Abstract
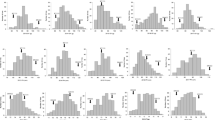
Both heading date and plant height are important traits related to grain yield in rice. In this study, a recombinant inbred lines (RILs) population was used to map quantitative trait loci (QTLs) for both traits under 3 long-day (LD) environments and 1 short-day (SD) environment. A total of eight QTLs for heading date and three QTLs for plant height were detected by composite interval mapping under LD conditions. Additional one QTL for heading date and three QTLs for plant height were identified by Two-QTL model under LD conditions. Among them, major QTLs qHd7.1, qHd7.2 and qHd8 for heading date, and qPh1 and qPh7.1 for plant height were commonly detected. qHd7.1 and qHd7.2 were mapped to small regions of less than 1 cM. Genome position comparison of previously cloned genes with QTLs detected in this study revealed that qHd5 and qPh3.1 were two novel QTLs. The alleles of these QTLs increasing trait values were dispersed in both parents, which well explained the transgressive segregation observed in this population. In addition, the interaction between qHd7.1 and qHd8 was detected under all LD conditions. Multiple-QTL model analysis revealed that all QTLs and their interactions explained over 80% of heading date variation and 50% of plant height variation. Two heading date QTLs were detected under SD condition. Of them, qHd10 were commonly identified under LD condition. The difference in QTL detection between LD and SD conditions indicated most heading date QTLs are sensitive to photoperiod. These findings will benefit breeding design for heading date and plant height in rice.




Similar content being viewed by others
References
Asano K, Takashi T, Miura K, Qian Q, Kitano H, Matsuoka M, Ashikari M (2007) Genetic and molecular analysis of utility of sd1 alleles in rice breeding. Breed Sci 57:53–58
Asano K, Yamasaki M, Takuno S, Miura K, Katagiri S, Ito T, Doi K, Wu J, Ebana K, Matsumoto T (2011) Artificial selection for a green revolution gene during japonica rice domestication. Proc Natl Acad Sci USA 108:11034–11039
Broman KW, Wu H, Sen Ś, Churchill GA (2003) R/qtl: QTL mapping in experimental crosses. Bioinformatics 19:889–890
Cavanagh C, Morell M, Mackay I, Powell W (2008) From mutations to MAGIC: resources for gene discovery, validation and delivery in crop plants. Curr Opin Plant Biol 11:215–221
Dalrymple DG (1986) Development and spread of high-yielding rice varieties in developing countries. Int Rice Res Inst 17–18
Dell’Acqua M, Gatti DM, Pea G, Cattonaro F, Coppens F, Magris G, Hlaing AL, Aung HH, Nelissen H, Baute J (2015) Genetic properties of the MAGIC maize population: a new platform for high definition QTL mapping in Zea mays. Genome Biol 16:1
Dempster AP, Laird NM, Rubin DB (1977) Maximum likelihood from incomplete data via the EM algorithm. J R Stat Soc Series B Stat Methodol 39:1–38
Doi K, Izawa T, Fuse T, Yamanouchi U, Kubo T, Shimatani Z, Yano M, Yoshimura A (2004) Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Gene Dev 18:926–936
Fujino K, Sekiguchi H (2005) Identification of QTLs conferring genetic variation for heading date among rice varieties at the northern-limit of rice cultivation. Breed Sci 55:141–146
Gao H, Jin M, Zheng X, Chen J, Yuan D, Xin Y, Wang M, Huang D, Zhang Z, Zhou K (2014) Days to heading 7, a major quantitative locus determining photoperiod sensitivity and regional adaptation in rice. Proc Natl Acad Sci USA 111:16337–16342
