Abstract
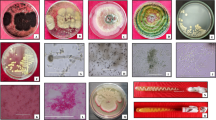
A soil microorganism, designated as P7, was characterized and investigated for its ability to degrade polyurethane (PU). This bacterial isolate was identified as Acinetobacter gerneri on the basis of 16 s rRNA sequencing and biochemical phenotype analysis. The ability of this organism to degrade polyurethane was characterized by the measurement of growth, SEM observation, measurement of electrophoretic mobility and the purification and characterization of a polyurethane degrading enzyme. The purified protein has a molecular weight of approximately 66 kDa as determined by SDS-PAGE. Substrate specificity was examined using p-nitrophenyl substrates with varying carbon lengths. The highest substrate specificity was observed using p-nitrophenyl-propanate with an activity of 37.58 ± 0.21 U mg−1. Additionally, the enzyme is inhibited by phenylmethylsulfonylfluoride and by ethylenediamine-tetra acetic acid. When grown on Impranil DLN™ YES medium, a lag phase was noted for the first 3 h which was followed by logarithmic growth for 5 h. For the linear portion of growth between 5 and 9 h, a μ value of 0.413 doublings h−1 was calculated. After 9 h of incubation the cell number dramatically decreased resulting in a chalky precipitate. Measurements of electrophoretic mobility indicated the formation of a complex between the PU and A. gerneri P7 cells. A hybrid zeta potential had been generated between the cells and polyurethane. Further evidence for a complex was provided by SEM observation where cells appeared to cluster along the surface of polyurethane particles and along edges of polyurethane films. Occasionally, the cells established an anchor-like structure that connected the cells to polyurethane particles.









Similar content being viewed by others
References
Akutsu Y, Nakajima-Kambe T, Nomura N, Nakahara T (1998) Purification and properties of a polyester polyurethane-degrading enzyme from Comamonas acidovorans TB-35. Appl Environ Microbiol 64:62–67
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Baumann P (1968) Isolation of Acinetobacter from soil and water. J Bacteriol 96:39–42
Bayer O (1947) Polyurethanes. Mod Plast 24:149–152
Bergogne-Bérézin E, Towner KJ (1996) Acinetobacter spp. as nosocomial pathogens: microbiological, clinical, and epidemiological features. Clin Microbiol Rev 9:148–165
Berlau J, Aucken HM, Houang E, Pitt TL (1999) Isolation of Acinetobacter spp. including A. baumannii from vegetables: implications for hospital-acquired infections. J Hosp Infect 42:201–204
Bernards AT, de Beaufort AT, Dijkshoorn L, van Boven CPA (1997) Outbreak of septicaemia in neonates caused by Acinetobacter junii investigated by amplified ribosomal DNA restriction analysis (ARDRA) and four typing methods. J Hosp Infect 35:129–140
Blake RC, Howard GT (1998) Adhesion and growth of a Bacillus sp. on a polyesterurethane. Int Biodeterio Biodegrad 42:63–73
Bouvet PJM, Grimont PAD (1986) Taxonomy of the genus Acinetobacter with the recognition of Acinetobacter baumannii sp. nov. Acinetobacter haemolyticus sp. nov. Acinetobacter johnsonii sp. nov. and Acinetobacter junii sp. nov. and emended descriptions of Acinetobacter calcoaceticus and Acinetobacter lwofii. Int J Syst Bacteriol 36:228–240
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Carr EL, Kampfer P, Patel BKC, Gurtler V, Seviour RJ (2003) Seven novel species of Acinetobacter isolated from activated sludge. Int J Syst Evol Microbiol 53:953–963
Crabbe JR, Campbell JR, Thompson L, Walz SL, Schultz WW (1994) Biodegradation of a collodial ester-based polyurethane by soil fungi. Int Biodeterio Biodegrad 33:103–113
DeSantis TZ, Hugenholtz P, Keller K, Brodie EL, Larsen N, Piceno YM, Phan R (2006) A server for comparative analysis of 16 s rRNA genes. Nucleic Acids Res 34:394–399
Deshpande MV, Eriksson KE, Peterson LG (1984) An assay for selective determination of exo-1,4-β-glucanases in a mixture of cellulolytic enzymes. Anal Biochem 138:481–487
El-Sayed AHMM, Mahmoud WM, Davis EM, Coughlin RW (1996) Biodegradation of polyurethane coatings by hydrocarbon-degrading bacteria. Int Biodeterio Biodegrad 37:69–79
Fried JR (1995) Polymer science and technology. Prentice Hall PTR, Englewood Cliffs
Gerner-Smidt P (1992) Ribotyping of the Acinetobacter calcoaceticus-Acinetobacter baumannii complex. J Clin Microbiol 30:2680–2685
Gerner-Smidt P, Tjernberg I, Ursing J (1991) Reliability of phenotypic tests for identification of Acinetobacter species. J Clin Microbiol 29:277–282
Han SJ, Back JH, Yoon MY, Shin PK, Cheong CS, Sung MH, Hong SP, Chung I, Han YS (2003) Expression and characterization of a novel enantioselective lipase from Acinetobacter species SY-01. Biochimie 85:501–510
Howard GT, Blake RC (1999) Growth of Pseudomonas fluorescens on a polyester-polyurethane and the purification and characterization of a polyurethanase-protease enzyme. Int Biodeterio Biodegrad 42:213–220
Howard GT, Ruiz C, Hilliard N (1998) Growth of Pseudomonas chlororaphis on a polyester-polyurethane and the purification and characterization of a polyurethanase-esterase enzyme. Int Biodeterio Biodegrad 43:7–12
