer, K.: Fractal Geometry. Mathematical Foundations and Applications. 3rd ed., Wiley (2014)
Thew, N.: A Dictionary of Driven Iterated Function Systems to Characterize Chaotic Time Series. Yale University Applied Mathematics Senior Thesis (2014)
Acknowledgements
This paper is an elaboration of the work in the first author’s Applied Mathematics Yale senior thesis [36]. In early stages of this project we benefitted from many thoughtful and energetic discussions with Rachel Lawrence and Hannah Otis. Comments by a very thoughtful anonymous referee led to improvements in the readability of this paper.
Funding
This research was supported by grant funding for the first author from the Science and Technology Research Scholars (STARS) II program at Yale University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
We have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article belongs to the Topical Collection: The Revolutionary Impact of Landscapes in Biology
Guest Editors: Robert Austin, Shyamsunder Erramilli, Sonya Bahar
Rights and permissions
About this article
Cite this article
Driver, N., Frame, M. Fitness landscapes for coupled map lattices. J Biol Phys 47, 215–235 (2021). https://doi.org/10.1007/s10867-021-09577-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10867-021-09577-6