Abstract
The binding domain of Plasmodium vivax Duffy binding protein (PvDBP-II) is a promising blood-stage vaccine candidate for vivax malaria. For the development of a successful vivax malaria vaccine based on DBP-II, the antigenic diversity and also naturally occurring functional antibodies to different PvDBP-II variant types in the various populations must be determined. However, similar to other blood-stage antigens, allelic variation within the PvDBP-II is a fundamental challenge for the development of a broadly efficient vaccine. The present study was performed to define whether the polymorphisms in PvDBP-II influence the nature of functional inhibitory activity of naturally acquired or induced anti-DBP-II antibodies in mice. In this investigation, five genetically distinct variants of PvDBP-II were transiently expressed on the COS-7 cell surface. Erythrocyte-binding inhibition assay (EBIA) was performed using human sera infected with corresponding and non-corresponding P. vivax variants as well as by the use of mice sera immunized with different expressed recombinant PvDBP-IIs. EBIA results showed that the inhibitory percentage varied between 50 and 63 % by using sera from infected individuals, and in case of mouse antisera, inhibition was in the range of 76–86 %. Interestingly, no significant difference was detected in red blood cell binding inhibition when different PvDBP-II variants on the COS-7 cell surfaces were incubated with heterologous and homologous sera infected with PvDBP-II variants. This suggests that the detected polymorphisms in all five forms of PvDBP-II may not affect functional activity of anti-DBP-II antibodies. In conclusion, our results revealed that there are functional cross-reactive antibody responses to heterologous PvDBP-II variants that might provide a broader inhibitory response against all, or at least the majority of strains compared to single allele of this protein that should be considered in development of PvDBP-II-based vaccine.
Similar content being viewed by others
Introduction
Over the past decade, by malaria roll-back strategy and elimination on the global health agenda [1–4], 34 malaria-eliminating countries have achieved remarkable success in reducing their malaria burden. Nowadays, in settings outside of sub-Saharan Africa, malaria cases are increasingly male, adult, clustered geographically, prevalent among migrant and other hard-to-reach groups, and caused mostly by Plasmodium vivax [5]. This parasite species was thought to cause a benign and often self-limiting infection. However, there are increasing reports on clinical severity with emerging virulent forms of P. vivax parasite [6, 7], widespread drug resistance [8–10], and recurrent clinical episodes as a result of reactivation of the dormant forms (hypnozoites) in the liver stage of the parasite’s life cycle [11]. This fact emphasizes the urgent need to create alternative prophylactic and therapeutic strategies, including the development of a vaccine to support the elimination program [12].
Studies in human and animal models have proved that immune responses to blood-stage antigens, mainly merozoite antigens, can control the parasitemia, thus protecting against disease [13]. The P. vivax Duffy binding protein (PvDBP) is one of the most promising blood-stage vaccine candidates. This protein is a 140-kDa microneme-secreted protein [14] that mediates the merozoite–red blood cell (RBC) irreversible attachment and junction formation by interacting with the Duffy antigen receptor for chemokines (DARC) expressed on RBCs [15]. Its amino terminal cysteine-rich region (denoted as region II or PvDBP-II) contains about 330 amino acids and plays an important role in binding to Duffy-positive human erythrocytes [16–18]. The PvDBP-II domain is hypervariable compared to other regions of the protein, and polymorphisms occur frequently at certain residues [19–22]. These polymorphisms consist of host immune selection pressure on this protein [19, 23, 24], which might create a bias toward strain-specific immunity in infected patients with P. vivax [25–28].
Moreover, naturally acquired antibodies to PvDBP-II are prevalent in residents of P. vivax malaria-endemic settings, and these antibodies can functionally inhibit the invasion of merozoites to human RBCs [29–31]. However, such individuals have developed significant quantitative and qualitative differences in their anti-DBP serological responses [23, 25, 29, 32, 33]. With respect to the presence of polymorphisms that might consequently be an obstacle to the development of a broadly effective DBP-II-based vaccine and also the differences in human host response to PvDBP-II among different ethnic groups, developing an effective DBP-II vaccine that could cover strain-specific immunity is in top priority. Interestingly, it has been reported that the essential domains or regions for binding to RBC are less polymorphic. These conserved regions may be suitable targets for cross-reactive antibodies [34–36], and antibodies to these conserved regions could inhibit binding of different variant forms of DBP-II to RBC and overcome the problem of antigenic diversity for designing a broadly effective vaccine.
A previous study in Iran reported a high rate of non-synonymous polymorphisms in Iranian pvdbp-II genes [22]. Moreover, naturally cross-reactive antibodies to the five expressed allelic forms of PvDBP-II were detected among individuals who are living in malaria-endemic areas of Iran by using immune-depletion assay [34]. Therefore, the present investigation was designed to determine whether naturally cross-reactive antibodies have immune protective activity that could have functional inhibitory effects on binding of all five PvDBP-II variant forms expressed on the surface of COS-7 cells to Duffy-positive RBC. Accordingly, it is expected that the results of the present investigation would finally resolve whether development of a PvDBP-II-based vaccine, including a single variant form of PvDBP-II, could induce broadly protective immune response that could be effective for almost all P. vivax strains.
Materials and methods
Human subjects and parasite detection
Blood samples (n = 202) were collected from symptomatic P. vivax-infected volunteers attending the Malaria Health Center in the Public Health Department, Chabahar district in Sistan and Baluchistan Province in Southeastern Iran, where malaria transmission is low and year round. Diagnosis of P. vivax infection was initially performed by light microscopic examination of Giemsa-stained blood smears. Before blood collection, an informed consent was obtained from adults or parents/legal guardians of children. In addition, a questionnaire was used to record each individual’s demographic data, including sex, age, episodes of malaria in the past 10 years, and travel history to highly malaria-endemic areas in neighboring countries, Afghanistan and Pakistan, 4 weeks before sampling. Then, a 2-ml venous blood sample was collected in tubes containing EDTA, on admission and after centrifugation at 4000 rpm for 10 min. Both blood and plasma were collected and stored at −20 °C until use. Next, all samples were transferred to the main laboratory in Tehran. They were then analyzed for final confirmation of microscopic assay for P. vivax DNA by nested polymerase chain reaction (PCR) amplification of a species-specific segment of the 18S rRNA gene of human malaria parasites as described before [37]. For providing negative controls for all experiments, blood samples were obtained from permanent adult residents of Tehran, where malaria transmission has not been reported.
