Abstract
The genotoxic potential of waters in six rivers and reservoirs from Serbia was monitored in different tissues of chub (Squalius cephalus L. 1758) with the alkaline comet assay. The comet assay, or single-cell gel electrophoresis, has a wide application as a simple and sensitive method for evaluating DNA damage in fish exposed to various xenobiotics in the aquatic environment. Three types of cells, erythrocytes, gill cells, and liver cells, were used for assessing DNA damage. Images of randomly selected cells were analyzed with a Leica fluorescence microscope and image analysis by software (Comet Assay IV Image analysis system, PI, UK). Three parameters (tail length—l, tail intensity—i, and Olive tail moment—m) were analyzed on 1,700 nuclei per cell type. The procedure for sum of ranking differences (SRD) was implemented to compare different types of cells and different parameters for estimation of DNA damage. Regarding our nine different estimations of genotoxicity: tail length, intensity, and moment in erythrocytes (rel, rei, rem), liver cells (rll, rli, rlm), and gill cells (rgl, rgi, rgm), the SRD procedure has shown that the Olive tail moment and tail intensity are (almost) equally good parameters; the SRD value was lower for the tail moment and tail intensity than for tail length in the case of all types of cells. The least reliable parameter was rel; close to the borderline case were rei, rll, and rgl (∼5 % probability of random ranking).

Comparison of comet assay parameters
Similar content being viewed by others
Introduction
The comet assay or single-cell gel electrophoresis assay is a rapid, sensitive, and relatively simple method for detecting DNA damage at the level of individual cells [1]. It combines the simplicity of biochemical techniques for detecting DNA single-strand breaks (strand breaks and incomplete excision repair sites), alkali labile sites, and cross-linking, with the single-cell approach, typical for cytogenetic assays [2]. The comet assay has found its application in basic research as well as in applied sciences such as medicine, ecogenotoxicology, monitoring studies, etc. [3–7]. The comet assay is based on the ability of negatively charged loops/fragments of DNA to be drawn through an agarose gel in response to an electric field. The extent of DNA migration depends directly on the DNA damage present in the cells. DNA lesions consisting of strand breaks after treatment with alkali either alone or in combination with certain enzymes (e.g., endonucleases) increase DNA migration compared to those in concurrent controls [8]. The determination of the shape, size, and amount of DNA within comets is a very important attribute of the assay if the damage is to be evaluated accurately.
In parallel with several technical and procedural evolutions to make the assay more robust, several approaches have also evolved to quantify the extent of damage more reliably, reproducibly, and meaningfully. Such quantification includes both visual examinations (i.e., photographic, occulometer, or nonspecific image analysis systems) and the usage of commercially available (or public domain specific) image analysis software packages. Such specific software packages also facilitate statistical analyses, plotting, and documentation of the data [2]. Besides that, an automated system provides an advantage over manual, not only for easier management, but also because of the lack of observer subjectivity.
As there are more parameters for the selection (tail moment, tail length, and tail intensity), it leads to controversy among researchers which is the most suitable parameter for assessing the damage of DNA. As SRD is entirely general, it is a suitable method to investigate which of the offered parameters is the most reliable to represent the damage of DNA molecules. SRD corresponds to the principle of parsimony; it provides an easy way to rank (e.g., techniques, models, and methods) in a unique, justified way [9]. The method based on ranking is nonparametric and robust in the common sense. SRD values can be used to compare data originating from normal and definitely not normal distributions [10]. Tailings (outliers) do not significantly influence the ranking, so no distributional assumption is needed.
The objective of this study was to find out which estimated parameters are the most reliable for the assessment of genotoxicity by sampling from sites with different anthropogenic impacts: the Uvac River “Zlatar” reservoir (protected natural area), Reservoir Garasi with low impact, and sites Bubanj Potok, Pestan, Beljanica, and Kolubara with a high degree of such impact. The data were obtained from comet assays performed on erythrocytes, and gill and liver cells of chub (Squalius cephalus L. 1758). The ranked parameters were tail length, tail intensity, and Olive tail moment—OTM [11].
