Abstract
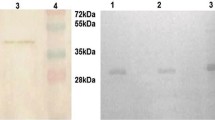
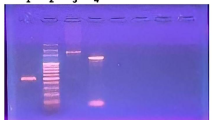
Streptokinase (SK), the heterogeneous protein family secreted by some groups of β-hemolytic streptococci (βHS), is a plasminogen activator and well-known drug for thrombolytic therapy. Differences in plasminogen activation property of streptococcal culture supernatants (SCS) have been traditionally used to identify superior producer strains and SK genes (skc) for recombinant SK (rSK) production. However, the role of SK heterogeneity and whether SK activities in SCS correlate with that of their corresponding rSK is a matter of debate. To address these concerns, SCS of nine group C streptococci (GCS) screened among 252 βHS clinical isolates were compared for plasminogen activation using S-2251 chromogenic assay. The GCS (Streptococcus equisimilis) showing the highest (GCS-S87) and lowest (GCS-S131) activities were selected for PCR-based isolation of skc, cloning and rSK production in Escherichia coli. The 6×His-tagged rSK proteins were purified by NI–NTA chromatography, analyzed by SDS-PAGE and Western blotting and their activities were determined. While SCS of GCS-S87 and GCS-S131 showed different plasminogen activations (95 and 35 %, respectively) compared to that of the reference strain (GCS-9542), but interestingly rSK of all three strains showed close specific activities (1.33, 1.70, and 1.55 × 104 IU mg−1). Accordingly, SKS87 and SKS131 had more than 90 % sequence identity at the amino acids level compared to SK9542. Therefore, SK heterogeneity by itself may not contribute to the differences in plasminogen activation properties of SCS and evaluation of this activity in SCS might not be a proper assay for screening superior skc.



Similar content being viewed by others
References
Abdelghani TTA, Kunamneni A, Ellaiah P (2005) Isolation and mutagenesis of streptokinase-producing bacteria. Am J Immunol 1(4):125–129
Al Sohaimy S, Aleem E, Hafez EE, Esmail SS, El-Saadani M, Moneim NA (2011) Expression of recombinant Streptokinase from local Egyptian Streptococcus sp SalMarEg. Afr J Biotechnol 10(45):9001–9011
Arabi R, Roohvand F, Norouzian D, Sardari S, Aghasadeghi MR, Khanahmad H, Memarnejadian A, Motevalli F (2011) A comparative study on the activity and antigenicity of truncated and full-length forms of streptokinase. Pol J Microbiol 60(3):243–251
Banerjee A, Chisti Y, Banerjee UC (2004) Streptokinase–a clinically useful thrombolytic agent. Biotechnol Adv 22(4):287–307
Chaudhary A, Vasudha S, Rajagopal K, Komath SS, Garg N, Yadav M, Mande SC, Sahni G (1999) Function of the central domain of streptokinase in substrate plasminogen docking and processing revealed by site-directed mutagenesis. Protein Sci 8(12):2791–2805
Cherish Babu P, Srinivas V, Krishna Mohan V, Krishna E (2008) Renaturation purification and characterization of streptokinase expressed as inclusion body in recombinant. E coli J Chromatogr B 861(2):218–226
Churchward G, Bates C, Gusa AA, Stringer V, Scott JR (2009) Regulation of streptokinase expression by CovR/S in Streptococcus pyogenes: covR acts through a single high-affinity binding site. Microbiology 155(2):566–575
Doss HM, Manohar M, Singh NA, Mohanasrinivasan V, Devi CS (2011) Studies on isolation, screening and strain improvement of streptokinase-producing hemolytic streptococci. WJST 1(03):07–11
Garrity GM, Brenner DJ, Krieg NR (2005) Bergey’s manual of systematic bacteriology, vol 2. Springer, Berlin Heidelberg New York
Gräfe S, Ellinger T, Malke H (1996) Structural dissection and functional analysis of the complex promoter of the streptokinase gene from Streptococcus equisimilis H46A. Med Microbiol Immunol 185(1):11–17
Huang TT, Malke H, Ferretti JJ (1989) Heterogeneity of the streptokinase gene in group a streptococci. Infect Immun 57(2):502–506
Hughes JM, Wilson ME, Brandt CM, Spellerberg B (2009) Human infections due to Streptococcus dysgalactiae subspecies equisimilis. Clin Infect Dis 49(5):766–772
Keramati M, Roohvand F, Eslaminejad Z, Mirzaie A, Nikbin VS, Aslani MM (2012) PCR/RFLP-based allelic variants of streptokinase and their plasminogen activation potencies. FEMS Microbiol Lett 335:79–85