Selection of the five different variant forms of the PvDBP-II
Based on a previous study [34], five genetically distinct variants of PvDBP-II were selected (DBPI, DBPV, DBPVI, DBPIX, and DBPX with GenBank accession nos. EU860428.1, EU860432.1, EU860433.1, EU860436.1, and KF318358, respectively). This selection was performed to test the presence or absence of functional cross-reactive antibodies using the erythrocyte-binding inhibition assay (EBIA). The common pattern of polymorphisms in the selected variants in contrast to DBP-II of Sal I strain (accession no. M61095) is summarized in Table 1. The most important criterion for the selection of these five variant forms was based on the occurrence of polymorphism in inhibitory B cell epitopes determined by Chootong et al. [29]. Therefore, in the present investigation based on Chootong et al. [29], inhibitory linear B cell epitopes were mapped on all five variant forms, and occurred polymorphisms were depicted on the selected variant forms in contrast to the reference strain, Sal I DBP-II (accession no. M61095).
Production of polyclonal antibody by immunizing mice with five variant forms of PvDBP-II
For the generation of antisera to those selected variants of PvDBP-II for EBIA, mice were immunized with expressed recombinant PvDBP-II (rPvDBP-II) as described previously [22]. Briefly, inbred female BALB/c mice (6–8 weeks old) were obtained from Laboratory Animal Science Department, Pasteur Institute of Iran. Mouse groups (1–5) were immunized with five different variant forms of rPvDBP-II (DBPI, DBPV, DBPVI, DBPIX, and DBPX) with 40 µg expressed proteins. The sixth group also received 40 µg of a mixture of five tested rPvDBP-II variant forms (8 µg of each). For priming, the antigens were emulsified in complete Freund’s adjuvant (CFA, 1:1 ratio, Sigma-Aldrich Co., USA) and administered subcutaneously at the base of tail. The animals were boosted with antigen solution emulsified in incomplete Freund’s adjuvant (IFA 1:1 ratio, Sigma-Aldrich Co., USA) on days 14 and 28. The mice in control groups were primed and boosted with PBS alone as well as with PBS in Freund’s adjuvant (CFA in priming, IFA in boosting). Bleeding from all mouse groups was carried out on days zero (pre-immune), 21, and 35 after the first immunization. Serum samples were collected from all groups and stored at −20 °C until use. Binding specificity, cross-reactivity, and anti-serum titers of each individual to the homologous as well as the heterologous antigens used for immunization were determined by ELISA.
PvDBP-II constructs for COS-7 surface expression
In this study, DNA fragments encoding the five selected pvdbp-II genes were cloned into the pHVDR22 vector (kindly provided by Professor C.E. Chitnis, Institut Pasteur Paris, France). pHVDR22 vector which is derived from pRE4 plasmid encodes the dbp-II gene (amino acids 198–522 of Sal I sequence) with the signal peptide and hydrophobic transmembrane stretch of the herpes simplex virus glycoprotein D1 (HSV gD1), a type I integral membrane protein [17]. This expression vector contains a SV40 origin of replication (SV40 ori), a Rous sarcoma virus LTR (RSV LTR) as a promoter and the SV40 early polyadenylation signal (SV40 polyA). HSV gD1 has unique Apa I and Pvu lI restriction sites. Insertion of the all sequences was accomplished by the amplification of the 1173-bp fragment of pvdbp-II corresponding to the base pairs 594–1767 (amino acids 198–589) of Sal I strain (M61095) with the following forward and reverse primers:
HVDRDBPF: 5-TGGCCCAGCTGGATTATGAGACATCTA-3_ and HVDRDBPR: 5_-TCCAAGGGCCCACTATCTACAGGCTG-3 to generate Pvu II and Apa I sites (underline and bold), respectively. After the excision of pHVDR22 vector and PCR products with Pvu II and Apa I restriction enzymes, the ligation of all five pvdbp-II genes was performed, and the DBP-II gene of pHVDR22 vector was replaced with all five pvdbp-II variants in separate reactions and confirmed by PCR. The accuracy of amino acid sequences in all recombinant constructs was confirmed by sequencing analysis using the HVDRDBPF and HVDRDBPR primers.
COS-7 culture and transfection
All recombinant plasmids were purified using endotoxin-free NucleoBond Xtra Maxi Plus plasmid purification kit (Macherey–Nagel, France). Recombinant plasmids were transfected into green monkey kidney cells (COS-7, American Type Culture Collection, and Manassas, VA, USA) by the use of Lipofectamine LTX transfection kit (Invitrogen Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s protocols. Briefly, COS-7 cells were seeded in six-well culture plates (0.45 × 105 cells/ml) in Dulbecco’s Modified Eagle Medium (DMEM, Sigma, USA) with 10 % fetal bovine serum (FBS, Gibco BRL, Life Technologies, Rockville, MS, USA) 24 h before transfection. In the next day, monolayer of COS-7 cells was transfected with recombinant plasmids (2.5 µg/well) as well as liposome complexes (2 % plus reagent and 4 % lipofectamine) in DMEM without serum and then incubated in a humidified incubator with 5 % CO2 at 37 °C for 7–8 h. Afterward, the transfection mixture was replaced with DMEM plus 10 % FBS (Gibco BRL, Life Technologies, Rockville, MS, USA) followed by incubation in a humidified incubator with 5 % CO2 at 37 °C for 36 h. A green fluorescent protein (GFP)-expressing vector (pEGFPN1, Invitrogen, Carlsbad, CA, USA) was used as positive control for checking the transfection efficiency.