Material and methods
Field sampling
Field sampling was conducted at four rivers (Bubanj Potok, Kolubara, Pestan, Beljanica) and two reservoirs (Lakes Zlatar and Garasi) from Serbia. The Uvac River “Zlatar” reservoir (reference site) is a protected natural area of great importance with very low anthropogenic impact. Reservoir Garasi was chosen as the site with low anthropogenic impact as it is used as a drinking water source. Rivers Pestan, Beljanica, and Kolubara are polluted sites at Kolubara basin, which has rich deposits of lignite (brown coal), and hence, the whole area is under intensive mining activity. Bubanj Potok is an example of a municipal creek with impact from industrial waste. As far as we know, there are no previous records of genotoxicity assessments of any kind in these rivers and reservoirs.
A total of 34 chub specimens were caught by an electrofishing device ELEMAX SHX 2000 (SAWAFUJI) on rivers and by gill and trammel nets in reservoirs. Fish specimens were anesthetized with clove oil, and samples of blood, gills, and liver were collected, stored on ice, and transferred to the laboratory in a dark cool box.
Comet assay
The alkaline comet assay procedure was performed under yellow light, in accordance with the method described by Singh et al. [1]. Microscopic slides were precoated with 0.5 % normal melting point (NMP) agarose and air-dried for 24 h. To form the second, supportive layer, 80 μl of 1 % NMP agarose was gently placed on the top of the 0.5 % NMP layer and spread over the slide using a coverslip. The slide was placed on ice for 5 min to allow complete polymerization of the agarose. After the coverslips were removed, 30 μl of cell pellet suspension, gently mixed with 70 μl of 1 % LMP (37 °C) agarose, was pipetted on the supportive layer of 1 % NMP agarose and covered with a coverslip. After keeping coverslips for 5 min on ice, they were removed, and the slides were placed into freshly made cold lysis buffer (2.5 M NaCl, 100 mM EDTA, 10 mM Tris, 1.5 % Triton X-100, pH 10 and with 10 % dimethyl sulfoxide for blood) for 1 h. To allow DNA unwinding, slides were put into the electrophoresis chamber containing cold alkaline electrophoresis buffer (300 mM NaOH, 1 mM EDTA, pH 13) for 20 min. Electrophoresis was performed at 0.75 V/cm at 4 °C for 20 min. After the electrophoresis, slides were placed into freshly made neutralizing buffer (0.4 M Tris, pH 7.5) for 15 min. Staining was performed with 20 μl per slide of EtBr (2 μg mL−1). The slides were examined with a fluorescence microscope (Leica, DMLS, Austria, ×400 magnifications, 510–560-nm excitation filter, and 590-nm barrier filter). Microscopic images of comets were scored using the Comet IV Computer Software (Perceptive Instruments, UK). Images of 50 cells were captured from each slide per sample, and among the parameters available for analyses, the tail length, tail intensity, and OTM were chosen as parameters to assess the DNA damage (Fig. 1).
Sum of ranking differences methodology
SRD methodology has already been applied in various scientific disciplines in the past. It can be used for column selection in chromatography [9], for QSAR, QSPR, and QSRR investigations [12–17], for testing performance for Raman spectra resolution [18], for grouping polarity measures [19, 20], for comparison of chemometric methods [21, 22], for checking evaluation panels in food chemistry [23, 24], for ranking polluted environmental sites [25], for comparing estimation methods of octanol–water partition coefficients [10, 26], for comparative evaluation of acidic dissociation constants [27], as well as for comparison of Elisa Veratox assay and liquid chromatography [28].
The SRD procedure is supervised in the sense that a reference (benchmark) ranking should be available. The data should be arranged in matrix form—the objects (rows; in this case, samples; 34 chub specimens) and the variables, nine different ways for estimations of genotoxicity with three types of evaluation: tail length, tail intensity, and Olive tail moment, in three different cells: erythrocytes, liver cells, and gill cells. The notations contain three letters; the first “r” indicates that rank numbers were analyzed to avoid scaling problems, the second letter indicates the cell types, and the third letter shows the evaluation type: rel, rei, rem, rll, rli, rlm, rgl, rgi, and rgm, respectively. The most apparent choice is to use the average of all estimations for genotoxicity as reference ranking (row average). The ranking by average values is used to call consensus, since the errors cancel each other. The maximum likelihood principle will ensure that the most probable ranking will be provided by the average. However, the average is not necessarily a bias-free solution, as the selection of models to be averaged is more or less arbitrary.
The absolute values of the differences between the average and individual rankings were calculated and summed for each evaluation method. The closer is the SRD value to zero (i.e., the closer is the ranking to the reference value), the “better” is the evaluation method in the sense that it is better suitable to replace all other methods. The proximity of SRD values shows that the methods behave similarly. Some groupings of evaluation methods can also be observed, whereas their distances show dissimilarity from the ordering point of view [9, 10].
