Abstract
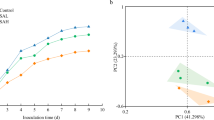
Saline–alkali soil can inhibit the growth of crops as a consequence of cellular damage through oxidation of lipids and proteins and degradation of nucleic acids, ultimately leading to cell death. The bacterial community composition and diversity in saline–alkali soil across different land uses, such as agricultural land, forest land, and grassland, were evaluated using high-throughput sequencing of the bacterial 16S rRNA gene. Significant differences in the soil physicochemical characteristics and bacterial community among different land uses were observed in this study. The soil pH value and electrical conductivity were much higher in grassland soil than in agricultural and forest soils. There were high proportions of Actinobacteria and Proteobacteria (phyla) in agricultural and forest soils, while Actinobacteria, Bacteroidetes, Gemmatimonadetes were more predominant in grassland soil. The relative abundance of dominant taxa exhibited a highly significant correlation with soil pH, water content, EC, and organic matter. The percentage of species that are shared among the different soil samples ranged from 5.3 to 30.5%. The haloalkaliphilic Actinobacterial genus Nitriliruptor was detected in grassland but not in areas with other types of land use. Results of both heatmap and principal component analysis (PCA) indicated that the soil properties and bacterial communities in the areas studied have been greatly influenced by long-term land use by different management.




Similar content being viewed by others
References
Nesme T, Colomb B, Hinsinger P, Watson CA (2014) Soil phosphorus management in organic cropping systems: from current practices to avenues for a more efficient use of P resources. In: Organic farming, prototype for sustainable agricultures. Springer, Berlin, pp 23–45
Heijden MGAvd, Bardgett RD, Straalen NMv (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11:296–310
Gyaneshwar P, Kumar GN, Parekh L, Poole P (2002) Role of soil microorganisms in improving P nutrition of plants. In: Food security in nutrient-stressed environments: exploiting plants’ genetic capabilities. Springer, Berlin, pp 133–143
Ding G-C, Piceno YM, Heuer H, Weinert N, Dohrmann AB, Carrillo A, Andersen GL, Castellanos T, Tebbe CC, Smalla K (2013) Changes of soil bacterial diversity as a consequence of agricultural land use in a semi-arid ecosystem. PLOS one 8(3):e59497
Bissett A, Richardson AE, Baker G, Thrall PH (2011) Long-term land use effects on soil microbial community structure and function. Appl Soil Ecol 51:66–78
Bossio DA, Girvan MS, Verchot L, Bullimore J, Borelli T, Albrecht A, Scow KM, Ball AS, Pretty JN, Osborn AM (2005) Soil microbial community response to land use change in an agricultural landscape of Western Kenya. Microb Ecol 49(1):50–62. doi:10.1007/s00248-003-0209-6
Lagerlöf J, Adolfsson L, Börjesson G, Ehlers K, Vinyoles GP, Sundh I (2014) Land-use intensification and agroforestry in the Kenyan highland: impacts on soil microbial community composition and functional capacity. Appl Soil Ecol 82:93–99
Li ZP, Wu XC, Chen BY (2007) Changes in transforamtion of soil organic carbon and functional diversity of soil microbial community under different land use patterns. Scienctia Agricultura Sinica 40 (8):1712–1721
Xun F, Xie B, Liu S, Guo C (2014) Effect of plant growth-promoting bacteria (PGPR) and arbuscular mycorrhizal fungi (AMF) inoculation on oats in saline-alkali soil contaminated by petroleum to enhance phytoremediation. Environ Sci Pollut Res 22(1):598–608
Kohler J, Hernández JA, Caravaca F, Roldán A (2009) Induction of antioxidant enzymes is involved in the greater effectiveness of a PGPR versus AM fungi with respect to increasing the tolerance of lettuce to severe salt stress. Environ Exp Bot 65(2–3):245–252
Wang X, Wang J, Liu H, Zou D, Zhao H (2013) Influence of natural saline-alkali stress on chlorophyll content and chloroplast ultrastructure of two contrasting rice (Oryza sativa L. japonica) cultivars. Aust J Crop Sci 7:289–292