Confirmation of PvDBP-II expression on the surface of transfected COS-7 cells with pHVDR22 vector
Transfected COS-7 cells, which have been seeded on the coverslips, were assayed for protein expression using immunofluorescent microscopy and rosetting assays 44 h after transfection. With immunofluorescent microscopy assay, both surface and internal expression were evaluated. The assay was performed as followed: The transfected COS-7 cells were washed by dipping coverslips in PBS 1× (pH 7.2) for 1 min, fixed in 2 % formaldehyde (in PBS) at room temperature (RT) for 15 min and washed again in PBS 1× (pH 7.2). The cells were then incubated at RT for 1 h with diluted anti-mouse anti-DBP-II antibody (1:200, produced in this study) in PBS + 3 % bovine serum albumin (BSA, Sigma-Aldrich Co., USA, for detection of surface expression), or PBS + 3 % BSA + 0.1 % saponin (Sigma-Aldrich Co., USA, for detection of internal expression). The cells were washed once in an excess of PBS 1× (pH 7.2) and incubated at RT for 45 min with 100 µl anti-mouse IgG-FITC-conjugated antibody (Sigma-Aldrich Co., USA) at 1:40 diluted in PBS + 3 % BSA, or PBS + 3 % BSA + 0.1 % saponin. Evans blue was added to this antibody mixture with the ratio of 1:100. The coverslips containing cells were washed in PBS for 5 min and then observed for the expression of protein on the cell surface (in the absence of saponin) or inside the cells (in the presence of saponin). Untransfected COS-7 cells were used as control.
The second test to check surface expression was rosetting assay. In this assay, the ability of expressed rPvDBP-II to bind to RBC was evaluated. For this purpose, Duffy-positive human erythrocytes were suspended in incomplete DMEM, and then the RBCs were added to each well (final cell suspension of 1 %). All plates were incubated with 5 % CO2 at 37 °C for 1 h, and subsequently unbound erythrocytes were removed by washing the wells three times with incomplete DMEM. Binding was quantified by examining rosette formation at magnification 40× with an inverted microscope. Rosettes were counted as positive when adherent erythrocytes covered more than 50 % of the COS-7 cell surface. Untransfected COS-7 cells were used as negative control for rosetting assays.
Erythrocyte-binding inhibition assay
For assessment of the inhibitory activity of both serum samples from patients infected with P. vivax (n = 20) and collected sera from the immunized mice, all five transfected COS-7 cells with recombinant pHVDR22 constructs were incubated at 37 °C for 1 h with corresponding and non-corresponding antisera diluted in incomplete DMEM. For human sera, 20 patients’ sera infected with P. vivax (15 males and 5 females, age range between 10 and 62 years, mean age: 28.5) were selected from 202 collected P. vivax-infected human’s sera on the basis of infected parasite sequence type and also from OD ≥ 1 of anti-PvDBP-II-IgG based on ELISA results [34]. In this test, various dilutions of sera (1:10, 1:20, 1:50, and 1:100) were examined to determine the best dilution for performing EBIA. In addition, sera obtained from each immunized mouse group were examined with homologous and heterologous PvDBP-II antigens expressed on the surface of transfected COS-7 cells. Similar to the human sera, various dilutions of mouse sera were evaluated (1:10, 1:100, 1:200, and 1:400). In the next step, all five transfected COS-7 cells were incubated with human erythrocytes (1 % final suspension in incomplete DMEM) at 37 °C with 5 % CO2 for 1 h. The cells were washed three times with PBS 1× to remove unbound erythrocytes. Erythrocyte binding was scored by counting the number of rosettes per 30 fields at magnification 40× under an inverted microscope. The erythrocyte-binding inhibition of all five expressed PvDBP-II on the COS-7 cell surface was calculated as (R c − R t)/R c × 100 [38], where R c is the average of the number of rosettes in the control wells (incubated with normal human sera or pre-immune mice sera) and R t is the average of the number of rosettes in the test wells (incubated with corresponding and non-corresponding antisera of either human infected with P. vivax or immunized mice).
Results
Parasite species infecting subjects and the selection of samples for EBIA
Plasmodium vivax infection was diagnosed initially by the light microscopic examination of Giemsa-stained blood smears in all 202 examined patients and further confirmed by a molecular approach. No mixed infections and P. falciparum mono-infection were detected among the examined samples. Furthermore, by using a standard ELISA, it was revealed that natural P. vivax infection produced IgG against all five examined variant forms of PvDBP-II with no significant difference (42.1 % for each variant, P > 0.05) [34]. In addition, 63 out of 202 P. vivax samples were sequenced, and 16 haplotypes were detected [22]. For performing EBIA, 20 out of 63 serum samples were selected based on the infected sequence type, and OD ≥ 1 of anti-PvDBP-II-IgG based on ELISA [34]. The pvdpb-II gene sequence analysis showed that these 20 selected P. vivax samples harbored eight different variant forms, namely DBPI, DBPIII, DBPIV, DBPV, DBPVI, DBPIX, DBPX, and DBPXI.
Inhibitory epitope mapping of five PvDBP-II variant forms
According to a previous study performed by Chootong et al. [29], 10 inhibitory epitopes were determined in the entire PvDBP-II region by using human serum infected with P. vivax from Papua New Guinea (PNG). By performing EBIA, those determined inhibitory epitopes were categorized into three distinct groups of high-(H1, H2, and H3), medium-(M1, M2, and M3), and low-(L1, L2, L3, and L4) binding inhibitory epitopes. In current study, in comparison with the reference sequence (Sal I, accession no. M61095), inhibitory B cell epitopes of all five selected PvDBP-II variant forms were mapped. The results revealed that two of the high inhibitory B cell epitopes (H1 and H3) and one medium inhibitory epitope (M2) harbored a higher rate of single nucleotide polymorphisms (SNPs) in contrast to H2, M3, L2, and L3 epitopes in five examined DBP-II variants. However, M1, L1, and L4 epitopes were conserved among all examined variant forms (Table 2).
