Abstract
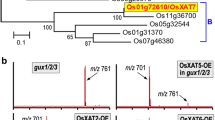
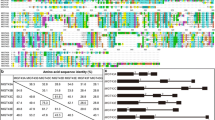
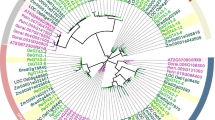
Glucuronoarabinoxylan is the major hemicellulose in grass cell walls, yet the mechanism of xylan synthesis in monocot plants is still unclear. Unraveling the genes involved in the biosynthesis of xylan in rice will be very important for the utilization of rice straw as a source of bioenergy in the future. In this report, we investigated the functional role of a rice gene homologous to Arabidopsis IRREGULAR XYLEM10 (IRX10), belonging to the glycosyl transferase (GT) gene family 47 (GT47), in the biosynthesis of xylan. The protein sequence of OsGT47A from rice exhibits a 93.49 % similarity to IRX10, which is involved in the biosynthesis of glucuronoxylan in Arabidopsis. Phylogenetic analysis of the GT47 glycosyl transferase family in the rice genome revealed that OsGT47A is a closely related homolog of IRX10 and IRX10L. Expression pattern analysis showed that the OsGT47A gene is highly expressed in the rice stem. Overexpression of OsGT47A in the irx10 irx10L double mutant rescued the plant growth phenotype and restored secondary wall thickness. Analysis of monosaccharides indicated that the rescued plants had levels of xylose identical to those of the wild type plants, and the fluorescence signals were restored in the complementation plants by xylan immunolocalization. The OsGT47A complementation under the native promoter of Arabidopsis IRX10L (ProIRX10L) partially rescued the double mutant, indicating that OsGT47A is functionally equivalent to IRX10L. Together, these results suggest that the IRX10 homolog OsGT47A exhibits functional conservation and is most likely involved in xylan synthesis in rice.







Similar content being viewed by others
Abbreviations
- IRX :
-
IRREGULAR XYLEM
- CaMV:
-
Cauliflower mosaic virus
- GAX:
-
Glucuronoarabinoxylan
- GlcA:
-
Glucuronic acid
- GT:
-
Glycosyltransferase
- qRT-PCR:
-
Quantitative real time-PCR
- GX:
-
Glucronoxylan
- PCR:
-
Polymerase chain reaction
- XylT:
-
Xylosyltransferase
References
Anders N, Wilkinson MD, Lovegrove A, Freeman J, Theodora T, Pellny TK, Weimar T, Mortimer JC, Stott K, Baker JM, Defoin-Platel M, Shewry PR, Dupree P, Mitchell RAC (2012) Glycosyl transferases in family 61 mediate arabinofuranosyl transfer on to xylan in grasses. Proc Natl Acad Sci USA 109:989–993
Andersson SI, Samuelson O, Ishihara M, Shimizu K (1983) Structure of the reducing end-groups in spruce xylan. Carbohydr Res 111:283–288
Bauer S, Vasu P, Persson S, Mort AJ, Somerville CR (2006) Development and application of a suite of polysaccharides degrading enzymes for analyzing plant cell walls. Proc Natl Acad Sci USA 103:11417–11422
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546
Brown DM, Zeef LAH, Ellis J, Goodacre R, Turner SR (2005) Identification of novel genes in Arabidopsis involved in secondary cell wall formation using expression profiling and reverse genetics. Plant Cell 17:2281–2295
Brown DM, Goubet F, Vicky WWA, Goodacre R, Stephens E, Dupree P, Turner SR (2007) Comparison of five xylan synthesis mutants reveals new insight into the mechanisms of xylan synthesis. Plant J 52:1154–1168
Brown DM, Zhang Z, Stephens E, Dupree P, Turner SR (2009) Characterization of IRX10 and IRX10-like reveals an essential role in glucuronoxylan biosynthesis in Arabidopsis. Plant J 57:732–746
Cao PJ, Bartley LE, Jung KH, Ronald PC (2008) Construction of a rice glycosyltransferase phylogenomic database and identification of rice-diverged glycosyltransferases. Mol Plant 1:858–877