Howard GT, Crother B, Vicknair J (2001) Cloning, nucleotide sequencing and characterization of a polyurethanase gene (pueB) from Pseudomonas chlororaphis. Int Biodeterio Biodegrad 47:141–149
Howard GT, Duos B, Watson E (2010) Characterization of the soil microbial community associated with the decomposition of a swine carcass. Int Biodeterio Biodegrad 64:300–304
Ibrahim A, Gerner-Smidt P, Liesack W (1997) Phylogenetic relationship of the twenty-one DNA groups the genus Acinetobacter as revealed by 16 s ribosomal DNA sequence analysis. J Bacterial 47:837–841
Ishii S, Koki J, Unno H, Hori K (2004) Two morphological types of cell appendages on a strongly adhesive bacterium, Acinetobacter sp. Strain Tol 5. Appl Environ Microbiol 70:5026–5029
Jaeger KE, Ransac S, Dijkstra BW, Colson C, Van Heuvel M, Misset O (1994) Bacterial lipases. FEMS Microbiol Rev 15:29–63
Kawai F (1995) Breakdown of plastics and polymers by microorganisms. Adv Biochem Eng Biotechnol 52:151–194
Knight GC, Seviouri RJ, Soddell OJA, McDonnell S, Bayl RC (1995) Metabolic variation among strains of Acinetobacter isolated from activated sludge. Water Res 29:2081–2084
Kok RG, Christoffels VM, Vosman B, Hellingwerf KJ (1993) Growth-phase dependent expression of the lipolytic system of Acinetobacter calcoaceticus BD413: cloning of a gene encoding one of the esterases. J Gen Microbiol 139:2329–2342
Kok RG, van Thor JJ, Nugteren-Roodzant IM, Brouwer MBW, Egmond MR, Nudel CB, Vosman B, Hellingwerf KJ (1995a) Characterization of the extracellular lipase, LipA, of Acinetobacter calcoaceticus BD413 and sequence analysis of the cloned structural gene. Mol Microbiol 15:803–818
Kok RG, van Thor JJ, Nugteren-Roodzant IM, Vosman B, Hellingwerf KJ (1995b) Characterization of lipase-deficient mutants of Acinetobacter calcoaceticus BD413: identification of a periplasmic lipase chaperone essential for the production of extracellular lipase. J Bacteriol 177:3295–3307
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature (London) 227:680–685
Nakajima-Kambe T, Onuma F, Kimpara N, Nakahara T (1995) Isolation and characterization of a bacterium which utilizes polyester polyurethane as a sole carbon and nitrogen source. FEMS Microbiol Lett 129:39–42
Nakajima-Kambe T, Onuma F, Akutsu Y, Nakahara T (1997) Determination of the polyester polyurethane breakdown products and distribution of the polyurethane degrading enzyme of Comamonas acidovorans strain TB-35. J Ferment Bioeng 83:456–460
Peleg AY, Seifert H, Paterson DL (2008) Acinetobacter baumannii: emergence of a successful pathogen. Clin Microbiol Rev 21:538–582
Persson B, Bentsson-Olivecrona G, Enerback S, Olivecrona T, Jornvall H (1989) Structure features of lipoprotein lipase: lipase family relationships, binding interactions, non-equivalence of lipase cofactors, vitellogenin similarities and functional subdivision of lipoprotein lipase. Eur J Biochem 179:39–45
Rowe L, Howard GT (2002) Growth of Bacillus subtilis on polyurethane and the purification and characterization of a polyurethanase-lipase enzyme. Int Biodeterio Biodegrad 50:33–40
Saunders JH, Frisch KC (1964) Polyurethanes: chemistry and technology, part II: technology. Interscience Publishers, New York
Seifert H, Strate A, Schulze A, Pulverer G (1993) Vascular catheterrelated bloodstream infection due to Acinetobacter johnsonii (formerly Acinetobacter calcoaceticus var. lwoffii): report of 13 cases. Clin Infect Dis 17:632–636
Shannon MJR, Unterman R (1993) Evaluating bioremediation: distinguishing fact from fiction. Ann Rev Microbiol 47:715–738
Stern RS, Howard GT (2000) The polyester polyurethanase gene (pueA) from Pseudomonas chlororaphis encodes a lipase. FEMS Microbiol Lett 185:163–168
Sullivan ER, Leahy JG, Colwell RR (1999) Cloning and sequence analysis of the lipase and lipase chaperone encoding genes from Acinetobacter calcoaceticus RAG-1, and redefinition of a proteobacterial lipase family and an analogous lipase chaperone family. Gene 230:277–285
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Towner KJ (1997) Clinical importance and antibiotic resistance of Acinetobacter spp. J Med Microbiol 46:721–746
Uhlig K (1999) Discovering polyurethanes. Hanser Publisher, Munich
Urbanski J, Czerwinski W, Janicka K, Majewska F, Zowall H (1977) Handbook of analysis of synthetic polymers and plastics. Ellis Horwood Limited, Chichester
Vega R, Main T, Howard GT (1999) Cloning and expression in Escherichia coli of a polyurethane-degrading enzyme from Pseudomonas fluorescens. Int Biodeterio Biodegrad 43:49–55
Winkler FK, D’Arcy A, Hunzinger W (1990) Structure of human pancreatic lipase. Nature (London) 343:7–13
Acknowledgments
This research was funded by the National Science Foundation (EPSCoR #53279) and by the Southeastern Louisiana University Research Initiative Program.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Howard, G.T., Norton, W.N. & Burks, T. Growth of Acinetobacter gerneri P7 on polyurethane and the purification and characterization of a polyurethanase enzyme. Biodegradation 23, 561–573 (2012). https://doi.org/10.1007/s10532-011-9533-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-011-9533-6