Abstract
Low phosphorus (P) tolerance in rice is a biologically and agronomically important character. Low P tolerant Indica-type rice genotypes, Sahbhagi Dhan (SD) and Chakhao Poreiton (CP), are adapted to acidic soils and show variable response to low P levels. Using RNAseq approach, transcriptome data was generated from roots of SD and CP after 15 days of low P treatment to understand differences and similarities at molecular level. In response to low P, number of genes up-regulated (1318) was more when compared with down-regulated genes (761). Eight hundred twenty-one genes found to be significantly regulated between SD and CP in response to low P. De novo assembly using plant database led to further identification of 1535 novel transcripts. Functional annotation of significantly expressed genes suggests two distinct methods of low P tolerance. While root system architecture in SD works through serine-threonine kinase PSTOL1, suberin-mediated cell wall modification seems to be key in CP. The transcription data indicated that CP relies more on releasing its internally bound Pi and coping with low P levels by transcriptional and translational modifications and using dehydration response-based signals. Role of P transporters seems to be vital in response to low P in CP while sugar- and auxin-mediated pathway seems to be preferred in SD. At least six small RNA clusters overlap with transcripts highly expressed under low P, suggesting role of RNA super clusters in nutrient response in plants. These results help us to understand and thereby devise better strategy to enhance low P tolerance in Indica-type rice.







Similar content being viewed by others
References
Akhtar MS, Yoko O, Tadashi A (2008) Intraspecific variations of phosphorus absorption and remobilization, P forms, and their internal buffering in Brassica cultivars exposed to a P-stressed environment. J Integr Plant Biol 50:703–716
Bagnaresi P, Biselli C, Orru L, Urso S, Crispino L, Abbruscato P, Piffanelli P, Lupotto E, Cattivelli L, Vale G (2012) Comparative transcriptome profiling of the early response to Magnaporthe oryzae in durable resistant vs susceptible rice (Oryza sativa L.) genotypes. PLoS One 7, e51609
Bengough AG, McKenzie BM, Hallet PD, Valentine TA (2011) Root elongation, water stress, and mechanical impedance: a review of limiting stresses and beneficial root tip traits. J Exp Bot 62:59–68
Cai HM, Lu YG, Xie WB, Zhu T, Lian XM (2012) Transcriptome response to phosphorus starvation in rice. Acta Physiol Plant 34:327–341
Challam C, Kharshing GA, Yumnam JS, Rai M, Tyagi W (2013) Association of qLTG3-1 with germination stage cold tolerance in diverse rice germplasm from the Indian subcontinent. Plant Genetic Resources 11:206–211
Chiou TJ, Lin SI (2011) Signaling network in sensing phosphate availability in plants. Annu Rev Plant Biol 62:185–206
Dkhar F, Rai M, Tyagi W (2014) Looking beyond PsTOL1: marker development for two novel rice genes showing differential expression in P deficient conditions. J Genet 93:573–577
Gamuyao R, Chin JH, Pariasca-Tanaka J, Pesaresi P, Catausan S, Dalid C, Slamet-Loedin I, Tecson-Mendoza EM, Wissuwa M, Heuer S (2012) The protein kinase Pstol1 from traditional rice confers tolerance of phosphorus deficiency. Nature 488:535–537
Garg R, Verma M, Agrawal S, Shankar R, Majee M, Jain M (2013) Deep transcriptome sequencing of wild halophyte rice, Porteresia coarctata, provides novel insights into the salinity and submergence tolerance factors. DNA Res 42:1–16
Hanin M, Brini F, Ebel C, Toda Y, Takeda S, Masmoudi K (2011) Plant dehydrins and stress tolerance; versatile proteins for complex mechanisms. Plant Signal Behav 10:1503–1509
Hanson WC (1950) The photometric determination of phosphorus in fertilizers using the phosphovanado-molybdate complex. J Sci Food Agric 1:172–173
Heuer S, Lu X, Chin JH, Tanaka JP, Kanamori H, Matsumoto T, De Leon T, Ulat VJ, Ismail AM, Yano M, Wissuwa M (2009) Comparative sequence analysis of the major quantitative trait locus phosphorus uptake 1 (Pup1) reveal a complex genetic structure. Plant Biotechnol J 7:456–471
Jain M, Nijhawan A, Tyagi AK, Khurana JP (2006) Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative realtime PCR. Biochem Biophys Res Commun 345:646–651