Kim DM, Lee SJ, Kim IC, Kim ST, Byun SM (2000) Asp41-His48 region of streptokinase is important in binding to a substrate plasminogen. Throm Res 99(1):93–98
Kim MR, Choeng YH, Chi WJ, Kang DK, Hong SK (2010) Heterologous production of streptokinase in secretory form in Streptomyces lividans and in nonsecretory form in Escherichia coli. J Microbiol Biotechnol 20(1):132–137
Kreikemeyer B, Boyle MDP, Buttaro BAL, Heinemann M, Podbielski A (2001) Group A streptococcal growth phase-associated virulence factor regulation by a novel operon (Fas) with homologies to two component type regulators requires a small RNA molecule. Mol Microbiol 39(2):392–406
Kunamneni A, Abdelghani TTA, Ellaiah P (2007) Streptokinase—the drug of choice for thrombolytic therapy. J Thromb Thrombolysis 23(1):9–23
Lizano S, Johnston KH (2005) Structural diversity of streptokinase and activation of human plasminogen. Infect Immun 73(7):4451–4453
Malke H, Steiner K (2004) Control of streptokinase gene expression in group A & C streptococci by two-component regulators. Indian J Med Res 119:48–56
Malke H, Steiner K, Gase K, Frank C (2000) Expression and regulation of the streptokinase gene. Methods 21(2):111–124
McArthur JD, McKay FC, Ramachandran V, Shyam P, Cork AJ, Sanderson-Smith ML, Cole JN, Ringdahl U, Sjöbring U, Ranson M (2008) Allelic variants of streptokinase from Streptococcus pyogenes display functional differences in plasminogen activation. The FASEB J 22(9):3146–3154
Pupo E, Baghbaderani BA, Lugo V, Fernández J, Páez R, Torréns I (1999) Two streptokinase genes are expressed with different solubility in Escherichia coli W3110. Biotechnol Lett 21(12):1119–1123
Sambrook J, Russell DW (2006) The condensed protocols from molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York
Sikri N, Bardia A (2007) A history of streptokinase use in acute myocardial infarction. Tex Heart Inst J 34(3):318–327
Steiner K, Malke H (2002) Dual control of streptokinase and streptolysin S production by the covRS and fasCAX two-component regulators in Streptococcus dysgalactiae subsp. equisimilis. Infect Immun 70(7):3627–3636
Sundram V, Nanda JS, Rajagopal K, Dhar J, Chaudhary A, Sahni G (2003) Domain truncation studies reveal that the streptokinase-plasmin activator complex utilizes long range protein–protein interactions with macromolecular substrate to maximize catalytic turnover. J Biol Chem 278(33):30569–30577
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24(8):1596–1599
Tewodros W, Norgren M, Kronvall G (1995) Streptokinase activity among group a streptococci in relation to streptokinase genotype, plasminogen binding, and disease manifestations. Microbial Pathog 18(1):53–65
Tharp AC, Laha M, Panizzi P, Thompson MW, Fuentes-Prior P, Bock PE (2009) Plasminogen substrate recognition by the streptokinase-plasminogen catalytic complex is facilitated by Arg253, Lys256, and Lys257 in the streptokinase-domain and kringle 5 of the substrate. J Biol Chem 284(29):19511–19521
Wohl RC, Summaria L, Robbins K (1980) Kinetics of activation of human plasminogen by different activator species at pH 7.4 and 37 degrees C. J Biol Chem 255(5):2005–2013
Yadav S, Aneja R, Kumar P, Datt M, Sinha S, Sahni G (2011) Identification through combinatorial random and rational mutagenesis of a substrate-interacting exosite in the domain of streptokinase. J Biol Chem 286(8):6458–6469
Acknowledgments
This work was financially supported by Pasteur Institute of Iran (PII). MK received a fellowship from graduate school of PII to pursue this study in partial fulfillment of her PhD thesis.
Conflict of interest
The authors declare that they have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Keramati, M., Roohvand, F., Aslani, M.M. et al. Pitfalls in screening streptococci for retrieving superior streptokinase (SK) genes: no activity correlation for streptococcal culture supernatant and recombinant SK. J Ind Microbiol Biotechnol 40, 151–158 (2013). https://doi.org/10.1007/s10295-012-1205-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-012-1205-y