Polyclonal antibody production to all five rPvDBP-II variants
The production of polyclonal antibody to rPvDBP-II was evaluated in BALB/c mice immunized with either each one of the five rPvDBP-II variants (DBPI, DBPV, DBPVI, DBPIX, and DBPX) or their combination in a single injection. Both single- and mixed-allele immunization strategies elicited high-level anti-DBP-II IgG responses in mice against homologous and heterologous rPvDBP-II [34]. No anti-DBP-II responses were observed in control groups, indicating the specificity of the antibody to rPvDBP-II.
Confirmation of surface expression of PvDBP-II by immunofluorescent microscopy and rosetting assay
COS-7 cells were successfully transfected with recombinant pHVDR22-PvDBP-II construct. Immunofluorescence test (IFAT) results showed that both surface expression (Fig. 1a, b) and internal distribution of all five PvDBP-II variants (Fig. 1c, d) in transfected COS-7 cells but not in untransfected cells (Fig. 1e) were detected. In case of performing the rosetting assay, the cells transfected with recombinant pHVDR22-PvDBP-II vector were covered by more than 50 % with bound Duffy-positive RBC (Fig. 2a) considered as positive. Approximately 6–10 cells in each microscopic field were formed rosettes (Fig. 2b), but in negative wells, no rosettes were detected with untransfected COS-7 cells (Fig. 2c).
Immunofluorescence assays showing the expression of PvDBP-IIs in transfected COS-7 cells. a, b The surface expression of PvDBP-II, staining in the absence of saponin. c, d The internal distribution of expressed proteins, staining in the presence of saponin. e Untransfected COS-7 cells as negative control
Representative erythrocyte-binding assay showing the expression of PvDBP-IIs on the surface of recombinant pHVDR22-PvDBP-II-transfected COS-7 cells. a Representative positive test for rosetting assay. b An appropriate protein expression level by using recombinant pHVDR22-PvDBP-II, c Cells transfected with control vector lacking PvDBP-II or untransfected COS-7 cells. Rosettes are indicated with arrows
Measurement of DBP erythrocyte-binding inhibition
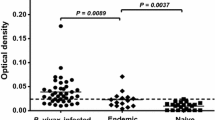
For evaluation of whether the inhibitory antibodies to expressed PvDBP-II are strain specific or strain transcending, both human sera infected with P. vivax and immunized mice with homologous and heterologous variant forms were used in the EBIA test. In this assay, based on sequencing data, 14 out of 20 examined patients’ sera were infected with one of the selected variants [DBPI (LS/GNH/NIW/I, n = 1, EU860428.1), DBPV (FR/DKR/KIR/K, n = 1, EU860432.1), DBPVI (FR/GKH/NLW/K, n = 5, EU860433.1), DBPIX (FR/DKR/NLW/I, n = 6, EU860436.1), and DBPX (FS/GNH/KIR/K, n = 1, KF318358)], and the rest of the sera were obtained from the patients infected with heterologous variant forms, including [DBPIII: (FR/GKR/KIR/I, n = 3, EU860430.1), DBPIV (FR/GKR/KIR/K, n = 1, EU860431.1), and DBPXI, (FR/GQR/KIR/K, n = 2, KF318359)]. In comparison with the control wells (normal human sera), EBIA results showed that the inhibitory percentage varied between 50 and 63 % (Fig. 3a) by using human sera infected with P. vivax (1:10 dilution). The lowest RBC-binding inhibition percentage (50 %) was observed when DBPI-expressing COS-7 cells were inhibited with heterologous sera (infected with DBPIII variant form). Surprisingly, in this study, the highest RBC-binding inhibition (63 %) was observed when DBPIX- and DBPX-expressing cells were inhibited with sera infected with heterologous sera infected with DBPIII and DBPVI variant forms, respectively (Fig. 3a).
Results of erythrocyte-binding inhibition results determining cross-reactive blocking antibodies in the sera of the patients infected with different PvDBP-II variant forms or immunized mice. a Sera from the patients infected with DBPI (LS/GNH/NIW/I, n = 1), DBPIII: (FR/GKR/KIR/I, n = 3), DBPIV (FR/GKR/KIR/K, n = 1), DBPV (FR/DKR/KIR/K, n = 1), DBPVI (FR/GKH/NLW/K, n = 5), DBPIX (FR/DKR/NLW/I, n = 6), DBPX (FS/GNH/KIR/K, n = 1), and DBPXI (FR/GQR/KIR/K, n = 2). Normal human sera with the same dilution (1:10) were used in control wells. b Sera from immunized mice with DBP-I, V, VI, IX, X, and a mixture of all five variant forms. Pre-immune mouse sera with the same dilution (1:100) were used in control wells
In the case of RBC-binding inhibition by mouse antisera to rPvDBP-II (1:100 dilution), inhibition was in the range of 76–86 % (Fig. 3b) in comparison with pre-immune sera as control. The lowest RBC-binding inhibition with mouse antisera (76 %) was observed when DBPI- and DBPIX-expressing COS-7 cells were inhibited with DBPV and DBPX mouse infected sera, respectively. In the present investigation, the highest inhibition (86 %) was calculated when DBPV- and DBPX-expressing cells were inhibited with mouse antisera infected with DBPIX and DBPVI variant forms, respectively. In addition, RBC-binding inhibition using sera collected from the mice immunized with a mixture of the five PvDBP-II variant forms in single injection revealed inhibition in the range of 77–82 %.
