Abstract
We present an improved method for the selection of single-stranded DNA aptamers that can recognize fumonisin B1 (FB1). FB1 is a carcinogenic mycotoxin mainly found in corn and corn-based food products worldwide, posing a global threat to feed and food safety. Selection was based on the mag-SELEX (magnetic bead systematic evolution of ligands by exponential enrichment) technology modified by adopting free analogs of targets rather than immobilized targets for counter selections. Firstly, aptamer candidates for FB1 were selected from an 80 nt random DNA library after 13 rounds of selection. Next, binding assays were performed for affinity evaluation, and circular dichroism spectroscopy was used to investigate their conformation. A high-affinity aptamer designated as F10 (with a dissociation constant of 62 ± 5 nM) was identified and tested for its specificity by competitive binding assays. The results demonstrate that this improved mag-SELEX technology facilitates aptamer screening because it avoids the tedious immobilization of counter-selection molecules on magnetic beads. The aptamers obtained by this technique open new possibilities for the detection of FB1 via aptasensors.

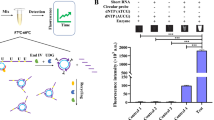
Illustration of SELEX selection procedure. An improved method for selection of ssDNA aptamers recognizing fumonisin B1 was developed based on magnetic beads. Free analogs of targets rather than immobilized were used for counter selections, facilitating the selection and leading to affinity-raised aptamers.






Similar content being viewed by others
References
Ross PF, Nelson PE, Richard JL, Osweiler GD, Rice LG, Plattner RD (1990) Production of fumonisins by Fusarium moniliforme and Fusarium proliferatum isolates associated with equine leukoencephalomalacia and a pulmonary edema syndrome in swine. Appl Environ Microbiol 56:3225–3226
Visconti A, Bruno Doko M, Solfrizzo M, Pascale M, Boenke A (1996) European intercomparison study for the determination of fumonisins in maize. Microchim Acta 123:55–61
Mallmann CA, Santurio JM, Almeida CAA, Dilkin P (2001) Fumonisin B1 levels in cereals and feeds from southern Brazil. Arq Inst Biol Sao Paulo 68:41–45
European Commission (2000) Opinion of the scientific committee of food on fusarium toxins part 3: fumonisin B1 (FB1) http://ec.europa.eu/food/fs/sc/scf/out73_en.pdf
International Agency for Research on Cancer (2002) Fumonisin B1. In: IARC monographs on the evaluation of carcinogenic risks to humans: some traditional herbal medicines, some mycotoxins, naphthalene and styrene. IARC, Lyon, France, 82: 301–366
Shephard GS (1998) Chromatographic determination of the fumonisin mycotoxins. J Chromatogr A 815:31–39
Gazzotti T, Lugoboni B, Zironi E, Barbarossa A, Serraino A, Pagliuca G (2009) Determination of fumonisin B1 in bovine milk by LC–MS/MS. Food Control 20:1171–1174
Gaspardo B, Del Zotto S, Torelli E, Cividino SR, Firrao G, Della Riccia G, Stefanon B (2012) A rapid method for detection of fumonisins B1 and B2 in corn meal using fourier transform near infrared (FT-NIR) spectroscopy implemented with integrating sphere. Food Chem 135:1608–1612
Quan Y, Zhang Y, Wang S, Lee N, Kennedy IR (2006) A rapid and sensitive chemiluminescence enzyme-linked immunosorbent assay for the determination of fumonisin B1 in food samples. Anal Chim Acta 580:1–8
Shiu CM, Wang JJ, Yu FY (2010) Sensitive enzyme-linked immunosorbent assay and rapid one-step immunochromatographic strip for fumonisin B1 in grain-based food and feed samples. J Sci Food Agric 90:1020–1026
Wang S, Quan Y, Lee N, Kennedy IR (2006) Rapid determination of fumonisin B1 in food samples by enzyme-Linked immunosorbent assay and colloidal gold immunoassay. J Agric Food Chem 54:2491–2495
O’Sullivan CK (2002) Aptasensors—the future of biosensing? Anal Bioanal Chem 372:44–48
Ellington AD, Szostak JW (1990) In vitro selection of RNA molecules that bind specific ligands. Nature 346:818–822
Tuerk C, Gold L (1990) Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249:505–510
Wang XF, Zhao Q (2012) A fluorescent sandwich assay for thrombin using aptamer modified magnetic beads and quantum dots. Microchim Acta 178:349–35