Koide Y, Tanaka JP, Rose T, Fukuo A, Konisho K, Yanagihara S, Fukuta Y, Wissuwa M (2013) QTLs for phosphorus deficiency tolerance detected in upland NERICA varieties. Plant Breed 132:259–265
Kotake T, Nakagawa N, Takeda K, Sakurai N (2000) Auxin-induced elongation growth and expressions of cell wall-bound exo- and endo-β-glucanases in barley coleoptiles. Plant Cell Physiol 41:1272–1278
Lata C, Prasad M (2011) Role of DREBs in regulation of abiotic stress responses in plants. J Exp Bot 62:4731–4748
Li Y-F, Luo A-C, Hassan MJ, Wei X-H (2006) Effect of phosphorus deficiency on leaf photosynthesis and carbohydrates partitioning in two rice genotypes with contrasting low phosphorus susceptibility. Rice Sci 13:283–290
Li L, Liu C, Lian X (2010) Gene expression profiles in rice roots under low phosphorus stress. Plant Mol Biol 72:423–432
Lynch JP (2007) Roots of the second green revolution. Aust J Bot 55:493–512
Lynch JP (2011) Root phenes for enhanced soil exploration and phosphorus acquisition: tools for future crops. Plant Physiol 156:1041–1049
Misson J, Raghothama KG, Jain A, Jouhet J, Block MA, Bligny R, Ortet P, Creff A, Somerville S, Rolland N, Doumas P, Nacry P, Herrerra-Estrella L, Nussaume L, Thibaud MC (2005) A genome-wide transcriptional analysis using Arabidopsis thaliana affymetrix gene chips determined plant responses to phosphate deprivation. Proc Natl Acad Sci U S A 102:11934–11939
Niu YF, Chai RS, Jin GL, Wang H, Tang CX, Zhang YS (2013) Responses of root architecture development to low phosphorus availability: a review. Ann Bot 112:391–408
Ohkama-Ohtsu N, Wasaki J (2010) Recent progress in plant nutrition research: cross-talk between nutrients, plant physiology and soil microorganisms. Plant Cell Physiol 51:1255–1264
Postma JA, Lynch JP (2010) Theoretical evidence for the functional benefit of root cortical aerenchyma in soils with low phosphorus availability. Ann Bot 107:829–841
Rausch C, Bucher M (2002) Molecular mechanisms of phosphate transport in plants. Planta 216:23–37
Richardson AE (2009) Regulating the phosphorus nutrition of plants: molecular biology meeting agronomic needs. Plant Soil 322:17–24
Rose TJ, Rose MT, Pariasca-Tanaka J, Heuer S, Wissuwa M (2011) The frustration with utilization: why have improvements in internal phosphorus utilization efficiency in crops remained so elusive? Front Plant Sci 2:73
Rouached H, Arpat AB, Poirier Y (2010) Regulation of phosphate starvation responses in plants: signaling players and cross-talks. Mol Plant 3:288–299
Song M, Xu W, Xiang Y, Jia H, Zhang L, Ma Z (2014) Association of jacalin-related lectins with wheat responses to stresses revealed by transcriptional profiling. Plant Mol Biol 84:95–110
Su CF, Wang YC, Hsieh TH, Lu CA, Tseng TH, Yu SM (2010) A novel MYBS3-dependent pathway confers cold tolerance in rice. Plant Physiol 153:145–158
Tiwari K, Singh A, Pattanaik S, Sandhu M, Kaur S, Jain S, Tiwari S, Mehrotra S, Anumalla M, Samal R et al (2015) Identification of a diverse mini-core panel of Indian rice germplasm based on genotyping using microsatellite markers. Plant Breed. doi:10.1111/pbr.12252
Tyagi W, Rai M, Dohling A (2012) Haplotype analysis for Pup1 locus in rice genotypes of north eastern and Eastern India to identify suitable donors tolerant to low phosphorus. SABRAO J Breed Genet 44:398–405
Wasaki J, Yonetani R, Kuroda S, Shinano T, Yazaki J, Fuji F, Shimbo K, Yamamoto K, Sakata K, Sasaki T, Kishimoto N, Kikuchi S, Yamagishi M, Osaki M (2003) Transcriptomic analysis of metabolic changes by phosphorus stress in rice plant roots. Plant Cell Environ 26:1515–1523
Wasaki J, Shinano T, Onishi K, Yonetani R, Yazaki J, Fujii F, Shimbo K, Ishikawa M, Shimatani Z, Nagata Y, Hashimoto A, Ohta T, Sato Y, Miyamoto C, Honda S, Kojima K, Sasaki T, Kishimoto N, Kikuchi S, Osaki M (2006) Transcriptomic analysis indicates putative metabolic changes caused by manipulation of phosphorus availability in rice leaves. J Exp Bot 57:2049–2059
Wissuwa M, Ae N (2001) Genotypic variation for tolerance to phosphorus deficiency in rice and the potential for its exploitation in rice improvement. Plant Breed 120:43–48
Wissuwa M, Yano M, Ae N (1998) Mapping for phosphorus-deficiency tolerance in rice (Oryza sativa L.). Theor Appl Genet 97:777–783