Carroll A, Somerville C (2009) Cellulosic biofuels. Annu Rev Plant Biol 60:165–182
Chen X, Vega-Sánchez ME, Verhertbruggen Y, Chiniquy D, Canlas PE, Fagerström A, Prak L, Christensen U, Oikawa A, Chern M, Zuo S, Lin F, Auer M, Willats WGT, Bartley L, Harholt J, Scheller HV, Ronald PC (2013) Inactivation of OsIRX10 leads to decreased xylan content in rice culm cell walls and improved biomass saccharification. Mol Plant 6:570–573
Chiniquy D, Sharma V, Schultink A, Baidoo EE, Rautengarten C, Cheng K, Carroll A, Ulvskov P, Harholt J, Keasling JD, Pauly M, Scheller HV, Ronald PC (2012) XAX1 from glycosyltransferase family 61 mediates xylosyltransfer to rice xylan. Proc Natl Acad Sci USA 109:17117–17122
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Earley KW, Haag JR, Pontes O, Opper K, Juehne T, Song K, Pikaard CS (2006) Gateway-compatible vectors for plant functional genomics and proteomics. Plant J 45:616–629
Ebringerová A, Heinze T (2000) Xylan and xylan derivatives—biopolymers with valuable properties. 1. Naturally occurring xylans structures, isolation procedures and properties. Macromol Rapid Commun 21:542–556
Ebringerová A, Heinze T (2005) Hemicellulose. Adv Polym Sci 186:1–67
Englyst HN, Cummings JH (1984a) Simplified method for the measurement of total non-starch polysaccharides by gas-liquid chromatography of constituent sugars as alditol acetates. Analyst (Lond) 109:937–942
Englyst HN, Cummings JH (1984b) Simplified method for the measurement of total non-starch polysaccharides by gas-liquid chromatography of constituent sugars as alditol acetates. Analyst (Lond) 109:937–942
Faik A (2010) Xylan biosynthesis: news from the grass. Plant Physiol 153:396–402
Forsthoefel NR, Wu Y, Schultz B, Bennett MJ, Feldmann KA (1992) T-DNA insertion mutagenesis in Arabidopsis: prospects and perspectives. Aust J Plant Physiol 19:353–366
Freshour G, Clay RP, Fuller MS, Albersheim P, Darvill AG, Hahn MG (1996) Developmental and tissue-specific structural alterations of the cell-wall polysaccharides of Arabidopsis thaliana roots. Plant Physiol 110:1413–1429
Freshour G, Bonin CP, Reiter WD, Albersheim P, Darvill AG, Hahn MG (2003) Distribution of fucose-containing xyloglucans in cell walls of the mur1 mutant of Arabidopsis. Plant Physiol 131:1602–1612
Johansson MH, Samuelson O (1977) Reducing end groups in Birch xylan and their alkaline degradation. Wood Sci Technol 11:251–263
Kong Y, Zhou G, Avic U, Gu X, Jones C, Yin Y, Xu Y, Hahn MG (2009) Two poplar glycosyltransferase genes, PdGATL1.1 and PdGATL1.2 are functional orthologs to PARVUS/AtGATL1 in Arabidopsis. Mol Plant 2:1040–1050
Lee C, O’Neill MA, Tsumuraya Y, Darvill AG, Ye Z-H (2007) The irregular xylem 9 mutant is deficient in xylan xylosyltransferase activity. Plant Cell Physiol 48:1624–1634
Lee C, Teng Q, Huang W, Zhong R, Ye Z-H (2010) The Arabidopsis family GT43 glycosyltransferases from two functionally nonredudant groups essential for elongation of glucuronoxylan backbone. Plant Physiol 153:526–541
Lee C, Zhong R, Ye Z-H (2012) Arabidopsis family GT43 members are xylan xylosyltransferases required for the elongation of the xylan backbone. Plant Cell Physiol 53:135–143
Lerouxel O, Cavalier DM, Liepman AH, Keegstra K (2006) Biosynthesis of plant cell wall polysaccharides—a complex process. Curr Opin Plant Biol 9:621–630
Liepman AH, Wightman R, Geshi N, Turner SR, Scheller HV (2010) Arabidopsis—a powerful model system for plant cell wall research. Plant J 61:1107–1121
Madson M, Christophe D, Li X, Verma R, Vanzin GF, Caplan J, Shoue DA, Carpita NC, Reiter WD (2003) The MUR3 gene of Arabidopsis encodes a xyloglucan galactosyltransferase that is evolutionarily related to animal exostosins. Plant Cell 15:1662–1670