Validation of the SRD procedure can be carried out in two ways: (1) using simulated random numbers in conjunction with the theoretical distribution of the SRD values (generated by random numbers and corresponding to the number of specimens) called comparison of ranks with random numbers (CRRN) procedure and (2) using cross validation (CV). The validation by CRRN procedure has been described earlier in detail in ref. [10]. Alternatively, leave-many-out CV has been used following the literature recommendations [29]. Approximately 1/7th of the objects (specimens) were left out, and the ranking was made on the remaining 6/7th number of objects. In such a way, standard deviation can be calculated from the seven SRD values for each model. As the CV underestimates the real variance, we may expect smaller CV variance than the true one. On the other hand, if we assign the variance for the ranking to full number of objects, e.g., we assign (much) larger variance than the true one. The two effects compensate each other; hence, less bias is expected. SRD corresponds to the principle of parsimony and provides an easy tool to rate the evaluation methods for genotoxicity.
Results
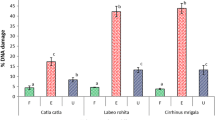
During our investigation, 34 specimens of chub were analyzed. Precisely from each specimen were analyzed three tissues: blood, liver, and gills. As shown in Table 1, results are presented for each site per tissue and for all three parameters: tail length, tail intensity, and Olive tail moment.
Regarding our nine different estimations of genotoxicity: tail length, intensity, and moment in erythrocytes (rel, rei, rem), liver cells (rll, rli, rlm), and gill cells (rgl, rgi, rgm), the SRD procedure showed that the tail moment and tail intensity are equally good parameters, more precisely that the SRD values were lower for the tail moment and tail intensity than for tail length (Fig. 2). According to our results, rel is the least reliable parameter. The overlapping is obvious: the ordering of rel cannot be distinguished from the random ranking. Close to the borderline case were rei, rll, and rgl (∼5 % probability of random ranking). Figure 3 illustrates the uncertainty of SRD values calculated by sevenfold cross validation. The figure naturally groups the similar ways for assessing genotoxicity. The last four ways (rll, rei, rgl, rel) are statistically not significant (at 5 % error level, according to sign test and Wilcoxon’s matched pair test.) Similarly, rgi and rlm as well as rlm and rgm are not significantly different according to the mentioned level and tests above.
SRD-CRRN test results of the data matrix given in Table 1. Y-axis, left-hand side, and X-axis—SRD-scaled values; Y-axis, right-hand side—relative frequencies (in percent) for the theoretical distribution function of random numbers
Discussion
Three measures of DNA migration are commonly used: tail length, OTM, and percent of the DNA in tail (tail intensity) [30–32]. Although OTM appeared to be the most statistically significant measurement [33], the inter-laboratory comparison of results seems to be difficult for this parameter [34]. OTM is calculated as a product of two factors: the percentage of DNA in the tail (tail intensity) and the distance between the intensity centroids (centers of gravity) of the head and the tail along the X-axis of the comet (Fig. 1). OTM calculation includes the distance between the intensity centroids of the head and the tail, which depends upon conditions of electrophoresis (e.g., electrophoresis time), and algorithms used to define the center of gravity of DNA distribution which vary among different software packages. Under these circumstances, it is advisable to use tail intensity for inter-laboratory comparisons [35–37]. When using derived measurements (e.g., tail moment), data on primary measurements (e.g., tail length and tail intensity) should also be presented in the analyses [8]. Tail length is considered unsatisfactory as a measure as it increases only while tails are first becoming established. Subsequently, the tail increases in intensity but not in length as the dose of damage increases. Tail length is also sensitive to the background or threshold setting of the image analysis program, as the end of the tail is defined by a certain excess of fluorescence over background [38]. Cells from different tissues or different species can differ substantially in tail length [35]. Still, tail length was, as a parameter, used in studies of genotoxicity in fish tissues [39–42]. Nevertheless, other parameters, especially, tail intensity, and tail moment, were more extensively used in similar works [43–45]. Kumaravel and Jha [33] recommend that both OTM and percent tail intensity could be used for scientific purposes.