Tang J, Wang L, Zhang S (2005) Investigating landscape pattern and its dynamics in Daqing, China. Int J Remote Sens 26(11):2259–2280
Wakelin SA, Barratt BI, Gerard E, Gregg AL, Brodie EL, Andersen GL, DeSantis TZ, Zhou J, He Z, Kowalchuk GA (2013) Shifts in the phylogenetic structure and functional capacity of soil microbial communities follow alteration of native tussock grassland ecosystems. Soil Biol Biochem 57:675–682
Schinner F, Ohlinger R, Kandeler E, Margesin R (1995) Methods in soil biology. Springer-Verlag, Berlin
Liu J, Sui Y, Yu Z, Shi Y, Chu H, Jin J, Liu X, Wang G (2014) High throughput sequencing analysis of biogeographical distribution of bacterial communities in the black soils of northeast China. Soil Biol Biochem 70:113–122
Maidak BL, Olsen GJ, Larsen N, Overbeek R, Mccaughey MJ, Woese CR (1997) The RDP (Ribosomal Database Project). Nucleic Acids Res 25(1):109–111
Chao A, Bunge J (2002) Estimating the number of species in a stochastic abundance model. Biometrics 58(3):531–539
Wu S, Wang G, Angert ER, Wang W, Li W, Zou H (2012) Composition, diversity, and origin of the bacterial community in grass carp intestine. PloS One 7(2):e30440
Acosta-Martinez V, Dowd S, Sun Y, Allen V (2008) Tag-encoded pyrosequencing analysis of bacterial diversity in a single soil type as affected by management and land use. Soil Biol Biochem 40(11):2762–2770
Roesch LF, Fulthorpe RR, Riva A, Casella G, Hadwin AK, Kent AD, Daroub SH, Canargo FA, Farmerie WG, Triplett EW (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1(4):283–290
Lauber C, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75(15):5111–5120
Chu HY, Fierer N, Lauber CL, Caporaso JG, Knight R, Grogan P (2010) Soil bacterial diversity in the Arctic is not fundamentally different from that found in other biomes. Environ Microbiol 12:2998–3006
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103(3):626–631
Abell GCJ, Bowman JP (2005) Colonization and community dynamics of class Flavobacteria on diatom detritus in experimental mesocosms based on Southern Ocean seawater. FEMS Microbiol Ecol 53:379–391
Chan OC, Casper P, Qing SL, Li FZ, Fu Y, Dong YX, Ulrich A, Ming ZX (2008) Vegetation cover of forest, shrub and pasture strongly influences soil bacterial community structure as revealed by 16 S rRNA gene T-RFLP analysis. FEMS Microbiol Ecol 64:449–458
Albuquerque L, França L, Rainey FA, Schumann P, Nobre MF, da Costa MS (2011) Gaiella occulta gen. nov., sp. nov., a novel representative of a deep branching phylogenetic lineage within the class Actinobacteria and proposal of Gaiellaceae fam. nov. and Gaiellales ord. nov. Syst Appl Microbiol 34(8):595–599
Navarrete AA, Kuramae EE, M DH, Pijl AS, van Veen JA, Tsai SM (2012) Acidobacterial community responses to agricultural management of soybean in Amazon forest soils. FEMS Microbiol Ecol 83(3):607–621
Sorokin DY, van Pelt S, Tourova TP, Evtushenko LI (2009) Nitriliruptor alkaliphilus gen. nov., sp. nov., a deep-lineage haloalkaliphilic actinobacterium from soda lakes capable of growth on aliphatic nitriles, and proposal of Nitriliruptoraceae fam. nov. and Nitriliruptorales ord. nov. Int J Syst Evol Microbiol 59(2):248–253
Pester M, Maixner F, Berry D, Rattei T, Koch H, Lücker S, Nowka B, Richter A, Spieck E, Lebedeva E (2014) NxrB encoding the beta subunit of nitrite oxidoreductase as functional and phylogenetic marker for nitrite-oxidizing Nitrospira. Environ Microbiol 16(10):3055–3071
Gruber-Dorninger C, Pester M, Kitzinger K, Savio DF, Loy A, Rattei T, Wagner M, Daims H (2014) Functionally relevant diversity of closely related Nitrospira in activated sludge. ISME J 9(3):643–655
Acknowledgements
The authors are grateful for the support by the Project “948” of the Chinese National Forestry Bureau, Grant (No. 2008-4-34) and the Fundamental Research Funds for the Central Universities (2572014CA22).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Peng, M., Jia, H. & Wang, Q. The Effect of Land Use on Bacterial Communities in Saline–Alkali Soil. Curr Microbiol 74, 325–333 (2017). https://doi.org/10.1007/s00284-017-1195-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-017-1195-0