Abstract
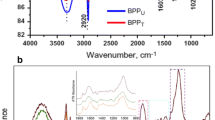
Morphologically different, three bacterial strains, capable of decolorizing Reactive Blue 59 were isolated from dye effluent contaminated soil sample, collected from Ichalkaranji, India. The individual bacterial strains viz. Bacillus odysseyi SUK3, Morganella morganii SUK5 and Proteus sp. SUK7 decolorized Reactive Blue 59 (50 mg l−1) completely within 60, 30, 24 h, respectively, while the bacterial consortium PMB11 of these strains required 3 h for the complete decolorization. The decolorization was confirmed by UV–Vis spectroscopy. Further, the biodegradation of Reactive Blue 59 in to different metabolites was confirmed by High performance liquid chromatography and Fourier transform infrared spectroscopy analysis. Significant increase in the activity of aminopyrine N-demethylase (AND) in the individual as well consortium cells, obtained after decolorization showed involvement of AND in the decolorization process. Phytotoxicity studies, revealed the nontoxic nature of the degraded metabolites of Reactive Blue 59 indicating effectiveness of bacterial consortium PMB11 for the treatment of textile effluent containing Reactive Blue 59.




Similar content being viewed by others
References
Aksu Z (2003) Reactive dye bioaccumulation by Saccharomyces cerevisiae. Process Biochem 38:1437–1444. doi:10.1016/S0032-9592(03)00034-7
Asgher M, Shah SAH, Ali M, Legge RL (2006) Decolorization of some reactive textile dyes by white rot fungi isolated in Pakistan. World J Microbiol Biotechnol 22:89–93. doi:10.1007/s11274-005-5743-6
Banat IM, Nigam P, Singh D, Marchant R (1996) Microbial decolorization of textile dye containing effluents, a review. Bioresour Technol 58:217–227. doi:10.1016/S0960-8524(96)00113-7
Cha C, Doerge DR, Cerniglia CE (2001) Biotransformation of malachite green by the fungus Cunninghamella elegans. Appl Environ Microbiol 67:4358–4360. doi:10.1128/AEM.67.9.4358-4360.2001
Chen KC, Jane YW, Liou DJ, Hwang SCJ (2003) Decolorization of the textile dyes by newly isolated bacterial strains. J Biotechnol 101:57–68. doi:10.1016/S0168-1656(02)00303-6
Chivukula M, Spadaro JT, Renganathan V (1995) Lignin peroxidase catalyzed oxidation of sulfonated azo dyes generates novel sulfophenyl hydroperoxides. Biochemistry 34:7765–7772. doi:10.1021/bi00023a024
Coughlin MF, Kinkle BK, Tepper A, Bishop PL (1997) Characterization of aerobic azo dye degrading bacteria and their activity in biofilms. Water Sci Technol 36:215–220. doi:10.1016/S0273-1223(97)00327-2
Cuoto SR, Rosales E, Sanroman MA (2006) Decolorization of synthetic dyes by Trametes hirsuta in expanded-bed reactors. Chemosphere 62:1558–1563
Da Silva CG, Faria JL (2003) Photochemical and photocatalytic degradation of an azo dye in aqueous solution by UV irradiation. J Photochem Photobiol Chem 155:133–143. doi:10.1016/S1010-6030(02)00374-X
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evol Int J Org Evol 39:783–791. doi:10.2307/2408678
Forgacs E, Cserhati T, Oros G (2004) Removal of synthetic dyes from wastewaters: a review. Environ Int 30:953–971. doi:10.1016/j.envint.2004.02.001
Hatvani N, Mecs I (2001) Production of laccase and manganese peroxidase by Lentinus edodes on malt containing byproduct of the brewing process. Process Biochem 37:491–496. doi:10.1016/S0032-9592(01)00236-9
Haug W, Schmidt A, Nortemann B, Hempel DC, Stolz A, Knackmuss HJ (1991) Mineralization of the sulfonated azo dye Mordant Yellow 3 by a 6-aminonaphthalene-2-sulfonate-degrading bacterial consortium. Appl Environ Microbiol 57:3144–3149
He F, Hu W, Li Y (2004) Biodegradation mechanisms and kinetics of azo dye 4BS by a microbial consortium. Chemosphere 57:293–301. doi:10.1016/j.chemosphere.2004.06.036
Hu T, Wu SC (2001) Assessment of the effect of azo dye RP2B on the growth of nitrogen fixing cyanobacterium Anabena sp. Bioresour Technol 77:93–95. doi:10.1016/S0960-8524(00)00124-3
Itoh K, Kitade Y, Kobayashi S, Nakanishi M, Yatome C (1998) Demethylation of acridine orange by Arthrobacter globiformis. Bull Environ Contam Toxicol 60:781–785. doi:10.1007/s001289900694
Itoh K, Kitade Y, Nakanishi M, Yatome C (2002) Decolorization of Methyl red by a mixed culture of Bacillus sp. and Pseudomonas stutzeri. J Environ Sci Health 37:415–421. doi:10.1081/ESE-120002838
Jadhav JP, Govindwar SP (2006) Biotransformation of malachite green by Saccharomyces cerevisiae MTCC 463. Yeast 23:315–323. doi:10.1002/yea.1356
Johnson RF, Zenhausen A, Zollinger H (1978) In: Mark HF, Mcketta JJ, Othmer DF Jr, Standen A (eds) Krik-Othmer, 2nd edn. Encyclopedia of chemical technology, vol 2. Wiley, Hoboken, pp 868–910
Junnarkar N, Murty DS, Bhatt NS, Madamwar D (2006) Decolorization of diazo dye Direct Red 81 by a novel bacterial consortium. World J Microbiol Biotechnol 22:163–168. doi:10.1007/s11274-005-9014-3