Regarding the tissue selection for the analyses, three cell types were selected, following previous research: erythrocytes, liver cells, and gill cells [31, 42, 46–48]. However, different tissues can accumulate pollutants to different degrees, depending on their biochemical characteristics [49]. Thus, the choice of tissues for the comet assay should also be related to the ability of the tissue to take up and metabolize xenobiotics. In fish, a tissue often chosen to perform the comet assay is the blood, because it is easy to collect and there is no need for cellular dissociation. In this work, the reasons for the choice of gill and liver cells, besides the red blood cells, was that the gills represent the first organ which is in direct contact with water and, consequently, with the pollutants present in it, and the liver has a great role in xenobiotic metabolism and accumulation [50]. According to our results, erythrocytes showed the smallest values compared with other tissues. This may be due to the regular cycles of change of blood cells in the bloodstream, which indicates that blood could be used as a biomarker only for acute contaminations. Gills proved to be the most sensitive tissue for monitoring due to pronounced variation in values.
Regarding the tissue selection for further analysis, although gills proved to be the best biomarker, it is advisable to recommend all mentioned tissues, as each one is specific to the intensity and mode of stress response. As for the parameters obtained using Comet IV Computer Software, it was shown that tail moment and tail intensity are the most suitable according to SRD method, which is correlated with the recommendations of scientists in the field mentioned above.
Conclusion
The comet assay has been used in biomonitoring and has proven to be a sensitive system for the study of environmental genotoxicity and could apply to a variety of different species, tissues, and cells. Unfortunately, researchers use a lot of different parameters when representing results. With the emergence of image analyzing systems, the comet assay has several adopted parameters (tail length, intensity, and moment). All three parameters have been employed so far among which tail intensity and moment were the most frequent ones, which we also recommend in accordance with our findings exploring SRD method.
References
Singh NP, McCoy MT, Tice RR, Schneider EL (1988) Exp Cell Res 175:184–191
Kumaravel TS, Vilhar B, Faux SP, Jha AN (2009) Cell Biol Toxicol 25:53–64
Frenzilli G, Nigro M, Lyons BP (2009) Mutation Res 681:80–92
Meybodi AM, Mozdarani H (2009) Iranian Biomed J 13:1–8
Kolarević S, Knežević-Vukčević J, Paunović M, Tomović J, Gačić Z, Vuković-Gačić B (2011) Arch Biol Sci 63(4):1209–1217
Sunjog K, Gačić Z, Kolarević S, Višnjić-Jeftić Ž, Jarić I, Knežević-Vukčević J, Vuković-Gačić B, Lenhardt M (2012) The Scientific World Journal. doi:10.1100/2012/351074
Bernardeschi M, Guidi P, Scarcelli V, Frenzilli G, Nigro M (2010) Anal Bioanal Chem 396:619–623
Tice RR, Agurell E, Anderson D, Burlinson B, Hartmann A, Kobayashi H, Miyamae Y, Rojas E, Ryu JC, Sasaki YF (2000) Environ Molec Mutagenesis 35:206–221
Héberger K (2010) Trends Anal Chem 29:101–109
Héberger K, Kollár-Hunek K (2011) J Chemometr 25:151–158
Olive PL, Banath JP, Durand RE (1990) Radiation Res 122:86–94
Bolboaca SD, Jaentschi L (2010) Studia Univ Babes-Bolyai Chem 55:69–76
Djakovic-Sekulic T, Mandic A, Trisovic N, Uscumlic G (2012) Current Computer Aided Drug Design 8:3–9
Garkani-Nejad Z, Ahmadvand M (2011) Chromatographia 73:733–742
Kar S, Roy K (2012) Chemosphere 87:339–355
Liu XH, Ren YR, Zhou P, Shang ZC (2011) J Molec Struct 995:163–172
Ojha PK, Roy K (2011) Chemometr Intell Lab Syst 109:146–161