Discussion
Success in malaria elimination and final eradication can be easily broken down, and for sustaining this achievement, continued effort and the establishment of new tools such as an effective vaccine are highly recommended. One of the reasons for the slow progress in developing an effective malaria vaccine is the strong capacity of Plasmodium parasites to evade the host’s immune response. This ability derives from the genetic complexity of the parasite, which exhibits genetic diversity as well as antigenic variation during the multistage life cycle that may induce strain-specific immunity. Therefore, multiple variant antigens are required to be included in malaria vaccine development to overcome antigenic diversity and variation [25, 26, 39–42]. Development of such a vaccine would be difficult due to consistency in formulation and an increase in the cost of vaccine manufacturing. In the present study, DBP as one of the most promising blood-stage vaccine candidate antigens with polymorphic nature, which could be a major challenge for developing a broadly efficient vivax vaccine, was selected as a target antigen. In continuation of our previous studies [34, 43], functional cross-protection of anti-PvDBP-II antibodies toward different variant forms of the target protein was evaluated to ensure whether a single variant of PvDBP-II could provide protection to diverse parasite strains. Therefore, in this study, we provide evidence that naturally acquired and vaccine-elicited antibodies to the five different variant forms of PvDBP-II can partially block in vitro interaction between the PvDBP-II ligand domain and its receptor present on the erythrocyte surface.
In any vaccine development, a reliable and functional test is highly required to assess the vaccine efficacy. Therefore, as P. vivax infects only reticulocytes, and there is no practical continuous in vitro culture [44], a reliable test for assessing the EBIA needed. In this regard, several biological assays have been developed and used. Among them, the COS-7 in vitro binding assay, as a reliable and widely accepted functional assay [17, 31], has been used in a variety of investigations [26, 28–30, 35, 45]. The main critical point in this functional assay is a high-level expression of target protein (PvDBP-II in our study) on the surface of COS-7 cells, because the positive test considers rosettes number when adherent erythrocytes cover more than 50 % of the COS-7 cell surface. In the present survey, the COS-7 in vitro binding assay was used to determine the efficacy of the naturally occurred and vaccine-elicited anti-PvDBP-II antibodies to inhibit DBP-II–erythrocyte interaction. In the first step, commercial pDisplay™ plasmid was used as mammalian expression vector (data not shown). Proteins expressed from pDisplay™ were fused at the N terminus to the murine Ig κ-chain leader sequence, and at the C terminus to the platelet-derived growth factor receptor (PDGFR) transmembrane domain, which displays the protein on the extracellular side. This vector was used successfully for the expression of a surface protein by another antibody engineering study [46]. However, in the present study, very low-level surface expression of the five recombinant variants of PvDBP-II, which previously identified as genetically or antigenically distinct [21, 22] on COS-7 cells, was detected. This low-level surface expression was inappropriate for RBC-binding inhibition assay, and thus, no actual rosettes were detected. Therefore, in the next step, pHVDR22 plasmid (a gift from Professor C.E. Chitnis, Institut Pasteur France) was used as a substitute for the mammalian expression vector with a high rate expression of all the five PvDBP-II variants on the COS-7 cell surface, in which it was appropriate for EBIA as shown by others [17, 26, 29, 31, 41].
Regarding determination and characterization of the actual protective immunity against PvDBP-II for vaccine development and based on various previous studies, there are two distinct concepts. In this regard, one group assumes that protective immunity against PvDBP-II could be strain transcending by considering conserved regions of the protein [30, 35, 36, 45, 47]. On this subject, Ntumngia et al. [36] showed that some monoclonal antibodies produced against distinct DBP-II variants recognized conserved epitopes on all tested DBP variants. Also in another study by the same group, a novel artificial DBP-II variant, termed DEKnull derived from Sal I DBP-II, could maintain erythrocyte-binding inhibitory of Sal I strain, indicating the stimulation of anti-PvDBP-II antibodies to shared epitopes of Sal I DBP-II [45]. Similarly, another research group [35] reported that antisera raised in laboratory animals against conserved sub-domains of PvDBP-II inhibited the binding of full length of PvDBP-II to its ligand, which supports the strain-transcending assumption [35]. In parallel, Chootong et al. [47] reported that inhibitory activity of the monoclonal antibody could equally block the RBC binding of different variant forms of Thai PvDBP-IIs. In fact, in the previous study by our group [34] by performing a series of antibody depletion ELISA, it has been shown that the occurrence of SNPs does not change the PvDBP structure that might alter antibody recognition. Moreover, the present RBC-binding inhibitory activity of naturally acquired sera (from patients infected with P. vivax in malaria-hypoendemic regions of Iran) or polyclonal antibodies (produced in mice) to corresponding and non-corresponding PvDBP-II variants expressed on the COS-7 cell surface further confirmed that inhibitory antibodies showed a high level of cross-reactivity that equally block erythrocyte binding of different distinct variant forms of PvDBP-II. These results support the first concept and confirm that the protective immunity against PvDBP-II could be strain transcending.
The second concept, on the other hand, which was from a number of groups that their work has shown protective immunity against PvDBP-II is strain specific [25, 26, 29]. VanBuskirk et al. [26] reported that by introducing three polymorphisms (N417K, W437R, and I503K) into Sal I DBP-II via site-directed mutagenesis, the ability of RBC-binding inhibition was significantly reduced using the original variant of anti-Sal I DBP-II serum produced in rabbit [26]. In addition, Chootong et al. [28, 29] have suggested that certain mutations at the dimer interface, either monomorphic (R308S and D384K) or dimorphic (D384K and K386N), in target epitopes of inhibitory anti-DBP-II IgG, particularly in H1 and H3 protective epitopes, can alter the inhibitory level of acquired neutralizing antibody to block DBP function. In contrary to these reports, all the five expressed antigens in this work had different combinations at 417, 437, and 503 residues. Furthermore, precisely three examined variant forms (DBP-I, DBP-VI, and DBP-X) harbored a number of polymorphic residues at determined inhibitory epitopes, especially in H1 and H3, but RBC-binding inhibitory level (percentage) of the examined variants did not change notably. Moreover, no difference was observed in RBC-binding inhibition when DBPI was expressed on COS-7 cells, which harbored two and five SNPs in H1 and H3, respectively, and incubated with infected sera with homologues and heterologous variant forms of PvDBP-II. These results are in line with EBIA results in our study that used the sera of the mice immunized with multi- or single variant forms of rPvDBP-II.