Gu X, Foley ME (2007) Epistatic interactions of three loci regulate flowering time under short and long daylengths in a backcross population of rice. Theor Appl Genet 114:745–754
Haley CS, Knott SA (1992) A simple regression method for mapping quantitative trait loci in line crosses using flanking markers. Heredity 69:315–324
Hargrove TR, Cabanilla VL (1979) The impact of semidwarf varieties on Asian rice-breeding programs. Bioscience 29:731–735
Hori K, Ogiso-Tanaka E, Matsubara K, Yamanouchi U, Ebana K, Yano M (2013) Hd16, a gene for casein kinase I, is involved in the control of rice flowering time by modulating the day-length response. Plant J 76:36–46. doi:10.1111/tpj.12268
Huang X, Feng Q, Qian Q, Zhao Q, Wang L, Wang A, Guan J, Fan D, Weng Q, Huang T (2009) High-throughput genotyping by whole-genome resequencing. Genome Res 19:1068–1076
Itoh H, Nonoue Y, Yano M, Izawa T (2010) A pair of floral regulators sets critical day length for Hd3a florigen expression in rice. Nat Genet 42:635–638. doi: 10.1038/ng.606
Kao CH, Zeng ZB, Teasdale RD (1999) Multiple interval mapping for quantitative trait loci. Genetics 152:1203–1216
Khush GS (1999) Green revolution: preparing for the 21st century. Genome 42:646–655
Kojima S, Takahashi Y, Kobayashi Y, Monna L, Sasaki T, Araki T, Yano M (2002) Hd3a, a rice ortholog of the Arabidopsis FT gene, promotes transition to flowering downstream of Hd1 under short-day conditions. Plant Cell Physiol 43:1096–1105
Lander ES, Botstein D (1989) Mapping Mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121:185–199
Lee S, Jia MH, Jia Y, Liu G (2014) Tagging quantitative trait loci for heading date and plant height in important breeding parents of rice (Oryza sativa). Euphytica 197:191–200
Li Z, Pinson S, Stansel J, Park W (1995) Identification of quantitative trait loci (QTLs) for heading date and plant height in cultivated rice (Oryza sativa L.). Theor Appl Genet 91:374–381
Li Z, Yu S, Lafitte H, Huang N, Courtois B, Hittalmani S, Vijayakumar C, Liu G, Wang G, Shashidhar H (2003) QTL × environment interactions in rice. I. Heading date and plant height. Theor Appl Genet 108:141–153
Li H, Hearne S, Bänziger M, Li Z, Wang J (2010) Statistical properties of QTL linkage mapping in biparental genetic populations. Heredity 105:257–267
Lu L, Yan W, Xue W, Shao D, Xing Y (2012) Evolution and association analysis of Ghd7 in rice. PLoS One 7:e34021. doi:10.1371/journal.pone.0034021
Ming SK (1987) Breeding of semi-dwarf rice. In: Yuang SR (ed) Rice. China Agriculture Press, Beijing, pp 66–67
Monna L, Kitazawa N, Yoshino R, Suzuki J, Masuda H, Maehara Y, Tanji M, Sato M, Nasu S, Minobe Y (2002) Positional cloning of rice semidwarfing gene, sd-1: rice “green revolution gene” encodes a mutant enzyme involved in gibberellin synthesis. DNA Res 9:11–17
Oikawa T, Koshioka M, Kojima K, Yoshida H, Kawata M (2004) A role of OsGA20ox1, encoding an isoform of gibberellin 20-oxidase, for regulation of plant stature in rice. Plant Mol Biol 55:687–700
Ryu CH, Lee S, Cho LH, Kim SL, Lee YS, Choi SC, Jeong HJ, Yi J, Park SJ, Han CD (2009) OsMADS50 and OsMADS56 function antagonistically in regulating long day (LD)-dependent flowering in rice. Plant Cell Environ 32:1412–1427. doi: 10.1111/j.1365-3040.2009.02008.x
Sasaki A, Ashikari M, Ueguchi-Tanaka M, Itoh H, Nishimura A, Swapan D, Ishiyama K, Saito T, Kobayashi M, Khush GS (2002) A mutant gibberellin-synthesis gene in rice. Nature 416:701–701
Sen Ś, Churchill GA (2001) A statistical framework for quantitative trait mapping. Genetics 159:371–387