Vajna B, Farkas A, Pataki H, Zsigmond Z, Igricz T, Marosi G (2012) Anal Chim Acta 712:45–55
Héberger K, Zenkevich IG (2010) J Chromatogr A 1217:2895–2902
Bielicka-Daszkiewicz K, Voelkel A, Pietrzyńska M, Héberger K (2010) J Chromatogr A 1217:5564–5570
Gowen AA, Downey G, Esquerre C, O’Donnell CP (2011) J Chemomet 25:375–381
Vajna B, Patyi G, Nagy Z, Bodis A, Farkas A, Marosi G, Raman J (2011) Spectr 42:1977–1986
Kollár-Hunek K, Heszberger J, Kókai Z, Láng-Lázi M, Papp E (2008) J Chemometr 22:218–226
Sipos L, Kovács Z, Szöllösi D, Kókai Z, Dalmádi I, Fekete A (2011) J Chemometr 25:275–286
Rocha MJ, Ferreira PC, Reis PA, Cruzeiro C, Rocha E (2011) J Chromatogr Sci 49:695–701
Acanski MM, Vujic DN, Jovanovic-Santa S (2011) Chem Ind Chem Eng Quart 17:535–542
Balogh GT, Tarcsay A, Keseru GM (2012) J Pharm Biomed Anal 67–68:63–70
Tangni EK, Motte JC, Callebaut A, Chandelier A, De Schrijver M, Marnix L, Pussemier L (2011) Mycotoxin Res 27:105–113
Hastie T, Tibshirani R, Friedman J (2009) The elements of statistical learning: data mining, inference, and prediction, 2nd edn. Springer, New York
Çok I, Ulutaş OK, Okuşluk O, Durmaz E, Demir N (2011) The Scientific World Journal 11:1455–1461
Vincent-Hubert F, Arini A, Gourlay-Francé C (2011) Mutation Res 723:26–35
Morin B, Filatreau J, Vicquelin L, Barjhoux I, Guinel S, Leray-Forget J, Cachot J (2011) Anal Bioanal Chem 399:2235–2242
Kumaravel TS, Jha AN (2006) Mutation Res 605:7–16
Rosenberger A, Rössler U, Hornhardt S, Sauter W, Bickeböller H, Wichmann HE, Gomolka M (2011) DNA Repair 10:322–337
Collins AR, Oscoz AA, Brunborg G, Gaivao I, Giovannelli L, Kruszewski M, Smith CC, Štetina R (2008) Mutagenesis 23:143–151
Speit G, Hartmann A (1999) Meth Mol Biol 113:203–212
Burlinson B, Tice RR, Speit G, Agurell E, Brendler-Schwaab SY, Collins AR, Escobar P, Honma M, Kumaravel TS, Nakajima M, Sasaki YF, Thybaud V, Uno Y, Vasquez M, Hartmann A (2007) Mutation Res 627:31–35
Collins AR (2004) Mol Biotechn 26:249–261
Rajaguru P, Suba S, Palanivel M, Kalaiselvi K (2003) Environ Molec Mutagenesis 41:85–91
De Andrade VM, Da Silva J, Da Silva FR, Heuser VD, Dias JF, Yoneama ML, De Freitas TRO (2004) Environ Molec Mutagenesis 44:459–468
Pandrangi R, Petras M, Ralph S, Vrzoc M (1995) Environ Molec Mutagenesis 26:345–356
Alink GM, Quik JTK, Penders EJM, Spenkelink A, Rotteveel SGP, Maas JL, Hoogenboezem W (2007) Mutation Res 631:93–100
Sharma S, Nagpure NS, Kumar R, Pandey S, Srivastava SK, Singh PJ, Mathur PK (2007) Arch Environ Contam Toxicol 53:617–623
Wirzinger G, Weltje L, Gercken J, Sordyl H (2007) Mutation Res 628:19–30
Pandeya AK, Nagpurea NS, Trivedib SP, Kumara R, Kushwahaa B (2011) Mutation Res 726:209–214
Kopjar N, Mustafic P, Zanella D, Buj I, Caleta M, Marcic Z, Milic M, Dolenec Z, Mrakovcic M (2008) Folia Zool 57:120–130
Cotelle S, Férard JF (1999) Environ Molec Mutagenesis 34:246–255
Abd-Allaha GA, El-Fayoumib RI, Smith MJ, Heckmann RA, O'Neill KL (1999) Mutation Res 446:181–188
Suicmez M, Kayim M, Koseoglu D, Hasdemir E (2006) Bull Environ Contam Toxicol 77:551–558
Kilemade MF, Hartl MGJ, Sheehan D, Mothersill C, Van Pelt FNAM, O’Halloran J, O’Brien NM (2004) Environ Mol Mutagen 44:56–64
Acknowledgments
This study represents a part of the activities within project no. 173045, funded by the Ministry of Education and Science of the Republic of Serbia.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sunjog, K., Kolarević, S., Héberger, K. et al. Comparison of comet assay parameters for estimation of genotoxicity by sum of ranking differences. Anal Bioanal Chem 405, 4879–4885 (2013). https://doi.org/10.1007/s00216-013-6909-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-013-6909-y