Previous reports from different populations living in vivax malaria-endemic areas have determined the presence of robust naturally acquired anti-PvDBP-II antibodies that were increased with exposure [25, 31, 48, 49] and age [33, 50]. Moreover, it has been demonstrated that these antibodies can inhibit PvDBP-II erythrocyte binding and accordingly block the invasion of erythrocytes in vitro [25, 31, 48, 49]. However, it is worth mentioning that relatively few individuals are capable of developing broadly inhibitory anti-PvDBP-II response against multiple allelic variants [29, 30] that surely is a great challenge in developing PvDBP-II-based vaccine. Our study confirms the efficiency of naturally acquired anti-DBP-II from infected patients in low and unstable transmission areas in controlling P. vivax infection by the partial inhibition of DBP-II binding to Duffy-positive erythrocytes. Also, these responses were significantly higher in vivax patients than uninfected residents out of malaria-endemic areas of Iran. Currently, in this concern, a few reports have evaluated naturally acquired functional inhibitory antibodies in different malaria-endemic regions [28, 30, 31, 48]. Among these reports, Michon et al. [31], by using pooled plasmas from infected patients with P. vivax from PNG reported that the sera from high responder subjects completely inhibit RBC binding (100 %), whereas those of low-responder subjects showed 50 % inhibition (at 1:10 dilution). However, in the present study, pooled sera from the subjects infected with P. vivax with a high level of anti-PvDBP-II antibodies [34] blocked RBC binding about 60 % (at 1:10 dilution). Such discrepancy in the level (percentage) of blocking activity by naturally occurred antibodies might be related to seasonality, low malaria transmission, the number of exposure to pathogen and the genetic background of the infected subjects in low and high vivax transmission areas. Besides, as pooled samples were used in the present study, individual variability in the inhibitory immune response (0–100 %), which was proposed to be a characteristic of unstable malaria transmission areas such as previous reports from Thailand and Brazil, needs further study [28, 48].
Interestingly, it has been suggested that DBP is a poor immunogen [48], but its moderate inhibition activity observed in this study (due to low level of endemicity) further support its inclusion in vaccine development [34]. However, Michon et al. [31] demonstrated that the secondary boosting of an initially low-responder immune response is important in congenic mice, which provide hope for PvDBP-II-based vaccine development in low-responder residents of malaria-endemic regions. This low responsiveness may arise due to just-in-time releasing of DBP in the invasion pathway [26, 51]. Therefore, based on the present results, by determining the variant form of the infected PvDBP-II of all tested human sera from low and seasonal transmission areas and more comprehensive cross-reactivity checking panel, the possibility that DBP-II cross-variant inhibitory activity reflects only on an accumulation of antibodies to strain-specific epitopes is unlikely.
In conclusion, an effective PvDBP-II vaccine will need to induce efficient functional inhibition against almost all PvDBP-II variants that are circulated in given malaria-endemic areas. In the present study, results showed the natural exposure to P. vivax in areas of low and unstable transmission induced anti-DBP antibody that inhibited the binding of diverse variants of DBP-II to reticulocytes. Therefore, such functional cross-reactive antibody responses to heterologous PvDBP-II variants might provide a broader inhibitory response against all, or at least the majority of strains compared to a single allele of this protein that should be considered in development of PvDBP-II-based vaccine.
References
Tanner M, de Savigny D (2008) Malaria eradication back on the table. Bull World Health Organ 86:82
WHO (2008) The world Malaria report. World Health Organization, Geneva
Mendis K, Rietveld A, Warsame M, Bosman A, Greenwood B, Wernsdorfer WH (2009) From malaria control to eradication: the WHO perspective. Trop Med Int Health 14:802–809
Kelly GC, Tanner M, Vallely A, Clements A (2012) Malaria elimination: moving forward with spatial decision support systems. Trends Parasitol 28:297–304
Cotter C, Sturrock HJ, Hsiang MS, Liu J, Phillips AA, Hwang J, Gueye CS, Fullman N, Gosling RD, Feachem RG (2013) The changing epidemiology of malaria elimination: new strategies for new challenges. Lancet 382:900–911
Sharma A, Khanduri U (2009) How benign is benign tertian malaria? J Vector Borne Dis 46:141–144
Alexandre MA, Ferreira CO, Siqueira AM, Magalhães BL, Mourão MP, Lacerda MV, Alecrim Md (2010) Severe Plasmodium vivax malaria, Brazilian Amazon. Emerg Infect Dis 16:1611–1614
Ketema T, Bacha K, Birhanu T, Petros B (2009) Chloroquine-resistant Plasmodium vivax malaria in Serbo town, Jimma zone, south-west Ethiopia. Malar J 8:177
Mohan K, Maithani MM (2010) Congenital malaria due to chloroquine-resistant Plasmodium vivax: a case report. J Trop Pediatr 56:454–455
Rijken MJ, Boel ME, Russell B, Imwong M, Leimanis ML, Phyo AP, Muehlenbachs A, Lindegardh N, McGready R, Rénia L, Snounou G, Singhasivanon P, Nosten F (2011) Chloroquine resistant vivax malaria in a pregnant woman on the western border of Thailand. Malar J 10:113