Yuan JL, Yu Y, Can L, Ma XY, Xia Y, Chen J, Wang ZP (2014) Visual detection and microplate assay for staphylococcus aureus based on aptamer recognition coupled to tyramine signal amplification. Microchim Acta 181:321–327
Hermann T, Patel DJ (2000) Adaptive recognition by nucleic acid aptamers. Science 287:820–825
Kawakami J, Imanaka H, Yokota Y, Naoki S (2000) In vitro selection of aptamers that act with Zn2+. J Inorg Biochem 82:197–206
Chen XJ, Huang YK, Duan N, Wu SJ, Ma XY, Xia Y, Zhu CQ, Jiang Y, Wang ZP (2013) Selection and identification of ssDNA aptamers recognizing zearalenone. Anal Bioanal Chem 405:6573–6581
Ylera F, Lurz R, Erdmann VA, Furste JP (2002) Selection of RNA Aptamers to the Alzheimer’s disease amyloid peptide. Biochem Bioph Res Commun 290:1583–1588
Bock LC, Griffin LC, Latham JA, Vermaas EH, Toole JJ (1992) Selection of single-stranded DNA molecules that bind and inhibit human thrombin. Nature 355:564–566
Park JW, Lee SJ, Choi EJ, Kim J, Song JY, Gu MB (2014) An ultra-sensitive detection of a whole virus using dual aptamers developed by immobilization-free screening. Biosens Bioelectron 51:324–329
Liu GQ, Yu XF, Xue F, Chen W, Ye YK, Yang XJ, Lian YQ, Yan Y, Zong K (2012) Screening and preliminary application of a DNA aptamer for rapid detection of Salmonella O8. Microchim Acta 178:237–244
Tang ZW, Shangguan DH, Wang KH, Shi K, Sefah K, Mallikratchy P, Chen HW, Li Y, Tan WH (2007) Selection of aptamers for molecular recognition and characterization of cancer cells. Anal Chem 79:4900–4907
Suh SH, Jaykus LA (2013) Nucleic acid aptamers for capture and detection of listeria spp. J Biotechnol 167:454–461
Duan N, Ding XY, He LX, Wu SJ, Wei YX, Wang ZP (2013) Selection, identification and application of a DNA aptamer against Listeria monocytogenes. Food Control 33:239–243
Bing T, Yang XJ, Mei HC, Cao ZH, Shangguan DH (2010) Conservative secondary structure motif of streptavidin-binding aptamers generated by different laboratories. Bioorg Med Chem 18:1798–1805
Cruz-Aguado JA, Penner G (2008) Determination of ochratoxin A with a DNA aptamer. J Agric Food Chem 56:10456–10461
Song KM, Cho M, Jo H, Min K, Jeon SH, Kim T, Han MS, Ku JK, Ban C (2011) Gold nanoparticle-based colorimetric detection of kanamycin using a DNA aptamer. Anal Biochem 415:175–181
McKeague M, Bradley CR, Girolamo AD, Visconti A, Miller JD, DeRosa MC (2010) Screening and initial binding assessment of Fumonisin B1 aptamers. Int J Mol Sci 11:4864–4881
Yang XJ, Bing T, Mei HC, Fang CL, Cao ZH, Shangguan DH (2011) Characterization and application of a DNA aptamer binding to L-tryptophan. Analyst 136:577–585
Wang L, Bao J, Wang L, Zhang F, Li YD (2006) One-Pot synthesis and bioapplication of amine-functionalized magnetite nanoparticles and hollow nanosphere. Chem Eur J 12:6341–6347
Azcona-Olivera JI, Abouzied MM, Plattner RD, Norred WP, Pestka JJ (1992) Generation of antibodies reactive with Fumonisin B1, B2 and B3 by using cholera toxin as the carrier-adjuvant. Appl Environ Microbiol 58:169–173
Avci-Adali M, Paul A, Wilhelm N, Ziemer G, Wendel HP (2010) Upgrading SELEX technology by using lambda exonuclease digestion for single-stranded DNA generation. Molecules 15:1–11
Bozza M, Sheardy RD, Dilone E, Scypinski S, Galazka M (2006) Characterization of the secondary structure and stability of an RNA aptamer that binds vascular endothelial growth factor. Biochemistry 45:7639–7643
Acknowledgments
This work was partly supported by NSFC (21375049), National S&T Support Program of China (2012BAK08B01), S&T Supporting Project of Jiangsu Province (BE2011621, BE2012614), Research Fund for the Doctoral Program of Higher Education (20110093110002), NCET-11-0663, JUSRP51309A and JUSRP21125.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 268 kb)
Rights and permissions
About this article
Cite this article
Chen, X., Huang, Y., Duan, N. et al. Selection and characterization of single stranded DNA aptamers recognizing fumonisin B1 . Microchim Acta 181, 1317–1324 (2014). https://doi.org/10.1007/s00604-014-1260-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00604-014-1260-3