Spielmeyer W, Ellis MH, Chandler PM (2002) Semidwarf (sd-1), “green revolution” rice, contains a defective gibberellin 20-oxidase gene. Proc Natl Acad Sci USA 99:9043–9048
Sugiyama F, Churchill GA, Higgins DC, Johns C, Makaritsis KP, Gavras H, Paigen B (2001) Concordance of murine quantitative trait loci for salt-induced hypertension with rat and human loci. Genomics 71:70–77
Tan C, Han Z, Yu H, Zhan W, Xie W, Chen X, Zhao H, Zhou F, Xing Y (2013) QTL scanning for rice yield using a whole genome SNP array. J Genet Genomics 40:629–638
Wei X, Xu J, Guo H, Jiang L, Chen S, Yu C, Zhou Z, Hu P, Zhai H, Wan J (2010) DTH8 suppresses flowering in rice, influencing plant height and yield potential simultaneously. Plant Physiol 153:1747–1758
Wu W, Zheng X, Lu G, Zhong Z, Gao H, Chen L, Wu C, Wang H, Wang Q, Zhou K (2013) Association of functional nucleotide polymorphisms at DTH2 with the northward expansion of rice cultivation in Asia. Proc Natl Acad Sci USA 110:2775–2780
Xie W, Feng Q, Yu H, Huang X, Zhao Q, Xing Y, Yu S, Han B, Zhang Q (2010) Parent-independent genotyping for constructing an ultrahigh-density linkage map based on population sequencing. Proc Natl Acad Sci USA 107:10578–10583
Xue W, Xing Y, Weng X, Zhao Y, Tang W, Wang L, Zhou H, Yu S, Xu C, Li X (2008) Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40:761–767
Yamamoto T, Kuboki Y, Lin S, Sasaki T, Yano M (1998) Fine mapping of quantitative trait loci Hd-1, Hd-2 and Hd-3, controlling heading date of rice, as single Mendelian factors. Theor Appl Genet 97:37–44
Yan WH, Wang P, Chen HX, Zhou HJ, Li QP, Wang CR, Ding ZH, Zhang YS, Yu SB, Xing YZ (2011) A major QTL, Ghd8, plays pleiotropic roles in regulating grain productivity, plant height, and heading date in rice. Mol Plant 4:319–330
Yan W, Liu H, Zhou X, Li Q, Zhang J, Lu L, Liu T, Liu H, Zhang C, Zhang Z (2013) Natural variation in Ghd7.1 plays an important role in grain yield and adaptation in rice. Cell Res 23:969–971
Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, Baba T, Yamamoto K, Umehara Y, Nagamura Y (2000) Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12:2473–2483
Yu J, Holland JB, McMullen MD, Buckler ES (2008) Genetic design and statistical power of nested association mapping in maize. Genetics 178:539–551
Zeng ZB, Kao CH, Basten CJ (1999) Estimating the genetic architecture of quantitative traits. Genet Res 74:279–289
Zhang Y, Luo L, Xu C, Zhang Q, Xing Y (2006) Quantitative trait loci for panicle size, heading date and plant height co-segregating in trait-performance derived near-isogenic lines of rice (Oryza sativa). Theor Appl Genet 113:361–368
Zhang L, Li Q, Dong H, He Q, Liang L, Tan C, Han Z, Yao W, Li G, Zhao H (2015) Three CCT domain-containing genes were identified to regulate heading date by candidate gene-based association mapping and transformation in rice. Sci Rep 5:7663
Acknowledgements
We thank Dr. Rukmini Mishra in India in Siksha O Anusandhan University, India for her language editing. This project was partly supported by the Natural Science Foundation of Hubei province, China (2015CFA006) and Wuhan Yellow Crane Talents (Special) Program.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Han, Z., Hu, W., Tan, C. et al. QTLs for heading date and plant height under multiple environments in rice. Genetica 145, 67–77 (2017). https://doi.org/10.1007/s10709-016-9946-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-016-9946-6