Krotoski WA, Collins WE, Bray RS, Garnham PC, Cogswell FB, Gwadz RW, Killick-Kendrick R, Wolf R, Sinden R, Koontz LC, Stanfill PS (1982) Demonstration of hypnozoites in sporozoite-transmitted Plasmodium vivax infection. Am J Trop Med Hyg 31:1291–1293
Ntumngia FB, King CL, Adams JH (2012) Finding the sweet spots of inhibition: understanding the targets of a functional antibody against Plasmodium vivax Duffy binding protein. Int J Parasitol 42:1055–1062
Richards JS, Beeson JG (2009) The future for blood-stage vaccines against malaria. Immunol Cell Biol 87:377–390. doi:10.1038/icb.2009.27
Adams JH, Sim BK, Dolan SA, Fang X, Kaslow DC, Miller LH (1992) A family of erythrocyte binding proteins of malaria parasites. Proc Natl Acad Sci USA 89:7085–7089
Miller LH, Mason SJ, Clyde DF, McGinniss MH (1976) The resistance factor to Plasmodium vivax in blacks: the Duffy-blood-group genotype, FyFy. N Engl J Med 295:302–304
Chitnis CE, Blackman MJ (2000) Host cell invasion by malaria parasites. Parasitol Today 16:411–415
Chitnis CE, Miller LH (1994) Identification of the erythrocyte binding domains of Plasmodium vivax and Plasmodium knowlesi proteins involved in erythrocyte invasion. J Exp Med 180:497–506
Ocampo M, Vera R, Eduardo Rodriguez L, Curtidor H, Urquiza M, Suarez J, Garcia J, Puentes A, Lopez R, Trujillo M, Torres E, Patarroyo ME (2002) Plasmodium vivax Duffy binding protein peptides specifically bind to reticulocytes. Peptides 23:13–22
Cole-Tobian J, King CL (2003) Diversity and natural selection in Plasmodium vivax Duffy binding protein gene. Mol Biochem Parasitol 127:121–132
Ampudia E, Patarroyo MA, Patarroyo ME, Murillo LA (1996) Genetic polymorphism of the Duffy receptor binding domain of Plasmodium vivax in Colombian wild isolates. Mol Biochem Parasitol 78:269–272
Babaeekho L, Zakeri S, Djadid ND (2009) Genetic mapping of the duffy bindingprotein (DBP) ligand domain of Plasmodium vivax from unstable malaria region in the Middle East. Am J Trop Med Hyg 80:112–118
Valizadeh V, Zakeri S, Mehrizi AA, Djadid ND (2014) Population genetics and natural selection in the gene encoding the Duffy binding protein II in Iranian Plasmodium vivax wild isolates. Infect Genet Evol 21:424–435
Xainli J, Baisor M, Kastens W, Bockarie M, Adams JH, King CL (2002) Age-dependent cellular immune responses to Plasmodium vivax Duffy binding protein in humans. J Immunol 169:3200–3207
Tsuboi T, Kappe SH, Al-Yaman F, Prickett MD, Alpers M, Adams JH (1994) Natural variation within the principal adhesion domain of the Plasmodium vivax Duffy binding protein. Infect Immun 62:5581–5586
Xainli J, Cole-Tobian JL, Baisor M, Kastens W, Bockarie M, Yazdani SS, Chitnis CE, Adams JH, King CL (2003) Epitope-specific humoral immunity to Plasmodium vivax Duffy binding protein. Infect Immun 71:2508–2515
VanBuskirk KM, Cole-Tobian JL, Baisor M, Sevova ES, Bockarie M, King CL, Adams JH (2004) Antigenic drift in the ligand domain of Plasmodium vivax Duffy binding protein confers resistance to inhibitory antibodies. J Infect Dis 190:1556–1562
Ceravolo IP, Sanchez BA, Sousa TN, Guerra BM, Soares IS, Braga EM, McHenry AM, Adams JH, Brito CF, Carvalho LH (2009) Naturally acquired inhibitory antibodies to Plasmodium vivax Duffy binding protein are short-lived and allele-specific following a single malaria infection. Clin Exp Immunol 156:502–510
Chootong P, Panichakul T, Permmongkol C, Barnes SJ, Udomsangpetch R, Adams JH (2012) Characterization of inhibitory anti-Duffy binding protein II immunity: approach to Plasmodium vivax vaccine development in Thailand. PLoS One 7:e35769
Chootong P, Ntumngia FB, VanBuskirk KM, Xainli J, Cole-Tobian JL, Campbell CO, Fraser TS, King CL, Adams JH (2010) Mapping epitopes of the Plasmodium vivax Duffy binding protein with naturally acquired inhibitory antibodies. Infect Immun 78:1089–1095
King CL, Michon P, Shakri AR, Marcotty A, Stanisic D, Zimmerman PA, Cole-Tobian JL, Mueller I, Chitnis CE (2008) Naturally acquired Duffy-binding protein-specific binding inhibitory antibodies confer protection from bloodstage Plasmodium vivax infection. Proc Natl Acad Sci USA 105:8363–8368
Michon P, Fraser T, Adams JH (2000) Naturally acquired and vaccine-elicited antibodies block erythrocyte cytoadherence of the Plasmodium vivax Duffy binding protein. Infect Immun 68:3164–3171
Cole-Tobian JL, Cortés A, Baisor M, Kastens W, Xainli J, Bockarie M, Adams JH, King CL (2002) Age-acquired immunity to a Plasmodium vivax invasion ligand, the duffy binding protein. J Infect Dis 186:531–539
Fraser T, Michon P, Barnwell JW, Noe AR, Al-Yaman F, Kaslow DC, Adams JH (1997) Expression and serologic activity of a soluble recombinant Plasmodium vivax Duffy binding protein. Infect Immun 65:2772–2777
Valizadeh V, Zakeri S, Mehrizi AA, Djadid ND (2014) Non-allele specific antibody responses to genetically distinct variant forms of Plasmodium vivax Duffy binding protein (PvDBP-II) in Iranians exposed to seasonal malaria transmission. Acta Trop 136:89–100
Siddiqui AA, Xainli J, Schloegel J, Carias L, Ntumngia F, Shoham M, Casey JL, Foley M, Adams JH, King CL (2012) Fine specificity of Plasmodium vivax duffy binding protein binding engagement of the duffy antigen on human erythrocytes. Infect Immun 80:2920–2928