McCartney L, Marcus SE, Knox JP (2005) Monoclonal antibodies to plant cell wall xylans and arabinoxylans. J Histochem Cytochem 53:543–546
Mitchell RA, Dupree P, Shewry PR (2007) A novel bioinformatics approach identifies candidate genes for the synthesis and feruloylation of arabinoxylan. Plant Physiol 144:43–53
Pauly M, Gille S, Liu L, Mansoori N, Souza AD, Schultink A (2013) Hemicellulose biosynthesis. Planta 238:627–642
Pena MJ, Zhong R, Zhou GK, Richardson EA, O’neill MA, Darvill AG, York WS, Ye Z-H (2007) Arabidopsis irregular xylem8 and irregular xylem9: implications for the complexity of glucuronoxylan biosynthesis. Plant Cell 19:549–563
Persson S, Hosmer Caffall K, Freshour G, Hilley MT, Bauer S, Poindexter P, Hahn MG, Mohnen D, Somerville C (2007) The Arabidopsis irregular xylem8 mutant is deficient in glucuronoxylan and homogalacturonan, which are essential for secondary cell wall integrity. Plant Cell 19:237–255
Scheible WR, Pauly M (2004) Glycosyltransferases and cell wall biosynthesis: novel players and insights. Curr Opin Plant Biol 7:285–295
Scheller HV, Ulvskov P (2010) Hemicelluloses. Annu Rev Plant Biol 61:263–289
Somerville CR (2006) Cellulose synthesis in higher plants. Annu Rev Cell Dev Biol 22:53–78
Turner S, Somerville C (1997) Collapsed xylem phenotype of Arabidopsis identifies mutants deficient in cellulose deposition in the secondary cell wall. Plant cell 9:689–701
Wu AM, Rihouey C, Seveno M, Hörnblad E, Singh SK, Matsunaga T, Ishii T, Lerouge P, Marchant A (2009) The Arabidopsis IRX10 and IRX10-LIKE glycosyltransferases are critical for glucuronoxylan biosynthesis during secondary cell wall formation. Plant J 57:718–731
Wu AM, Hornblad E, Voxeur A, Gerber L, Rihouey C, Lerouge P, Marchant A (2010) Analysis of the Arabidopsis IRX9/IRX9-L and IRX14/IRX14-L pairs of glycosyltransferase genes reveals critical contributions to biosynthesis of the hemicellulose glucuronoxylan. Plant Physiol 153:542–554
Zeng W, Jiang N, Nadella R, Killen TL, Nadella V, Faik A (2010) A glucurono(arabino)xylan synthase complex from wheat contains members of the GT43, GT47, and GT75 families and functions cooperatively. Plant Physiol 154:78–97
Zhong R, Ye Z-H (2003) Unraveling the functions of glycosyltransferase family 47 in plants. Trends Plant Sci 8:565–568
Zhong R, Pena MJ, Zhou G-K, Naim CJ, Wood-Jones A, Richardson EA, Morrison WH, Darvill AG, York WS, Ye Z-H (2005) The FRA8 gene which encodes a putative glucuronyltransferase is essential for normal secondary wall synthesis. Plant Cell 17:3390–3408
Zhou GK, Zhong R, Richardson EA, Morrison WH, Nairn CJ, Wood-Jones A, Ye Z-H (2006) The poplar glycosyltransferase GT47C is functionally conserved with Arabidopsis Fragile Fiber8. Plant Cell Physiol 47:1229–1240
Zhou GK, Zhong R, Himmelsbach DS, McPhail BT, Ye Z-H (2007) Molecular characterization of PoGT8D and PoGT43B, two secondary wall-associated glycosyltransferases in poplar. Plant Cell Physiol 48:689–699
Acknowledgments
We thank two anonymous reviewers for their constructive comments and suggestions. This work was supported by the National Natural Science Foundation of China (Grant Number 31170165) and the Natural Science Foundation supported by Guangdong Province (Grant Number S2011010001138).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, B., Zhao, T., Yu, W. et al. Functional conservation of the glycosyltransferase gene GT47A in the monocot rice. J Plant Res 127, 423–432 (2014). https://doi.org/10.1007/s10265-014-0631-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-014-0631-5