Yi K, Wu Z, Zhou J, Du L, Guo L, Wu Y, Wu P (2005) OsPTF1, a novel transcription factor involved in tolerance to phosphate starvation in rice. Plant Physiol 138:2087–2096
Yoshida S, Forno DA, Cook JH, Gomez KA (1976) Laboratory manual for physiological studies of rice. International Rice Research Institute, Manila, pp 61–67
Yumnam JS, Rai M, Tyagi W (2015) Allele mining across PSTOL1 and Dirigent-like genes reveals novel haplotypes in rice genotypes adapted to acidic soils. Plant Genetic Resources. doi:10.1017/S1479262115000544
Acknowledgments
This work was supported by grant from Indian Council of Agricultural Research, National Agricultural Innovative Project [C30033/415101-036 to W.T.]. We are grateful to Drs. T. Mohapatra and A.K. Singh, NAIP, for helpful suggestions during initial design of the experiment and for sharing of seeds of SD, respectively. We are also grateful to Dr. P. Dasaradhi for fruitful discussions. Julia S. Yumnam and Aibanshan K. Dohling are acknowledged for technical assistance.
Note: The RNAseq data reported are available in NCBI sequence read archive under the accession number SRP058964 (PRJNA285664).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Bhumi Nath Tripathi
Supplementary materials
Below is the link to the electronic supplementary material.
Figure S1
Experimental design and bioinformatics flow. (PPT 168 kb)
Figure S2
cDNA library bioanalyzer profile on high sensitivity chip of the four samples. (PPT 422 kb)
Figure S3
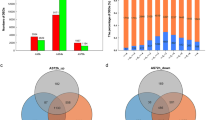
Venn diagram showing the number of genes expressed under different conditions in SD (a) and CP (b). Venn diagrams display the overlap between expressed genes in roots following treatment with low P (green circles, right) compared with control (peach circles, left). The total gene count for each category is shown within each circle. (PPT 186 kb)
Figure S4
Volcano plot showing significantly different genes between the two genotypes, SD and CP. (PPT 126 kb)
Figure S5
K-means clustering of differentially expressed genes under low P conditions into five distinct clusters. (PPT 142 kb)
Figure S6
Global overview and hierarchical clustering of Sahbhagi Dhan (SD) and Chakhao Poreiton (CP) transcriptional changes. Mean expression versus log fold change plots (MA-plots) were computed for significantly expressed transcripts in SD and CP; normalized expression mean values are plotted versus log2 fold changes and called DEGs (FDR<0.05) are plotted in red. Bar at the top represents log2 transformed values, thereby values 0, 40 and 200 represent low, intermediate and high expression, respectively (PPT 388 kb)
Figure S7
Selected genomic origins showing target for small RNAs. The potential target transcript is written at the right side of each snapshot with locus ID (LOC_). The number of individual clusters within a region is indicated on top of the line. (PPT 244 kb)
Table S1
List of transcripts differentially regulated in SD and CP under low P conditions. (XLS 640 kb)
Table S2
List of GOSlim term assignment to SD and CP transcripts in different categories of biological process under control (C; normal P levels) and treatment conditions (T; low P levels). (XLS 204 kb)
Table S3
The mapping statistics of SD and CP treated samples on Kasalath BAC clones. (DOC 26 kb)
Table S4
Denovo assembly of unused reads for genotype, SD. (XLS 2104 kb)
Table S5
Denovo assembly of unused reads for genotype, CP. (XLS 7542 kb)
Table S6
List of functionally annotated transcripts based on denovo assembly. (XLS 240 kb)
Table S7
List and functional annotation of highly upregulated transcripts under low P conditions in genotypes, SD and CP. (XLS 33 kb)
Table S8
Log2 fold expression of putative copper oxidase transcripts under low P conditions in genotypes, SD and CP. (XLS 185 kb)
Table S9
List of cis-acting elements. (TXT 1 kb)
Rights and permissions
About this article
Cite this article
Tyagi, W., Rai, M. Root transcriptomes of two acidic soil adapted Indica rice genotypes suggest diverse and complex mechanism of low phosphorus tolerance. Protoplasma 254, 725–736 (2017). https://doi.org/10.1007/s00709-016-0986-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-016-0986-7