Ntumngia FB, Schloegel J, Barnes SJ, McHenry AM, Singh S, King CL, Adams JH (2012) Conserved and variant epitopes of Plasmodium vivax Duffy binding protein as targets of inhibitory monoclonal antibodies. Infect Immun 80:1203–1208
Snounou G, Viriyakosol S, Zhu XP, Jarra W, Pinheiro L, do Rosario VE, Thaithong S, Brown KN (1993) High sensitivity of detection of human malaria parasites by the use of nested polymerase chain reaction. Mol Biochem Parasitol 61:315–320
Ntumngia FB, Schloegel J, McHenry AM, Barnes SJ, George MT, Kennedy S, Adams JH (2013) Immunogenicity of single versus mixed allele vaccines of Plasmodium vivax Duffy binding protein region II. Vaccine 31:4382–4388
Genton B, D’Acremont V, Rare L, Baea K, Reeder JC, Alpers MP, Müller I (2008) Plasmodium vivax and mixed infections are associated with severe malaria in children: a prospective cohort study from Papua New Guinea. PLoS Med 5:e127
Polley SD, Tetteh KK, Lloyd JM, Akpogheneta OJ, Greenwood BM, Bojang KA, Conway DJ (2007) Plasmodium falciparum merozoite surface protein 3 is a target of allele-specific immunity and alleles are maintained by natural selection. J Infect Dis 195:279–287
McHenry AM, Barnes SJ, Ntumngia FB, King CL, Adams JH (2011) Determination of the molecular basis for a limited dimorphism, N417K, in the Plasmodium vivax Duffy-binding protein. PLoS One 6:e20192
Cole-Tobian JL, Michon P, Biasor M, Richards JS, Beeson JG, Mueller I, King CL (2009) Strain-specific duffy binding protein antibodies correlate with protection against infection with homologous compared to heterologous Plasmodium vivax strains in Papua New Guinean children. Infect Immun 77:4009–4017
Zakeri S, Babaeekhou L, Mehrizi AA, Abbasi M, Djadid ND (2011) Antibody responses and avidity of naturally acquired anti-Plasmodium vivax Duffy binding protein (PvDBP) antibodies in individuals from an area with unstable malaria transmission. Am J Trop Med Hyg 84:944–950
Russell B, Suwanarusk R, Borlon C, Costa FT, Chu CS, Rijken MJ, Sriprawat K, Warter L, Koh EG, Malleret B, Colin Y, Bertrand O, Adams JH, D’Alessandro U, Snounou G, Nosten F, Rénia L (2011) A reliable ex vivo invasion assay of human reticulocytes by Plasmodium vivax. Blood 118:e74–e81
Ntumngia FB, Adams JH (2012) Design and immunogenicity of a novel synthetic antigen based on the ligand domain of the Plasmodium vivax duffy binding protein. Clin Vaccine Immunol 19:30–36
Ho M, Pastan I (2009) Display and selection of scFv antibodies on HEK-293T cells. Methods Mol Biol 562:99–113
Chootong P, McHenry AM, Ntumngia FB, Sattabongkot J, Adams JH (2014) The association of Duffy binding protein region II polymorphisms and its antigenicity in Plasmodium vivax isolates from Thailand. Parasitol Int 63:858–864
Ceravolo IP, Souza-Silva FA, Fontes CJ, Braga EM, Madureira AP, Krettli AU, Souza JM, Brito CF, Adams JH, Carvalho LH (2008) Inhibitory properties of the antibody response to Plasmodium vivax Duffy binding protein in an area with unstable malaria transmission. Scand J Immunol 67:270–278
Grimberg BT, Udomsangpetch R, Xainli J, McHenry A, Panichakul T, Sattabongkot J, Cui L, Bockarie M, Chitnis C, Adams J, Zimmerman PA, King CL (2007) Plasmodium vivax invasion of human erythrocytes inhibited by antibodies directed against the Duffy binding protein. PLoS Med 4:e337
Michon PA, Arevalo-Herrera M, Fraser T, Herrera S, Adams JH (1998) Serologic responses to recombinant Plasmodium vivax Duffy binding protein in a Colombian village. Am J Trop Med Hyg 59:597–599
Adams JH, Hudson DE, Torii M, Ward GE, Wellems TE, Aikawa M, Miller LH (1990) The Duffy receptor family of Plasmodium knowlesi is located within the micronemes of invasive malaria merozoites. Cell 63:141–145
Acknowledgments
The authors are grateful to Professor C.E. Chitnis, Institut Pasteur Paris, France, for providing pHVDR22 plasmid. We acknowledge with deep respect the generos cooperation of Zahedan University of Medical Sciences, staff in Public Health Department, Sistan and Baluchestan Province and Chabahar District, for their help in the collection of human blood samples. We are thankful to the individuals and their families in Sistan and Baluchistan for their willingness to participate in this study. This project received financial support from Pasteur Institute of Iran to S. Zakeri (Project No. 590).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in this study regarding involving human participants were in accordance with the ethical standards, which has been approved and certified by the Ethical Review Committee of Research of Pasteur Institute of Iran. Also, all animal handling was in accordance with the ethical standards of the Laboratory Animal Science Department, Pasteur Institute of Iran.
Informed consent
An informed consent was obtained from all individual participants, including adults or parents/legal guardians of children.
Rights and permissions
About this article
Cite this article
Valizadeh, V., Zakeri, S., Mehrizi, A.A. et al. Natural acquired inhibitory antibodies to Plasmodium vivax Duffy binding protein (PvDBP-II) equally block erythrocyte binding of homologous and heterologous expressed PvDBP-II on the surface of COS-7 cells. Med Microbiol Immunol 205, 85–95 (2016). https://doi.org/10.1007/s00430-015-0429-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00430-015-0429-7