Kalme SD, Parshetti GK, Jadhav SU, Govindwar SP (2007) Biodegradation of benzidine based dye Direct Blue-6 by Pseudomonas desmolyticum NCIM 2112. Bioresour Technol 98:1405–1410. doi:10.1016/j.biortech.2006.05.023
Kapanen A, Itavaara M (2001) Ecotoxicity tests for compost applications. Ecotoxicol Environ Saf 49:1–16. doi:10.1006/eesa.2000.1927
Knapp JS, Newby PS (1995) The microbiological decolorization of an industrial effluent containing a diazo linked chromophore. Water Res 29:1807–1809. doi:10.1016/0043-1354(94)00341-4
Liu S, Suffita JM (1993) Ecology and evolution of microbial populations for bioremediation. Trends Biotechnol 11:344–352. doi:10.1016/0167-7799(93)90157-5
Mohandass R, Bhaskar A, Kalavathy S, Devilaksmi S (2007) Biodecolorization and biodegradation of Reactive Blue by Aspergillus sp. Afr J Biotechnol 6:1441–1445
Moller P, Wallin H (2000) Genotoxic hazards of azo pigments and other colorants related to 1-phenylazo-2-hydroxynaphthalene. Mutat Res 462:13–30. doi:10.1016/S1383-5742(99)00090-3
Moosvi S, Keharia H, Madamwar D (2005) Decolorization of textile dye Reactive Violet by a newly isolated bacterial consortium RVM 11.1. World J Microbiol Biotechnol 21:667–672. doi:10.1007/s11274-004-3612-3
Moosvi S, Kher X, Madamwar D (2007) Isolation characterization and decolorization of textile dyes by a mixed bacterial consortium JW-2. Dyes Pigments 74:723–729. doi:10.1016/j.dyepig.2006.05.005
Nigam P, Banat IM, Singh D, Marchant R (1996) Microbial process for the decolorization of textile effluent containing azo, diazo and reactive dyes. Process Biochem 31:435–442. doi:10.1016/0032-9592(95)00085-2
Nyanhongo GS, Gomes J, Guebitz GM, Zvauya R, Read J, Steiner W (2002) Decolorization of textile dyes by laccases from a newly isolated strain of Trametes modesta. Water Res 36:1449–1456. doi:10.1016/S0043-1354(01)00365-7
O’Neill C, Lopez A, Esteves S, Hawkes FR, Hawkes DL, Wilcox S (2000) Azo-dye degradation in an anaerobic–aerobic treatment system operating on simulated textile effluent. Appl Microbiol Biotechnol 53:249–254. doi:10.1007/s002530050016
Okazaki S, Nagasawa S, Goto M, Furusaki S, Wariishi H, Tanaka H (2002) Decolorization of azo and anthraquinone dyes in hydrophobic organic media using microperoxidase-11 entrapped in reversed micelles. Biochem Eng J 12:237–241. doi:10.1016/S1369-703X(02)00074-8
Parshetti GK, Kalme SD, Saratale GD, Govindwar SP (2006) Biodegradation of Malachite green by Kocuria rosea MTCC 1532. Acta Chim Slov 53:492–498
Paszcezynski A, Pasti-Grigsby M, Goszceynski S, Crawford R, Crawford DL (1992) Mineralization of sulfonated azo dyes and sulfanilic acid by Phanerochaete chrysosporium and Streptomyces chromofuscus. Appl Environ Microbiol 58:3598–3604
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Salokhe MD, Govindwar SP (1999) Effect of carbon source on the biotransformation enzymes in Serratia marcescens. World J Microbiol Biotechnol 15:259–263. doi:10.1023/A:1008875404889
Senan RC, Abraham TE (2004) Bioremediation of textile azo dyes by aerobic bacterial consortium. Biodegradation 15:275–280. doi:10.1023/B:BIOD.0000043000.18427.0a
Shanmugam V, Kumari M, Yadav KD (1999) n-Propanol as a substrate for assaying the lignin peroxidase activity of Phaenerochaete chrysosporium. Indian J Biochem Biophys 36:39–43
Takezaki N, Rzhetsky A, Nei M (2004) Phylogenetic test of the molecular clock and linearized trees. Mol Biol Evol 12:823–833
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci USA 101:11030–11035. doi:10.1073/pnas.0404206101
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599. doi:10.1093/molbev/msm092
Verma P, Madamwar D (2003) Decolorization of synthetic dyes by a newly isolated strain of Serratia marcescens. World J Microbiol Biotechnol 19:615–618. doi:10.1023/A:1025115801331
Vijaya PP, Sandhya S (2003) Decolorization and complete degradation of methyl red by a mixed culture. Environmentalist 23:145–149. doi:10.1023/A:1024839805387
Watanabe K, Baker PW (2000) Environmentally relevant microorganisms. J Biosci Bioeng 89:1–11. doi:10.1016/S1389-1723(00)88043-3
Yatome C, Yamada S, Ogawa T, Matsui M (1993) Degradation of crystal violet by Nocardia coralline. Appl Microbiol Biotechnol 38:565–569
Zhang X, Flurkey W (1997) Phenoloxidases in Portabella mushrooms. J Food Sci 62:97–100. doi:10.1111/j.1365-2621.1997.tb04376.x
Acknowledgments
Ms P.S. Patil one of the authors is thankful to Department of Microbiology, Shivaji University, Kolhapur for awarding the Departmental Research Fellowship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Patil, P.S., Shedbalkar, U.U., Kalyani, D.C. et al. Biodegradation of Reactive Blue 59 by isolated bacterial consortium PMB11. J Ind Microbiol Biotechnol 35, 1181–1190 (2008). https://doi.org/10.1007/s10295-008-0398-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-008-0398-6